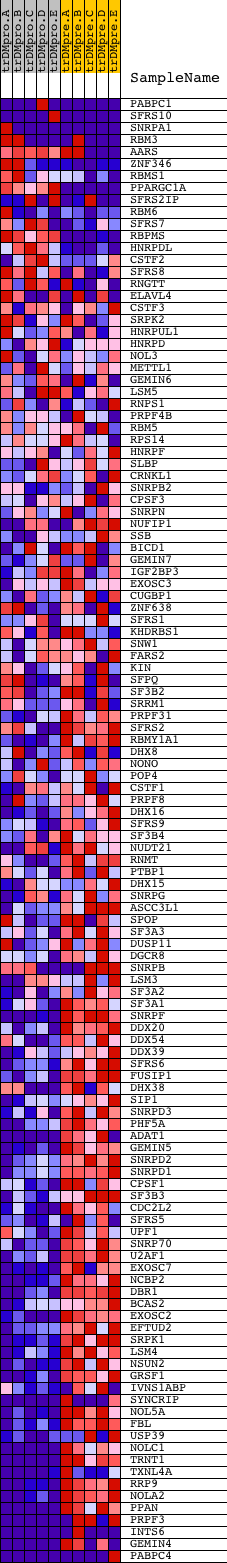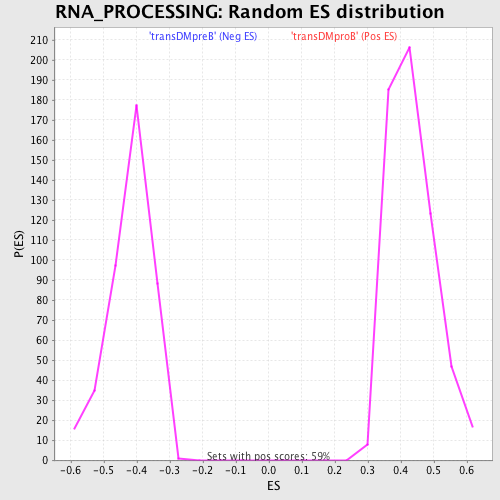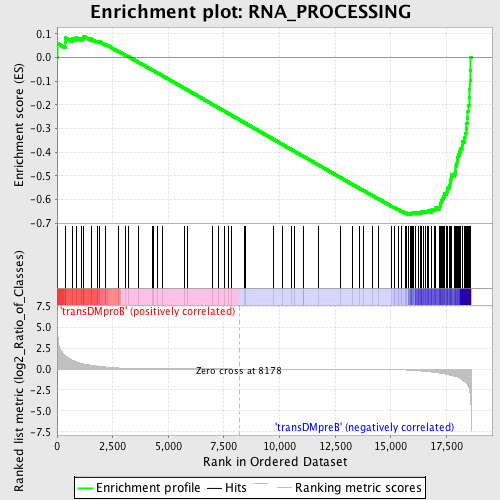
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_transDMpreB.phenotype_transDMproB_versus_transDMpreB.cls #transDMproB_versus_transDMpreB.phenotype_transDMproB_versus_transDMpreB.cls #transDMproB_versus_transDMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_transDMpreB.cls#transDMproB_versus_transDMpreB_repos |
| Upregulated in class | transDMpreB |
| GeneSet | RNA_PROCESSING |
| Enrichment Score (ES) | -0.6635379 |
| Normalized Enrichment Score (NES) | -1.577526 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.16727784 |
| FWER p-Value | 0.699 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PABPC1 | 5219 9522 9523 23572 | 19 | 4.416 | 0.0592 | No | ||
| 2 | SFRS10 | 9818 1722 5441 | 368 | 1.583 | 0.0620 | No | ||
| 3 | SNRPA1 | 1373 18216 | 381 | 1.570 | 0.0827 | No | ||
| 4 | RBM3 | 5366 2625 | 690 | 1.034 | 0.0802 | No | ||
| 5 | AARS | 10630 6151 | 853 | 0.873 | 0.0833 | No | ||
| 6 | ZNF346 | 3163 6517 3193 | 1089 | 0.659 | 0.0796 | No | ||
| 7 | RBMS1 | 14580 | 1190 | 0.598 | 0.0823 | No | ||
| 8 | PPARGC1A | 16533 | 1193 | 0.597 | 0.0904 | No | ||
| 9 | SFRS2IP | 7794 13009 | 1523 | 0.451 | 0.0787 | No | ||
| 10 | RBM6 | 9707 3095 3063 2995 | 1803 | 0.344 | 0.0683 | No | ||
| 11 | SFRS7 | 22889 | 1898 | 0.314 | 0.0675 | No | ||
| 12 | RBPMS | 5369 | 2196 | 0.231 | 0.0546 | No | ||
| 13 | HNRPDL | 11947 3551 6954 | 2768 | 0.118 | 0.0253 | No | ||
| 14 | CSTF2 | 24257 | 3071 | 0.083 | 0.0102 | No | ||
| 15 | SFRS8 | 10543 6089 | 3195 | 0.072 | 0.0045 | No | ||
| 16 | RNGTT | 10769 2354 6272 | 3673 | 0.041 | -0.0208 | No | ||
| 17 | ELAVL4 | 15805 4889 9137 | 4280 | 0.021 | -0.0532 | No | ||
| 18 | CSTF3 | 10449 6002 | 4328 | 0.020 | -0.0555 | No | ||
| 19 | SRPK2 | 5513 | 4494 | 0.017 | -0.0642 | No | ||
| 20 | HNRPUL1 | 6118 17923 1608 10577 1345 | 4740 | 0.014 | -0.0772 | No | ||
| 21 | HNRPD | 8645 | 5733 | 0.007 | -0.1308 | No | ||
| 22 | NOL3 | 18496 915 921 | 5882 | 0.007 | -0.1387 | No | ||
| 23 | METTL1 | 19860 3400 3404 | 6967 | 0.003 | -0.1972 | No | ||
| 24 | GEMIN6 | 12492 7415 | 7238 | 0.002 | -0.2118 | No | ||
| 25 | LSM5 | 7286 | 7525 | 0.002 | -0.2272 | No | ||
| 26 | RNPS1 | 9730 23361 | 7696 | 0.001 | -0.2364 | No | ||
| 27 | PRPF4B | 3208 5295 | 7854 | 0.001 | -0.2449 | No | ||
| 28 | RBM5 | 13507 3016 | 8413 | -0.001 | -0.2751 | No | ||
| 29 | RPS14 | 9751 | 8450 | -0.001 | -0.2770 | No | ||
| 30 | HNRPF | 13607 | 9741 | -0.004 | -0.3467 | No | ||
| 31 | SLBP | 9822 | 10114 | -0.005 | -0.3668 | No | ||
| 32 | CRNKL1 | 14407 | 10515 | -0.006 | -0.3883 | No | ||
| 33 | SNRPB2 | 5470 | 10690 | -0.006 | -0.3976 | No | ||
| 34 | CPSF3 | 7038 | 11073 | -0.007 | -0.4182 | No | ||
| 35 | SNRPN | 1715 9844 5471 | 11086 | -0.007 | -0.4187 | No | ||
| 36 | NUFIP1 | 21955 | 11765 | -0.010 | -0.4552 | No | ||
| 37 | SSB | 907 14998 | 12757 | -0.016 | -0.5086 | No | ||
| 38 | BICD1 | 8657 4447 | 13288 | -0.020 | -0.5370 | No | ||
| 39 | GEMIN7 | 17944 | 13580 | -0.024 | -0.5524 | No | ||
| 40 | IGF2BP3 | 4699 8932 | 13754 | -0.026 | -0.5614 | No | ||
| 41 | EXOSC3 | 15890 | 14184 | -0.034 | -0.5841 | No | ||
| 42 | CUGBP1 | 2805 8819 4576 2924 | 14465 | -0.041 | -0.5987 | No | ||
| 43 | ZNF638 | 9476 | 15023 | -0.063 | -0.6280 | No | ||
| 44 | SFRS1 | 8492 | 15175 | -0.071 | -0.6352 | No | ||
| 45 | KHDRBS1 | 5405 9778 | 15336 | -0.081 | -0.6427 | No | ||
| 46 | SNW1 | 7282 | 15501 | -0.094 | -0.6503 | No | ||
| 47 | FARS2 | 21666 | 15647 | -0.110 | -0.6566 | No | ||
| 48 | KIN | 15120 | 15687 | -0.114 | -0.6572 | No | ||
| 49 | SFPQ | 12936 | 15789 | -0.127 | -0.6609 | No | ||
| 50 | SF3B2 | 23974 | 15809 | -0.129 | -0.6602 | No | ||
| 51 | SRRM1 | 6958 | 15872 | -0.139 | -0.6616 | Yes | ||
| 52 | PRPF31 | 7594 | 15904 | -0.143 | -0.6614 | Yes | ||
| 53 | SFRS2 | 9807 20136 | 15923 | -0.146 | -0.6603 | Yes | ||
| 54 | RBMY1A1 | 5368 | 15926 | -0.146 | -0.6585 | Yes | ||
| 55 | DHX8 | 20649 | 15940 | -0.148 | -0.6572 | Yes | ||
| 56 | NONO | 11993 | 15961 | -0.152 | -0.6562 | Yes | ||
| 57 | POP4 | 7261 | 16030 | -0.163 | -0.6576 | Yes | ||
| 58 | CSTF1 | 12515 | 16090 | -0.175 | -0.6584 | Yes | ||
| 59 | PRPF8 | 20780 1371 | 16105 | -0.178 | -0.6567 | Yes | ||
| 60 | DHX16 | 23247 | 16112 | -0.179 | -0.6546 | Yes | ||
| 61 | SFRS9 | 16731 | 16235 | -0.203 | -0.6585 | Yes | ||
| 62 | SF3B4 | 22269 | 16255 | -0.207 | -0.6567 | Yes | ||
| 63 | NUDT21 | 12665 | 16345 | -0.226 | -0.6584 | Yes | ||
| 64 | RNMT | 7501 | 16372 | -0.234 | -0.6566 | Yes | ||
| 65 | PTBP1 | 5303 | 16386 | -0.238 | -0.6540 | Yes | ||
| 66 | DHX15 | 8842 | 16394 | -0.240 | -0.6511 | Yes | ||
| 67 | SNRPG | 12622 | 16463 | -0.258 | -0.6513 | Yes | ||
| 68 | ASCC3L1 | 14867 | 16562 | -0.285 | -0.6527 | Yes | ||
| 69 | SPOP | 5498 | 16637 | -0.305 | -0.6526 | Yes | ||
| 70 | SF3A3 | 16091 | 16675 | -0.313 | -0.6503 | Yes | ||
| 71 | DUSP11 | 17086 7785 | 16706 | -0.321 | -0.6476 | Yes | ||
| 72 | DGCR8 | 22644 | 16820 | -0.351 | -0.6489 | Yes | ||
| 73 | SNRPB | 9842 5469 2736 | 16845 | -0.360 | -0.6453 | Yes | ||
| 74 | LSM3 | 12565 | 16847 | -0.360 | -0.6404 | Yes | ||
| 75 | SF3A2 | 19938 | 16967 | -0.396 | -0.6414 | Yes | ||
| 76 | SF3A1 | 7450 | 17019 | -0.417 | -0.6385 | Yes | ||
| 77 | SNRPF | 7645 | 17025 | -0.419 | -0.6331 | Yes | ||
| 78 | DDX20 | 15213 | 17173 | -0.466 | -0.6347 | Yes | ||
| 79 | DDX54 | 7778 | 17209 | -0.482 | -0.6300 | Yes | ||
| 80 | DDX39 | 18551 | 17216 | -0.486 | -0.6237 | Yes | ||
| 81 | SFRS6 | 14751 | 17246 | -0.503 | -0.6184 | Yes | ||
| 82 | FUSIP1 | 4715 16036 | 17254 | -0.507 | -0.6119 | Yes | ||
| 83 | DHX38 | 48 | 17271 | -0.512 | -0.6057 | Yes | ||
| 84 | SIP1 | 21263 | 17307 | -0.533 | -0.6004 | Yes | ||
| 85 | SNRPD3 | 12514 | 17340 | -0.547 | -0.5946 | Yes | ||
| 86 | PHF5A | 22194 | 17362 | -0.559 | -0.5881 | Yes | ||
| 87 | ADAT1 | 11400 6633 | 17411 | -0.582 | -0.5828 | Yes | ||
| 88 | GEMIN5 | 20439 | 17423 | -0.586 | -0.5754 | Yes | ||
| 89 | SNRPD2 | 8412 | 17505 | -0.632 | -0.5712 | Yes | ||
| 90 | SNRPD1 | 23622 | 17533 | -0.647 | -0.5638 | Yes | ||
| 91 | CPSF1 | 22241 | 17547 | -0.654 | -0.5556 | Yes | ||
| 92 | SF3B3 | 18746 | 17567 | -0.669 | -0.5475 | Yes | ||
| 93 | CDC2L2 | 15964 2419 | 17650 | -0.712 | -0.5422 | Yes | ||
| 94 | SFRS5 | 9808 2062 | 17659 | -0.716 | -0.5329 | Yes | ||
| 95 | UPF1 | 9718 18855 | 17675 | -0.729 | -0.5238 | Yes | ||
| 96 | SNRP70 | 1186 | 17695 | -0.739 | -0.5147 | Yes | ||
| 97 | U2AF1 | 8431 | 17718 | -0.750 | -0.5057 | Yes | ||
| 98 | EXOSC7 | 19256 | 17740 | -0.768 | -0.4964 | Yes | ||
| 99 | NCBP2 | 12643 | 17873 | -0.871 | -0.4916 | Yes | ||
| 100 | DBR1 | 19341 | 17894 | -0.890 | -0.4806 | Yes | ||
| 101 | BCAS2 | 12662 | 17926 | -0.912 | -0.4698 | Yes | ||
| 102 | EXOSC2 | 15044 | 17927 | -0.912 | -0.4574 | Yes | ||
| 103 | EFTUD2 | 1219 20203 | 17956 | -0.931 | -0.4462 | Yes | ||
| 104 | SRPK1 | 23049 | 17977 | -0.952 | -0.4343 | Yes | ||
| 105 | LSM4 | 18583 | 17992 | -0.964 | -0.4219 | Yes | ||
| 106 | NSUN2 | 21610 | 18022 | -1.003 | -0.4098 | Yes | ||
| 107 | GRSF1 | 16487 | 18080 | -1.073 | -0.3982 | Yes | ||
| 108 | IVNS1ABP | 4384 4050 | 18124 | -1.135 | -0.3851 | Yes | ||
| 109 | SYNCRIP | 3078 3035 3107 | 18203 | -1.259 | -0.3721 | Yes | ||
| 110 | NOL5A | 12474 | 18233 | -1.313 | -0.3558 | Yes | ||
| 111 | FBL | 8955 | 18317 | -1.488 | -0.3400 | Yes | ||
| 112 | USP39 | 1116 1083 11373 | 18335 | -1.516 | -0.3202 | Yes | ||
| 113 | NOLC1 | 7704 | 18381 | -1.612 | -0.3007 | Yes | ||
| 114 | TRNT1 | 17344 | 18417 | -1.731 | -0.2790 | Yes | ||
| 115 | TXNL4A | 6567 11329 | 18451 | -1.865 | -0.2553 | Yes | ||
| 116 | RRP9 | 19328 | 18464 | -1.970 | -0.2291 | Yes | ||
| 117 | NOLA2 | 11972 | 18485 | -2.087 | -0.2017 | Yes | ||
| 118 | PPAN | 19546 | 18541 | -2.549 | -0.1699 | Yes | ||
| 119 | PRPF3 | 12899 12898 1877 7703 | 18551 | -2.691 | -0.1337 | Yes | ||
| 120 | INTS6 | 5184 126 | 18558 | -2.798 | -0.0959 | Yes | ||
| 121 | GEMIN4 | 6592 6591 | 18571 | -3.083 | -0.0545 | Yes | ||
| 122 | PABPC4 | 10510 2418 2548 | 18600 | -4.171 | 0.0009 | Yes |