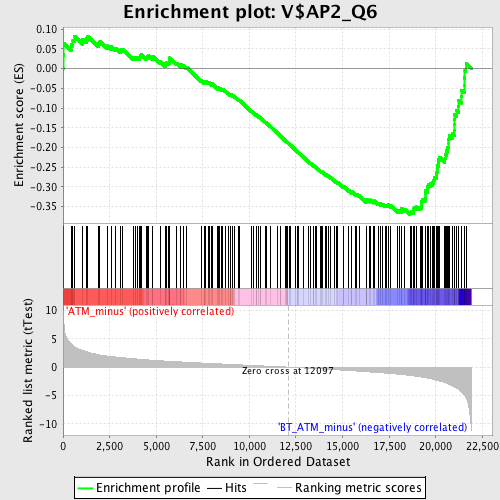
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_02_ATM_minus_versus_BT_ATM_minus.phenotype_ATM_minus_versus_BT_ATM_minus.cls #ATM_minus_versus_BT_ATM_minus.phenotype_ATM_minus_versus_BT_ATM_minus.cls #ATM_minus_versus_BT_ATM_minus_repos |
| Phenotype | phenotype_ATM_minus_versus_BT_ATM_minus.cls#ATM_minus_versus_BT_ATM_minus_repos |
| Upregulated in class | BT_ATM_minus |
| GeneSet | V$AP2_Q6 |
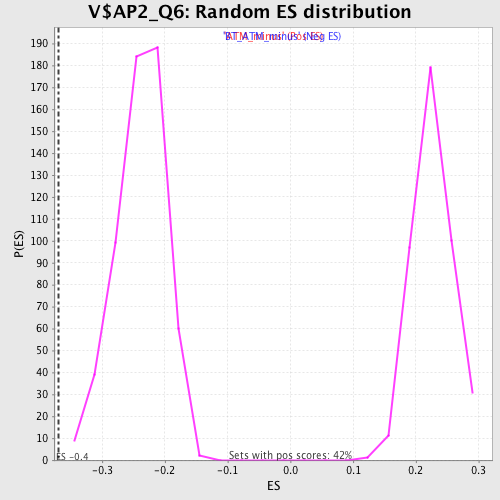
| Enrichment Score (ES) | -0.36997974 |
| Normalized Enrichment Score (NES) | -1.5559524 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.12307789 |
| FWER p-Value | 0.801 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CDC25A | 1417131_at 1417132_at 1445995_at | 8 | 9.163 | 0.0353 | No | ||
| 2 | SDC1 | 1415943_at 1415944_at 1437279_x_at 1448158_at | 44 | 7.679 | 0.0636 | No | ||
| 3 | SHB | 1434153_at | 450 | 4.012 | 0.0606 | No | ||
| 4 | ORC6L | 1417037_at | 527 | 3.842 | 0.0720 | No | ||
| 5 | EBAG9 | 1422669_at | 611 | 3.621 | 0.0823 | No | ||
| 6 | HDLBP | 1415987_at 1415988_at 1419806_at 1419869_s_at 1443546_at 1449615_s_at | 1051 | 2.946 | 0.0736 | No | ||
| 7 | CDK8 | 1460389_at | 1240 | 2.746 | 0.0756 | No | ||
| 8 | E2F3 | 1427462_at 1434564_at | 1331 | 2.642 | 0.0818 | No | ||
| 9 | RCOR2 | 1417302_at | 1897 | 2.194 | 0.0643 | No | ||
| 10 | CRLF1 | 1418476_at | 1978 | 2.142 | 0.0690 | No | ||
| 11 | HPN | 1420712_a_at | 2396 | 1.940 | 0.0574 | No | ||
| 12 | WNT1 | 1425377_at | 2583 | 1.866 | 0.0561 | No | ||
| 13 | GRK5 | 1446361_at 1446998_at 1449514_at | 2835 | 1.773 | 0.0515 | No | ||
| 14 | ARHGAP5 | 1423194_at 1450896_at 1450897_at 1457410_at | 3073 | 1.699 | 0.0472 | No | ||
| 15 | FAF1 | 1418242_at 1434370_s_at | 3164 | 1.673 | 0.0496 | No | ||
| 16 | PBX3 | 1421193_a_at 1440154_at 1442270_at 1447640_s_at | 3752 | 1.495 | 0.0284 | No | ||
| 17 | GALNT14 | 1453630_at | 3889 | 1.455 | 0.0278 | No | ||
| 18 | NRG1 | 1456524_at | 3978 | 1.429 | 0.0293 | No | ||
| 19 | EPHB1 | 1455188_at | 4098 | 1.396 | 0.0293 | No | ||
| 20 | PPP6C | 1424346_at 1424347_at | 4136 | 1.384 | 0.0330 | No | ||
| 21 | IGFBP7 | 1423584_at 1437804_at | 4198 | 1.367 | 0.0355 | No | ||
| 22 | RASA1 | 1426476_at 1426477_at 1426478_at 1438998_at | 4454 | 1.298 | 0.0288 | No | ||
| 23 | APLP1 | 1416134_at 1435857_s_at | 4521 | 1.280 | 0.0308 | No | ||
| 24 | PPM1E | 1434990_at 1445741_at | 4594 | 1.261 | 0.0324 | No | ||
| 25 | SLC30A5 | 1422497_at | 4817 | 1.204 | 0.0269 | No | ||
| 26 | RRBP1 | 1426123_a_at 1436660_at 1449221_a_at 1452767_at | 4819 | 1.203 | 0.0315 | No | ||
| 27 | INSM1 | 1421399_at 1455865_at | 5223 | 1.118 | 0.0173 | No | ||
| 28 | SFXN2 | 1439966_x_at 1440786_x_at | 5489 | 1.069 | 0.0093 | No | ||
| 29 | GATA6 | 1425463_at 1425464_at 1443081_at | 5511 | 1.063 | 0.0125 | No | ||
| 30 | EFEMP2 | 1417018_at 1447668_x_at | 5561 | 1.054 | 0.0143 | No | ||
| 31 | H1F0 | 1423702_at 1450522_a_at | 5675 | 1.031 | 0.0131 | No | ||
| 32 | RASIP1 | 1428016_a_at | 5705 | 1.027 | 0.0158 | No | ||
| 33 | MAPK6 | 1419168_at 1419169_at 1429963_at 1459146_at | 5714 | 1.023 | 0.0194 | No | ||
| 34 | WNT7A | 1423367_at 1447647_at 1458334_at | 5724 | 1.021 | 0.0230 | No | ||
| 35 | IGSF4 | 1417376_a_at 1417377_at 1417378_at 1431611_a_at | 5725 | 1.021 | 0.0270 | No | ||
| 36 | GPC4 | 1421088_at 1436514_at 1443620_at | 6105 | 0.955 | 0.0133 | No | ||
| 37 | CREB3L1 | 1419295_at | 6312 | 0.921 | 0.0074 | No | ||
| 38 | PLCB3 | 1448661_at | 6320 | 0.920 | 0.0106 | No | ||
| 39 | RAP1GDS1 | 1425266_a_at 1459768_x_at | 6476 | 0.888 | 0.0070 | No | ||
| 40 | KCNN2 | 1445676_at 1448927_at | 6623 | 0.864 | 0.0036 | No | ||
| 41 | SOX12 | 1439754_at 1455562_at 1456882_at | 7436 | 0.725 | -0.0309 | No | ||
| 42 | FKBP14 | 1433481_at 1458225_at | 7590 | 0.700 | -0.0352 | No | ||
| 43 | ARPC1A | 1416079_a_at | 7609 | 0.697 | -0.0333 | No | ||
| 44 | EXOSC8 | 1420115_at 1428291_at 1449738_s_at | 7653 | 0.690 | -0.0326 | No | ||
| 45 | KCNH5 | 1441742_at | 7783 | 0.670 | -0.0359 | No | ||
| 46 | BCL3 | 1418133_at | 7849 | 0.658 | -0.0364 | No | ||
| 47 | LHX2 | 1418317_at | 7986 | 0.633 | -0.0401 | No | ||
| 48 | TRIM28 | 1415869_a_at 1456308_x_at | 8006 | 0.631 | -0.0386 | No | ||
| 49 | MBD3 | 1417728_at | 8299 | 0.585 | -0.0497 | No | ||
| 50 | ATP5G2 | 1415980_at 1456128_at | 8330 | 0.581 | -0.0488 | No | ||
| 51 | HCST | 1419119_at | 8423 | 0.566 | -0.0509 | No | ||
| 52 | ACTR1A | 1415873_a_at 1458696_at 1459855_x_at | 8521 | 0.550 | -0.0532 | No | ||
| 53 | HS3ST3B1 | 1421331_at 1421332_at 1433977_at 1442816_at 1444055_at | 8570 | 0.542 | -0.0533 | No | ||
| 54 | LMX1A | 1421554_at | 8701 | 0.521 | -0.0572 | No | ||
| 55 | CTNND2 | 1422592_at 1438891_at 1443638_at 1445114_at 1447552_s_at 1456116_at | 8862 | 0.493 | -0.0627 | No | ||
| 56 | C1ORF24 | 1422567_at 1454942_at | 8998 | 0.475 | -0.0670 | No | ||
| 57 | ANGPTL2 | 1421002_at 1431848_at | 9002 | 0.474 | -0.0653 | No | ||
| 58 | POLD3 | 1426838_at 1426839_at 1443733_x_at 1459647_at | 9110 | 0.456 | -0.0684 | No | ||
| 59 | PLXNC1 | 1423213_at 1423214_at 1443433_at 1443434_s_at 1450905_at 1450906_at | 9198 | 0.444 | -0.0707 | No | ||
| 60 | SALL1 | 1450489_at | 9429 | 0.410 | -0.0797 | No | ||
| 61 | BSN | 1422023_at 1436123_at 1450467_at | 9480 | 0.402 | -0.0804 | No | ||
| 62 | ZDHHC2 | 1452655_at 1452656_at 1453481_at 1457204_at | 10117 | 0.306 | -0.1085 | No | ||
| 63 | SLC7A10 | 1421093_at | 10243 | 0.285 | -0.1131 | No | ||
| 64 | GPR3 | 1460275_at | 10361 | 0.265 | -0.1175 | No | ||
| 65 | ZDHHC14 | 1423668_at 1437614_x_at 1437616_x_at 1438151_x_at 1438619_x_at 1438975_x_at | 10363 | 0.265 | -0.1165 | No | ||
| 66 | BCL2 | 1422938_at 1427818_at 1437122_at 1440770_at 1443837_x_at 1457687_at | 10517 | 0.241 | -0.1226 | No | ||
| 67 | WFDC3 | 1430224_at 1460012_at | 10585 | 0.233 | -0.1247 | No | ||
| 68 | SFRP2 | 1448201_at | 10604 | 0.231 | -0.1247 | No | ||
| 69 | SOX2 | 1416967_at | 10886 | 0.188 | -0.1369 | No | ||
| 70 | KLC2 | 1418214_at 1431487_at | 10941 | 0.179 | -0.1386 | No | ||
| 71 | PDGFB | 1450413_at 1450414_at | 11145 | 0.147 | -0.1474 | No | ||
| 72 | ANKRD27 | 1425112_at 1454717_at | 11492 | 0.095 | -0.1629 | No | ||
| 73 | MMD | 1423488_at 1423489_at | 11648 | 0.070 | -0.1698 | No | ||
| 74 | MAPRE3 | 1427079_at | 11668 | 0.066 | -0.1704 | No | ||
| 75 | SUFU | 1420848_at 1431713_at 1440004_at 1446489_at | 11933 | 0.024 | -0.1824 | No | ||
| 76 | DUSP15 | 1426189_at 1435126_at | 12004 | 0.015 | -0.1856 | No | ||
| 77 | SLC25A12 | 1428440_at 1444489_at | 12015 | 0.013 | -0.1860 | No | ||
| 78 | RASAL2 | 1436910_at 1444671_at | 12064 | 0.005 | -0.1882 | No | ||
| 79 | HCN1 | 1450193_at | 12067 | 0.005 | -0.1883 | No | ||
| 80 | CENPB | 1426051_a_at 1435944_s_at | 12150 | -0.008 | -0.1920 | No | ||
| 81 | WDR12 | 1417270_at 1448646_at 1459762_x_at | 12204 | -0.016 | -0.1944 | No | ||
| 82 | STAG2 | 1421849_at 1450396_at | 12491 | -0.069 | -0.2073 | No | ||
| 83 | WAC | 1437426_at 1439249_at 1444241_at 1451494_at 1451495_at 1457559_at 1458471_at | 12582 | -0.088 | -0.2110 | No | ||
| 84 | FMNL1 | 1445636_at 1450212_at | 12585 | -0.088 | -0.2108 | No | ||
| 85 | UBE2B | 1423106_at 1423107_at 1430177_at 1439477_at 1460132_at | 12633 | -0.095 | -0.2126 | No | ||
| 86 | FZD5 | 1422937_at 1430809_at | 12928 | -0.144 | -0.2255 | No | ||
| 87 | GUCY1A2 | 1459336_at | 13191 | -0.188 | -0.2369 | No | ||
| 88 | VPS35 | 1415783_at 1415784_at | 13294 | -0.206 | -0.2407 | No | ||
| 89 | WNT3 | 1450763_x_at | 13441 | -0.230 | -0.2466 | No | ||
| 90 | MAFB | 1451715_at 1451716_at | 13457 | -0.233 | -0.2463 | No | ||
| 91 | GATA2 | 1428816_a_at 1450333_a_at | 13544 | -0.247 | -0.2493 | No | ||
| 92 | PITX3 | 1449917_at | 13617 | -0.262 | -0.2516 | No | ||
| 93 | CRBN | 1423094_at 1423095_s_at 1455734_at | 13837 | -0.298 | -0.2605 | No | ||
| 94 | ARID1A | 1447104_at | 13897 | -0.307 | -0.2620 | No | ||
| 95 | VEGFB | 1451803_a_at | 13924 | -0.312 | -0.2620 | No | ||
| 96 | CUGBP1 | 1423932_at 1425932_a_at 1426407_at 1426408_at 1427413_a_at | 14115 | -0.344 | -0.2694 | No | ||
| 97 | APBA1 | 1437940_at 1455369_at 1459605_at | 14119 | -0.345 | -0.2682 | No | ||
| 98 | CNNM2 | 1423475_at 1451007_at 1458132_at | 14224 | -0.366 | -0.2716 | No | ||
| 99 | MAST2 | 1417324_at 1440321_at 1441847_at 1444813_at 1456927_at 1459558_at | 14369 | -0.393 | -0.2767 | No | ||
| 100 | MAPRE1 | 1422764_at 1422765_at 1428819_at 1428820_at 1450740_a_at | 14578 | -0.434 | -0.2845 | No | ||
| 101 | FBXL11 | 1435329_at 1443336_at 1444358_at 1455942_at 1458449_at | 14681 | -0.455 | -0.2875 | No | ||
| 102 | RAB10 | 1422664_at 1429296_at 1442463_at 1453095_at | 14755 | -0.470 | -0.2890 | No | ||
| 103 | POU2F1 | 1419716_a_at 1427695_a_at 1427835_at | 15046 | -0.533 | -0.3002 | No | ||
| 104 | GAS7 | 1417859_at 1431400_a_at 1445685_at | 15068 | -0.536 | -0.2991 | No | ||
| 105 | ICA1 | 1417901_a_at 1431644_a_at | 15350 | -0.593 | -0.3097 | No | ||
| 106 | CTCF | 1418330_at 1449042_at | 15489 | -0.623 | -0.3136 | No | ||
| 107 | SP4 | 1421504_at 1437508_at | 15502 | -0.626 | -0.3118 | No | ||
| 108 | FGF11 | 1421793_at 1439959_at | 15699 | -0.672 | -0.3181 | No | ||
| 109 | MNT | 1418192_at | 15763 | -0.683 | -0.3184 | No | ||
| 110 | PABPC1 | 1418883_a_at 1419500_at 1453840_at | 15919 | -0.719 | -0.3227 | No | ||
| 111 | HMGN2 | 1433507_a_at | 16269 | -0.804 | -0.3356 | No | ||
| 112 | LHX1 | 1421951_at 1450428_at | 16302 | -0.812 | -0.3339 | No | ||
| 113 | ZZZ3 | 1434332_at 1438487_s_at 1441748_at | 16313 | -0.814 | -0.3312 | No | ||
| 114 | THRAP1 | 1436904_at 1438725_at 1444068_at 1446840_at 1453160_at 1453161_at | 16470 | -0.854 | -0.3351 | No | ||
| 115 | CREB3 | 1419979_s_at 1424740_at 1424741_s_at | 16503 | -0.860 | -0.3332 | No | ||
| 116 | COL11A2 | 1423578_at | 16671 | -0.905 | -0.3374 | No | ||
| 117 | UBE2S | 1416726_s_at 1430962_at | 16712 | -0.916 | -0.3356 | No | ||
| 118 | SIRT1 | 1418640_at 1458538_at | 16926 | -0.962 | -0.3417 | No | ||
| 119 | CBX6 | 1424407_s_at 1429290_at | 17037 | -0.985 | -0.3429 | No | ||
| 120 | TERF2 | 1421147_at 1455651_at 1459650_at | 17135 | -1.007 | -0.3434 | No | ||
| 121 | SP3 | 1431804_a_at 1459360_at | 17292 | -1.035 | -0.3466 | No | ||
| 122 | SPATA6 | 1418650_at 1418651_at 1438023_at 1445355_at | 17380 | -1.057 | -0.3465 | No | ||
| 123 | TEAD2 | 1448519_at | 17451 | -1.075 | -0.3455 | No | ||
| 124 | CRK | 1416201_at 1425855_a_at 1436835_at 1448248_at 1460176_at | 17587 | -1.112 | -0.3474 | No | ||
| 125 | SPTBN4 | 1425116_a_at | 17959 | -1.222 | -0.3597 | No | ||
| 126 | CNNM4 | 1421635_at 1431233_at | 18074 | -1.259 | -0.3600 | No | ||
| 127 | HOXB2 | 1444204_at 1449397_at | 18176 | -1.292 | -0.3596 | No | ||
| 128 | MYADM | 1423321_at 1439389_s_at | 18189 | -1.294 | -0.3551 | No | ||
| 129 | PPFIA1 | 1435861_at 1455478_at | 18345 | -1.348 | -0.3570 | No | ||
| 130 | DNMT3A | 1423063_at 1423064_at 1423065_at 1423066_at 1442309_at 1460324_at | 18628 | -1.463 | -0.3643 | Yes | ||
| 131 | IRF1 | 1448436_a_at | 18702 | -1.496 | -0.3618 | Yes | ||
| 132 | NXPH3 | 1419710_at | 18826 | -1.548 | -0.3615 | Yes | ||
| 133 | PPP2R5E | 1428461_at 1428462_at 1428463_a_at 1439401_x_at 1443533_at 1443949_at 1446356_at 1452788_at | 18837 | -1.551 | -0.3559 | Yes | ||
| 134 | GLTP | 1419027_s_at | 18876 | -1.571 | -0.3515 | Yes | ||
| 135 | EP300 | 1434765_at | 18998 | -1.622 | -0.3508 | Yes | ||
| 136 | RAB35 | 1433922_at | 19168 | -1.693 | -0.3519 | Yes | ||
| 137 | PLAGL2 | 1417517_at 1417518_at 1417519_at 1442235_at | 19245 | -1.730 | -0.3487 | Yes | ||
| 138 | MYST2 | 1433433_at 1440746_at 1447631_at | 19257 | -1.738 | -0.3424 | Yes | ||
| 139 | NRF1 | 1420978_at 1424787_a_at 1434627_at 1439545_at 1445914_at | 19264 | -1.741 | -0.3359 | Yes | ||
| 140 | SLC25A28 | 1424776_a_at | 19317 | -1.764 | -0.3315 | Yes | ||
| 141 | LDB1 | 1452024_a_at | 19436 | -1.817 | -0.3298 | Yes | ||
| 142 | CNNM1 | 1420354_at 1441312_at 1445979_at | 19457 | -1.834 | -0.3236 | Yes | ||
| 143 | ACTB | 1419734_at 1436722_a_at 1440365_at AFFX-b-ActinMur/M12481_3_at AFFX-b-ActinMur/M12481_5_at AFFX-b-ActinMur/M12481_M_at | 19459 | -1.836 | -0.3165 | Yes | ||
| 144 | FBXO11 | 1434108_at | 19477 | -1.843 | -0.3101 | Yes | ||
| 145 | CPEB4 | 1420617_at 1420618_at 1449931_at | 19564 | -1.895 | -0.3067 | Yes | ||
| 146 | TLN1 | 1436042_at 1448402_at 1457782_at | 19574 | -1.902 | -0.2997 | Yes | ||
| 147 | NOTCH3 | 1421964_at 1421965_s_at 1443300_at | 19636 | -1.951 | -0.2949 | Yes | ||
| 148 | CDK9 | 1417269_at 1457240_at | 19725 | -2.022 | -0.2911 | Yes | ||
| 149 | TRIM39 | 1418851_at | 19831 | -2.106 | -0.2877 | Yes | ||
| 150 | UBTF | 1453097_a_at 1455490_at 1460304_a_at | 19907 | -2.171 | -0.2827 | Yes | ||
| 151 | USP48 | 1419277_at 1419278_at 1424056_at | 19934 | -2.198 | -0.2754 | Yes | ||
| 152 | ALG5 | 1424160_at 1430854_at | 20036 | -2.285 | -0.2711 | Yes | ||
| 153 | PDE5A | 1445963_at 1455970_at | 20069 | -2.312 | -0.2636 | Yes | ||
| 154 | MTA2 | 1423165_a_at | 20085 | -2.319 | -0.2552 | Yes | ||
| 155 | RASGRP2 | 1417804_at 1438932_at 1438933_x_at 1442264_at 1449613_at | 20092 | -2.330 | -0.2464 | Yes | ||
| 156 | ARL6IP2 | 1416793_at 1416794_at 1431197_at 1435594_at | 20137 | -2.374 | -0.2392 | Yes | ||
| 157 | CAMK4 | 1421941_at 1426167_a_at 1439843_at 1442822_at 1452572_at | 20147 | -2.379 | -0.2304 | Yes | ||
| 158 | ATF7IP | 1446323_at 1449192_at 1454973_at 1459096_at | 20221 | -2.430 | -0.2243 | Yes | ||
| 159 | CNP | 1418980_a_at 1437341_x_at 1449296_a_at | 20506 | -2.707 | -0.2268 | Yes | ||
| 160 | DNTTIP1 | 1417666_at 1456985_at | 20525 | -2.734 | -0.2170 | Yes | ||
| 161 | POFUT1 | 1443881_at 1450137_at 1456323_at | 20582 | -2.804 | -0.2086 | Yes | ||
| 162 | RAB3IP | 1434306_at 1436103_at | 20652 | -2.892 | -0.2006 | Yes | ||
| 163 | DDX5 | 1419653_a_at 1423645_a_at 1433809_at 1433810_x_at 1438371_x_at 1454793_x_at | 20673 | -2.918 | -0.1901 | Yes | ||
| 164 | SMOX | 1424268_at 1446509_at | 20676 | -2.921 | -0.1789 | Yes | ||
| 165 | ATP5H | 1423676_at 1435112_a_at | 20724 | -2.996 | -0.1694 | Yes | ||
| 166 | PHF12 | 1434922_at 1453579_at | 20886 | -3.262 | -0.1641 | Yes | ||
| 167 | FST | 1421365_at 1434458_at | 21001 | -3.464 | -0.1558 | Yes | ||
| 168 | CRSP7 | 1452282_at 1458377_at | 21009 | -3.477 | -0.1426 | Yes | ||
| 169 | RAB22A | 1424503_at 1424504_at 1451379_at 1454988_s_at | 21013 | -3.483 | -0.1292 | Yes | ||
| 170 | BAHD1 | 1433703_s_at | 21028 | -3.512 | -0.1162 | Yes | ||
| 171 | PSCD2 | 1450103_a_at | 21138 | -3.734 | -0.1066 | Yes | ||
| 172 | RELB | 1417856_at | 21234 | -3.907 | -0.0958 | Yes | ||
| 173 | BRPF1 | 1460402_at | 21250 | -3.958 | -0.0811 | Yes | ||
| 174 | CAMTA2 | 1426901_s_at 1452229_at | 21389 | -4.291 | -0.0707 | Yes | ||
| 175 | BANP | 1452462_a_at | 21404 | -4.358 | -0.0544 | Yes | ||
| 176 | REV3L | 1424632_a_at 1426194_x_at 1451960_a_at | 21540 | -4.857 | -0.0417 | Yes | ||
| 177 | GALNT10 | 1418194_at 1418195_at 1440493_at | 21544 | -4.877 | -0.0229 | Yes | ||
| 178 | SMARCD2 | 1426192_at 1448400_a_at 1448401_at | 21578 | -5.047 | -0.0048 | Yes | ||
| 179 | SPRY2 | 1421656_at 1436584_at | 21635 | -5.415 | 0.0137 | Yes |