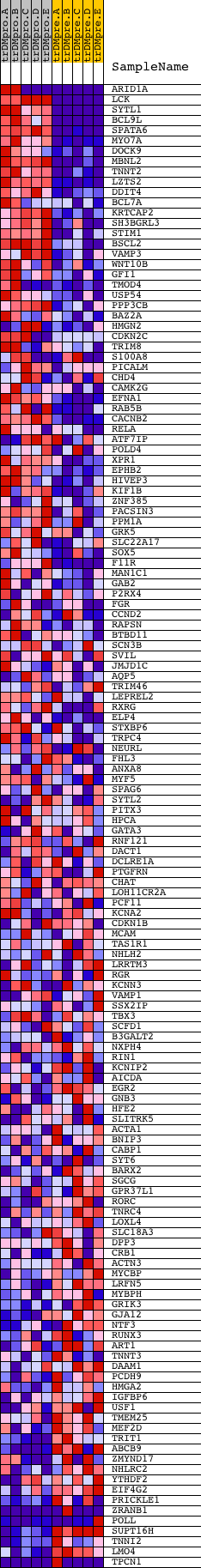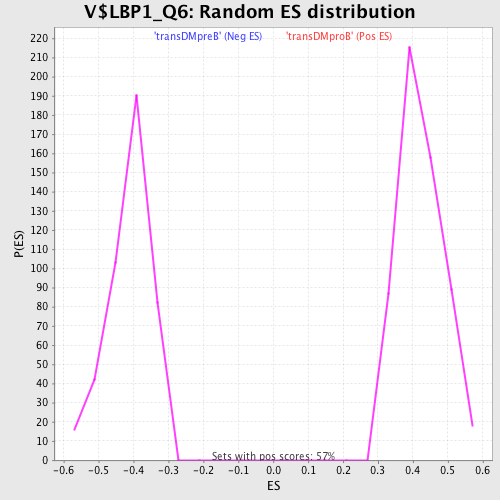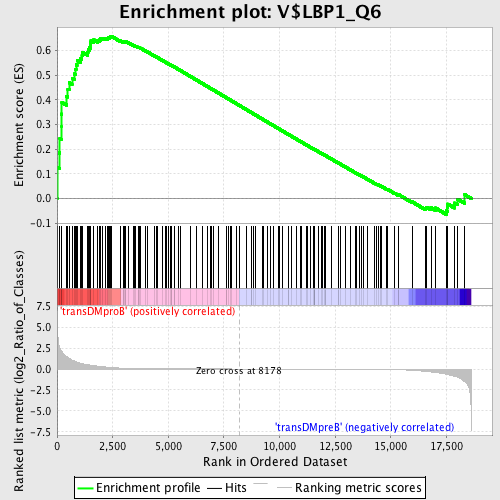
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_transDMpreB.phenotype_transDMproB_versus_transDMpreB.cls #transDMproB_versus_transDMpreB.phenotype_transDMproB_versus_transDMpreB.cls #transDMproB_versus_transDMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_transDMpreB.cls#transDMproB_versus_transDMpreB_repos |
| Upregulated in class | transDMproB |
| GeneSet | V$LBP1_Q6 |
| Enrichment Score (ES) | 0.65854615 |
| Normalized Enrichment Score (NES) | 1.5607365 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.06439653 |
| FWER p-Value | 0.453 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ARID1A | 8214 2501 13554 | 6 | 5.419 | 0.1278 | Yes | ||
| 2 | LCK | 15746 | 101 | 2.651 | 0.1854 | Yes | ||
| 3 | SYTL1 | 15728 | 124 | 2.463 | 0.2424 | Yes | ||
| 4 | BCL9L | 19475 | 184 | 2.209 | 0.2915 | Yes | ||
| 5 | SPATA6 | 16134 | 190 | 2.187 | 0.3429 | Yes | ||
| 6 | MYO7A | 9439 | 214 | 2.051 | 0.3902 | Yes | ||
| 7 | DOCK9 | 8369 | 426 | 1.512 | 0.4145 | Yes | ||
| 8 | MBNL2 | 21932 | 484 | 1.376 | 0.4440 | Yes | ||
| 9 | TNNT2 | 14113 | 547 | 1.243 | 0.4700 | Yes | ||
| 10 | LZTS2 | 10362 | 674 | 1.057 | 0.4882 | Yes | ||
| 11 | DDIT4 | 19762 | 772 | 0.948 | 0.5053 | Yes | ||
| 12 | BCL7A | 8063 | 837 | 0.890 | 0.5229 | Yes | ||
| 13 | KRTCAP2 | 15541 | 855 | 0.871 | 0.5426 | Yes | ||
| 14 | SH3BGRL3 | 15722 | 897 | 0.832 | 0.5600 | Yes | ||
| 15 | STIM1 | 18164 | 1042 | 0.691 | 0.5686 | Yes | ||
| 16 | BSCL2 | 23940 | 1117 | 0.642 | 0.5798 | Yes | ||
| 17 | VAMP3 | 15660 | 1147 | 0.619 | 0.5928 | Yes | ||
| 18 | WNT10B | 5874 | 1369 | 0.514 | 0.5930 | Yes | ||
| 19 | GFI1 | 16451 | 1392 | 0.506 | 0.6038 | Yes | ||
| 20 | TMOD4 | 1816 1884 15509 | 1455 | 0.482 | 0.6118 | Yes | ||
| 21 | USP54 | 21910 | 1494 | 0.463 | 0.6207 | Yes | ||
| 22 | PPP3CB | 5285 | 1499 | 0.461 | 0.6314 | Yes | ||
| 23 | BAZ2A | 4371 | 1521 | 0.451 | 0.6409 | Yes | ||
| 24 | HMGN2 | 9095 | 1653 | 0.396 | 0.6432 | Yes | ||
| 25 | CDKN2C | 15806 2321 | 1832 | 0.332 | 0.6414 | Yes | ||
| 26 | TRIM8 | 23824 | 1921 | 0.310 | 0.6440 | Yes | ||
| 27 | S100A8 | 15527 | 1958 | 0.298 | 0.6491 | Yes | ||
| 28 | PICALM | 18191 | 2061 | 0.266 | 0.6499 | Yes | ||
| 29 | CHD4 | 8418 17281 4225 | 2169 | 0.239 | 0.6497 | Yes | ||
| 30 | CAMK2G | 21905 | 2283 | 0.210 | 0.6486 | Yes | ||
| 31 | EFNA1 | 15279 | 2291 | 0.208 | 0.6531 | Yes | ||
| 32 | RAB5B | 359 19591 3430 | 2355 | 0.191 | 0.6542 | Yes | ||
| 33 | CACNB2 | 4466 8678 | 2406 | 0.180 | 0.6558 | Yes | ||
| 34 | RELA | 23783 | 2432 | 0.175 | 0.6585 | Yes | ||
| 35 | ATF7IP | 12028 13038 | 2835 | 0.108 | 0.6393 | No | ||
| 36 | POLD4 | 12822 | 2969 | 0.094 | 0.6344 | No | ||
| 37 | XPR1 | 5383 | 2977 | 0.093 | 0.6362 | No | ||
| 38 | EPHB2 | 4675 2440 8910 | 3035 | 0.087 | 0.6351 | No | ||
| 39 | HIVEP3 | 4973 | 3051 | 0.085 | 0.6364 | No | ||
| 40 | KIF1B | 15671 2476 4950 15673 | 3088 | 0.082 | 0.6363 | No | ||
| 41 | ZNF385 | 22106 2297 | 3215 | 0.070 | 0.6312 | No | ||
| 42 | PACSIN3 | 14948 8149 2776 2665 | 3442 | 0.054 | 0.6202 | No | ||
| 43 | PPM1A | 5279 9607 | 3495 | 0.050 | 0.6186 | No | ||
| 44 | GRK5 | 9036 23814 | 3534 | 0.048 | 0.6177 | No | ||
| 45 | SLC22A17 | 21830 3096 | 3635 | 0.043 | 0.6133 | No | ||
| 46 | SOX5 | 9850 1044 5480 16937 | 3677 | 0.041 | 0.6120 | No | ||
| 47 | F11R | 9199 | 3703 | 0.040 | 0.6116 | No | ||
| 48 | MAN1C1 | 10514 6064 15714 | 3748 | 0.038 | 0.6101 | No | ||
| 49 | GAB2 | 1821 18184 2025 | 3992 | 0.029 | 0.5976 | No | ||
| 50 | P2RX4 | 3520 16711 | 4069 | 0.027 | 0.5942 | No | ||
| 51 | FGR | 4723 | 4382 | 0.019 | 0.5777 | No | ||
| 52 | CCND2 | 16987 | 4469 | 0.018 | 0.5735 | No | ||
| 53 | RAPSN | 2879 14951 | 4530 | 0.017 | 0.5706 | No | ||
| 54 | BTBD11 | 13147 | 4736 | 0.014 | 0.5599 | No | ||
| 55 | SCN3B | 39 3117 6163 10646 | 4872 | 0.013 | 0.5529 | No | ||
| 56 | SVIL | 1990 1942 10334 1976 23626 1946 1945 2041 | 4938 | 0.012 | 0.5496 | No | ||
| 57 | JMJD1C | 11531 19996 | 4997 | 0.012 | 0.5468 | No | ||
| 58 | AQP5 | 22365 | 5118 | 0.011 | 0.5405 | No | ||
| 59 | TRIM46 | 1779 15281 | 5130 | 0.011 | 0.5402 | No | ||
| 60 | LEPREL2 | 17001 | 5161 | 0.011 | 0.5388 | No | ||
| 61 | RXRG | 14063 | 5255 | 0.010 | 0.5340 | No | ||
| 62 | ELP4 | 8094 | 5258 | 0.010 | 0.5342 | No | ||
| 63 | STXBP6 | 21076 | 5470 | 0.009 | 0.5229 | No | ||
| 64 | TRPC4 | 15593 | 5540 | 0.008 | 0.5194 | No | ||
| 65 | NEURL | 5164 | 5997 | 0.006 | 0.4949 | No | ||
| 66 | FHL3 | 16092 2457 | 6248 | 0.005 | 0.4815 | No | ||
| 67 | ANXA8 | 22046 | 6539 | 0.004 | 0.4659 | No | ||
| 68 | MYF5 | 19632 | 6749 | 0.004 | 0.4546 | No | ||
| 69 | SPAG6 | 22650 | 6895 | 0.003 | 0.4469 | No | ||
| 70 | SYTL2 | 18190 1539 3730 | 6956 | 0.003 | 0.4437 | No | ||
| 71 | PITX3 | 9571 | 7039 | 0.003 | 0.4393 | No | ||
| 72 | HPCA | 9118 | 7231 | 0.002 | 0.4290 | No | ||
| 73 | GATA3 | 9004 | 7617 | 0.002 | 0.4082 | No | ||
| 74 | RNF121 | 7952 373 | 7618 | 0.001 | 0.4083 | No | ||
| 75 | DACT1 | 7203 | 7716 | 0.001 | 0.4031 | No | ||
| 76 | DCLRE1A | 23641 | 7805 | 0.001 | 0.3983 | No | ||
| 77 | PTGFRN | 15224 | 7823 | 0.001 | 0.3974 | No | ||
| 78 | CHAT | 21882 | 8042 | 0.000 | 0.3856 | No | ||
| 79 | LOH11CR2A | 19498 | 8194 | -0.000 | 0.3774 | No | ||
| 80 | PCF11 | 121 | 8517 | -0.001 | 0.3600 | No | ||
| 81 | KCNA2 | 15458 | 8727 | -0.001 | 0.3488 | No | ||
| 82 | CDKN1B | 4512 8730 | 8756 | -0.001 | 0.3473 | No | ||
| 83 | MCAM | 3109 19479 | 8833 | -0.002 | 0.3432 | No | ||
| 84 | TAS1R1 | 15657 | 8934 | -0.002 | 0.3378 | No | ||
| 85 | NHLH2 | 9462 | 9248 | -0.002 | 0.3209 | No | ||
| 86 | LRRTM3 | 19744 | 9277 | -0.003 | 0.3195 | No | ||
| 87 | RGR | 21874 3042 | 9434 | -0.003 | 0.3111 | No | ||
| 88 | KCNN3 | 15538 1913 | 9578 | -0.003 | 0.3034 | No | ||
| 89 | VAMP1 | 5846 1043 | 9719 | -0.004 | 0.2959 | No | ||
| 90 | SSX2IP | 13617 | 9938 | -0.004 | 0.2842 | No | ||
| 91 | TBX3 | 16723 | 10008 | -0.004 | 0.2806 | No | ||
| 92 | SCFD1 | 21278 2157 | 10149 | -0.005 | 0.2731 | No | ||
| 93 | B3GALT2 | 6498 | 10391 | -0.005 | 0.2602 | No | ||
| 94 | NXPH4 | 19602 | 10421 | -0.005 | 0.2588 | No | ||
| 95 | RIN1 | 10349 | 10547 | -0.006 | 0.2521 | No | ||
| 96 | KCNIP2 | 23655 3725 | 10752 | -0.006 | 0.2412 | No | ||
| 97 | AICDA | 17295 1087 | 10949 | -0.007 | 0.2308 | No | ||
| 98 | EGR2 | 8886 | 10977 | -0.007 | 0.2295 | No | ||
| 99 | GNB3 | 9027 | 11201 | -0.008 | 0.2176 | No | ||
| 100 | HFE2 | 1777 15495 | 11254 | -0.008 | 0.2150 | No | ||
| 101 | SLITRK5 | 21939 | 11387 | -0.009 | 0.2080 | No | ||
| 102 | ACTA1 | 18715 | 11541 | -0.009 | 0.2000 | No | ||
| 103 | BNIP3 | 17581 | 11561 | -0.009 | 0.1992 | No | ||
| 104 | CABP1 | 16412 | 11571 | -0.009 | 0.1989 | No | ||
| 105 | SYT6 | 13437 7042 12040 | 11583 | -0.009 | 0.1985 | No | ||
| 106 | BARX2 | 8649 | 11738 | -0.010 | 0.1904 | No | ||
| 107 | SGCG | 21798 | 11870 | -0.011 | 0.1836 | No | ||
| 108 | GPR37L1 | 13825 | 11939 | -0.011 | 0.1802 | No | ||
| 109 | RORC | 9732 | 12025 | -0.011 | 0.1758 | No | ||
| 110 | TNRC4 | 15514 | 12070 | -0.012 | 0.1737 | No | ||
| 111 | LOXL4 | 23672 | 12077 | -0.012 | 0.1737 | No | ||
| 112 | SLC18A3 | 21883 | 12315 | -0.013 | 0.1611 | No | ||
| 113 | DPP3 | 7954 3746 | 12631 | -0.015 | 0.1444 | No | ||
| 114 | CRB1 | 5031 | 12649 | -0.015 | 0.1439 | No | ||
| 115 | ACTN3 | 23967 8540 | 12724 | -0.015 | 0.1402 | No | ||
| 116 | MYCBP | 7093 | 12953 | -0.017 | 0.1283 | No | ||
| 117 | LRFN5 | 6213 | 13196 | -0.019 | 0.1156 | No | ||
| 118 | MYBPH | 14130 | 13415 | -0.022 | 0.1044 | No | ||
| 119 | GRIK3 | 16083 | 13472 | -0.022 | 0.1018 | No | ||
| 120 | GJA12 | 20433 1323 | 13609 | -0.024 | 0.0951 | No | ||
| 121 | NTF3 | 16991 | 13671 | -0.025 | 0.0923 | No | ||
| 122 | RUNX3 | 8702 | 13765 | -0.026 | 0.0879 | No | ||
| 123 | ART1 | 660 4414 | 13951 | -0.030 | 0.0786 | No | ||
| 124 | TNNT3 | 18004 | 14287 | -0.036 | 0.0613 | No | ||
| 125 | DAAM1 | 5536 | 14355 | -0.038 | 0.0586 | No | ||
| 126 | PCDH9 | 21736 | 14423 | -0.040 | 0.0559 | No | ||
| 127 | HMGA2 | 4857 9098 3341 3444 | 14519 | -0.043 | 0.0518 | No | ||
| 128 | IGFBP6 | 9161 | 14581 | -0.044 | 0.0495 | No | ||
| 129 | USF1 | 5832 10260 10261 | 14823 | -0.052 | 0.0377 | No | ||
| 130 | TMEM25 | 2992 19141 | 14841 | -0.053 | 0.0381 | No | ||
| 131 | MEF2D | 9379 | 15172 | -0.071 | 0.0219 | No | ||
| 132 | TRIT1 | 16098 | 15325 | -0.080 | 0.0156 | No | ||
| 133 | ABCB9 | 3576 3546 7096 | 15337 | -0.081 | 0.0169 | No | ||
| 134 | ZMYND17 | 21909 | 15954 | -0.150 | -0.0129 | No | ||
| 135 | NHLRC2 | 23828 | 16561 | -0.283 | -0.0390 | No | ||
| 136 | YTHDF2 | 5623 5624 | 16606 | -0.295 | -0.0344 | No | ||
| 137 | EIF4G2 | 1908 8892 | 16813 | -0.350 | -0.0373 | No | ||
| 138 | PRICKLE1 | 8379 | 17006 | -0.410 | -0.0380 | No | ||
| 139 | ZRANB1 | 6884 11702 | 17496 | -0.628 | -0.0497 | No | ||
| 140 | POLL | 23658 3688 | 17537 | -0.649 | -0.0365 | No | ||
| 141 | SUPT16H | 8541 | 17560 | -0.663 | -0.0220 | No | ||
| 142 | TNNI2 | 18005 | 17863 | -0.865 | -0.0179 | No | ||
| 143 | LMO4 | 15151 | 18004 | -0.982 | -0.0022 | No | ||
| 144 | TPCN1 | 16398 6383 10915 | 18324 | -1.494 | 0.0158 | No |