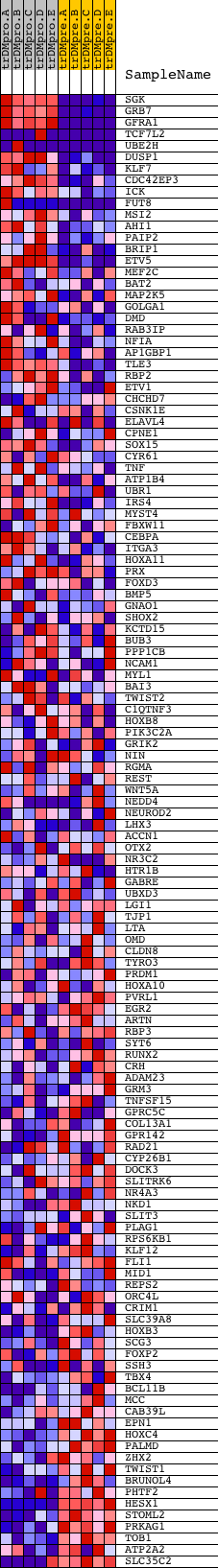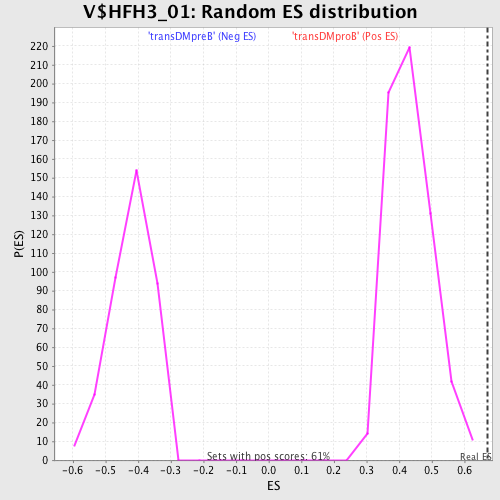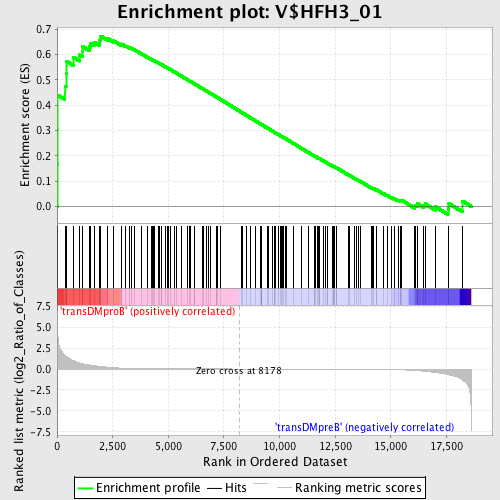
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_transDMpreB.phenotype_transDMproB_versus_transDMpreB.cls #transDMproB_versus_transDMpreB.phenotype_transDMproB_versus_transDMpreB.cls #transDMproB_versus_transDMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_transDMpreB.cls#transDMproB_versus_transDMpreB_repos |
| Upregulated in class | transDMproB |
| GeneSet | V$HFH3_01 |
| Enrichment Score (ES) | 0.6721751 |
| Normalized Enrichment Score (NES) | 1.5518423 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.06593963 |
| FWER p-Value | 0.508 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SGK | 9809 3419 | 12 | 4.998 | 0.1650 | Yes | ||
| 2 | GRB7 | 20673 | 25 | 4.169 | 0.3025 | Yes | ||
| 3 | GFRA1 | 9015 | 28 | 4.130 | 0.4393 | Yes | ||
| 4 | TCF7L2 | 10048 5646 | 354 | 1.605 | 0.4749 | Yes | ||
| 5 | UBE2H | 5823 5822 | 402 | 1.542 | 0.5235 | Yes | ||
| 6 | DUSP1 | 23061 | 416 | 1.523 | 0.5733 | Yes | ||
| 7 | KLF7 | 8205 | 734 | 0.981 | 0.5886 | Yes | ||
| 8 | CDC42EP3 | 6436 | 1007 | 0.722 | 0.5978 | Yes | ||
| 9 | ICK | 7135 12146 | 1150 | 0.618 | 0.6106 | Yes | ||
| 10 | FUT8 | 2163 21233 | 1155 | 0.617 | 0.6309 | Yes | ||
| 11 | MSI2 | 8030 1268 | 1451 | 0.482 | 0.6309 | Yes | ||
| 12 | AHI1 | 20077 | 1505 | 0.460 | 0.6433 | Yes | ||
| 13 | PAIP2 | 12593 | 1668 | 0.389 | 0.6474 | Yes | ||
| 14 | BRIP1 | 20311 | 1888 | 0.315 | 0.6460 | Yes | ||
| 15 | ETV5 | 22630 | 1903 | 0.314 | 0.6557 | Yes | ||
| 16 | MEF2C | 3204 9378 | 1945 | 0.302 | 0.6634 | Yes | ||
| 17 | BAT2 | 23006 | 1966 | 0.296 | 0.6722 | Yes | ||
| 18 | MAP2K5 | 19088 | 2276 | 0.211 | 0.6625 | No | ||
| 19 | GOLGA1 | 14595 | 2532 | 0.156 | 0.6538 | No | ||
| 20 | DMD | 24295 2647 | 2886 | 0.102 | 0.6381 | No | ||
| 21 | RAB3IP | 19623 | 2894 | 0.101 | 0.6411 | No | ||
| 22 | NFIA | 16172 5170 | 3093 | 0.082 | 0.6331 | No | ||
| 23 | AP1GBP1 | 20725 | 3259 | 0.066 | 0.6264 | No | ||
| 24 | TLE3 | 5767 19416 | 3343 | 0.060 | 0.6238 | No | ||
| 25 | RBP2 | 19347 | 3467 | 0.052 | 0.6189 | No | ||
| 26 | ETV1 | 4688 8925 8924 | 3792 | 0.036 | 0.6026 | No | ||
| 27 | CHCHD7 | 16280 | 4047 | 0.027 | 0.5898 | No | ||
| 28 | CSNK1E | 6570 2211 11332 | 4238 | 0.022 | 0.5802 | No | ||
| 29 | ELAVL4 | 15805 4889 9137 | 4280 | 0.021 | 0.5787 | No | ||
| 30 | CPNE1 | 11186 | 4352 | 0.020 | 0.5755 | No | ||
| 31 | SOX15 | 9848 | 4359 | 0.020 | 0.5758 | No | ||
| 32 | CYR61 | 9159 15145 9311 5024 | 4566 | 0.016 | 0.5652 | No | ||
| 33 | TNF | 23004 | 4599 | 0.016 | 0.5640 | No | ||
| 34 | ATP1B4 | 12586 | 4675 | 0.015 | 0.5605 | No | ||
| 35 | UBR1 | 5828 | 4693 | 0.015 | 0.5600 | No | ||
| 36 | IRS4 | 9183 4926 | 4849 | 0.013 | 0.5521 | No | ||
| 37 | MYST4 | 7027 7026 12014 2739 | 4971 | 0.012 | 0.5459 | No | ||
| 38 | FBXW11 | 20926 | 4989 | 0.012 | 0.5454 | No | ||
| 39 | CEBPA | 4514 | 5101 | 0.011 | 0.5398 | No | ||
| 40 | ITGA3 | 20284 | 5278 | 0.010 | 0.5306 | No | ||
| 41 | HOXA11 | 17146 | 5351 | 0.009 | 0.5270 | No | ||
| 42 | PRX | 3729 9628 | 5578 | 0.008 | 0.5150 | No | ||
| 43 | FOXD3 | 9088 | 5849 | 0.007 | 0.5007 | No | ||
| 44 | BMP5 | 19376 | 5941 | 0.007 | 0.4960 | No | ||
| 45 | GNAO1 | 4785 3829 | 6003 | 0.006 | 0.4929 | No | ||
| 46 | SHOX2 | 1802 5432 | 6163 | 0.006 | 0.4845 | No | ||
| 47 | KCTD15 | 17864 | 6178 | 0.006 | 0.4839 | No | ||
| 48 | BUB3 | 18045 | 6540 | 0.004 | 0.4645 | No | ||
| 49 | PPP1CB | 5282 3529 3504 | 6599 | 0.004 | 0.4615 | No | ||
| 50 | NCAM1 | 5149 | 6694 | 0.004 | 0.4566 | No | ||
| 51 | MYL1 | 4015 | 6797 | 0.004 | 0.4512 | No | ||
| 52 | BAI3 | 5580 | 6896 | 0.003 | 0.4460 | No | ||
| 53 | TWIST2 | 14185 | 7150 | 0.003 | 0.4324 | No | ||
| 54 | C1QTNF3 | 2224 22505 | 7174 | 0.003 | 0.4312 | No | ||
| 55 | HOXB8 | 9111 | 7206 | 0.003 | 0.4296 | No | ||
| 56 | PIK3C2A | 17667 | 7330 | 0.002 | 0.4231 | No | ||
| 57 | GRIK2 | 3318 | 8286 | -0.000 | 0.3714 | No | ||
| 58 | NIN | 9463 2103 | 8343 | -0.000 | 0.3684 | No | ||
| 59 | RGMA | 18213 | 8532 | -0.001 | 0.3583 | No | ||
| 60 | REST | 16818 | 8708 | -0.001 | 0.3488 | No | ||
| 61 | WNT5A | 22066 5880 | 8917 | -0.002 | 0.3376 | No | ||
| 62 | NEDD4 | 19384 3101 | 9119 | -0.002 | 0.3268 | No | ||
| 63 | NEUROD2 | 20262 | 9167 | -0.002 | 0.3244 | No | ||
| 64 | LHX3 | 2761 4996 | 9175 | -0.002 | 0.3241 | No | ||
| 65 | ACCN1 | 20327 1194 | 9191 | -0.002 | 0.3233 | No | ||
| 66 | OTX2 | 9519 | 9449 | -0.003 | 0.3095 | No | ||
| 67 | NR3C2 | 18562 13798 | 9488 | -0.003 | 0.3076 | No | ||
| 68 | HTR1B | 19051 | 9669 | -0.003 | 0.2979 | No | ||
| 69 | GABRE | 24140 | 9773 | -0.004 | 0.2925 | No | ||
| 70 | UBXD3 | 15701 | 9779 | -0.004 | 0.2924 | No | ||
| 71 | LGI1 | 23867 | 9798 | -0.004 | 0.2915 | No | ||
| 72 | TJP1 | 17803 | 9967 | -0.004 | 0.2826 | No | ||
| 73 | LTA | 23003 | 10055 | -0.004 | 0.2780 | No | ||
| 74 | OMD | 21643 | 10107 | -0.005 | 0.2754 | No | ||
| 75 | CLDN8 | 22549 1747 | 10152 | -0.005 | 0.2732 | No | ||
| 76 | TYRO3 | 5811 | 10170 | -0.005 | 0.2724 | No | ||
| 77 | PRDM1 | 19775 3337 | 10270 | -0.005 | 0.2672 | No | ||
| 78 | HOXA10 | 17145 989 | 10319 | -0.005 | 0.2648 | No | ||
| 79 | PVRL1 | 3006 19482 12211 7192 | 10622 | -0.006 | 0.2486 | No | ||
| 80 | EGR2 | 8886 | 10977 | -0.007 | 0.2297 | No | ||
| 81 | ARTN | 15783 2430 2532 | 11301 | -0.008 | 0.2125 | No | ||
| 82 | RBP3 | 22047 | 11559 | -0.009 | 0.1989 | No | ||
| 83 | SYT6 | 13437 7042 12040 | 11583 | -0.009 | 0.1980 | No | ||
| 84 | RUNX2 | 4480 8700 | 11607 | -0.009 | 0.1971 | No | ||
| 85 | CRH | 8784 | 11723 | -0.010 | 0.1912 | No | ||
| 86 | ADAM23 | 10699 4057 | 11755 | -0.010 | 0.1898 | No | ||
| 87 | GRM3 | 16929 | 11772 | -0.010 | 0.1893 | No | ||
| 88 | TNFSF15 | 15863 | 11807 | -0.010 | 0.1878 | No | ||
| 89 | GPRC5C | 12864 | 11964 | -0.011 | 0.1797 | No | ||
| 90 | COL13A1 | 8763 | 12057 | -0.011 | 0.1751 | No | ||
| 91 | GPR142 | 20604 | 12148 | -0.012 | 0.1706 | No | ||
| 92 | RAD21 | 22298 | 12173 | -0.012 | 0.1698 | No | ||
| 93 | CYP26B1 | 17093 | 12357 | -0.013 | 0.1603 | No | ||
| 94 | DOCK3 | 9920 5537 10751 | 12401 | -0.013 | 0.1584 | No | ||
| 95 | SLITRK6 | 21724 | 12420 | -0.013 | 0.1579 | No | ||
| 96 | NR4A3 | 9473 16212 5183 | 12466 | -0.014 | 0.1559 | No | ||
| 97 | NKD1 | 8228 | 12481 | -0.014 | 0.1556 | No | ||
| 98 | SLIT3 | 20925 | 12553 | -0.014 | 0.1522 | No | ||
| 99 | PLAG1 | 7148 15949 12161 | 12574 | -0.014 | 0.1516 | No | ||
| 100 | RPS6KB1 | 7815 1207 13040 | 13079 | -0.018 | 0.1250 | No | ||
| 101 | KLF12 | 4960 2637 21732 9228 | 13156 | -0.019 | 0.1215 | No | ||
| 102 | FLI1 | 4729 | 13349 | -0.021 | 0.1118 | No | ||
| 103 | MID1 | 5097 5098 | 13477 | -0.022 | 0.1056 | No | ||
| 104 | REPS2 | 24018 | 13554 | -0.023 | 0.1023 | No | ||
| 105 | ORC4L | 11172 6460 | 13622 | -0.024 | 0.0995 | No | ||
| 106 | CRIM1 | 403 | 14139 | -0.033 | 0.0727 | No | ||
| 107 | SLC39A8 | 12547 1776 | 14177 | -0.034 | 0.0718 | No | ||
| 108 | HOXB3 | 20685 | 14240 | -0.035 | 0.0696 | No | ||
| 109 | SCG3 | 19061 | 14367 | -0.038 | 0.0641 | No | ||
| 110 | FOXP2 | 4312 17523 8523 | 14370 | -0.039 | 0.0652 | No | ||
| 111 | SSH3 | 3709 10890 | 14689 | -0.048 | 0.0496 | No | ||
| 112 | TBX4 | 5633 | 14830 | -0.053 | 0.0438 | No | ||
| 113 | BCL11B | 7190 | 15046 | -0.064 | 0.0343 | No | ||
| 114 | MCC | 552 | 15167 | -0.071 | 0.0301 | No | ||
| 115 | CAB39L | 21992 | 15331 | -0.081 | 0.0240 | No | ||
| 116 | EPN1 | 18401 | 15413 | -0.087 | 0.0225 | No | ||
| 117 | HOXC4 | 4864 | 15454 | -0.090 | 0.0233 | No | ||
| 118 | PALMD | 15181 | 15496 | -0.094 | 0.0242 | No | ||
| 119 | ZHX2 | 11842 | 16078 | -0.171 | -0.0015 | No | ||
| 120 | TWIST1 | 21291 | 16092 | -0.175 | 0.0036 | No | ||
| 121 | BRUNOL4 | 2010 4229 | 16200 | -0.195 | 0.0042 | No | ||
| 122 | PHTF2 | 16599 | 16203 | -0.195 | 0.0106 | No | ||
| 123 | HESX1 | 22070 | 16470 | -0.259 | 0.0048 | No | ||
| 124 | STOML2 | 15902 | 16537 | -0.277 | 0.0104 | No | ||
| 125 | PRKAG1 | 22135 | 16997 | -0.407 | -0.0009 | No | ||
| 126 | TOB1 | 20703 | 17571 | -0.670 | -0.0097 | No | ||
| 127 | ATP2A2 | 4421 3481 | 17610 | -0.694 | 0.0113 | No | ||
| 128 | SLC35C2 | 10463 14339 | 18224 | -1.300 | 0.0212 | No |