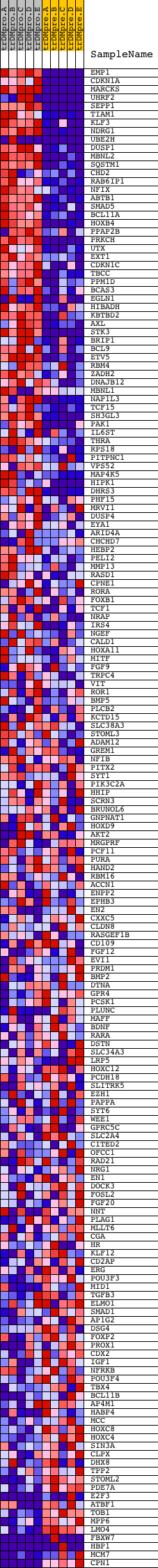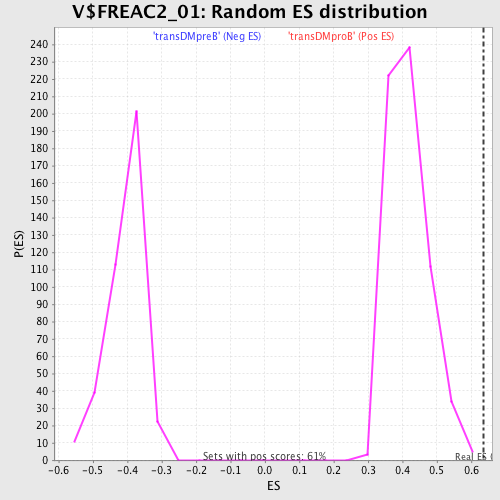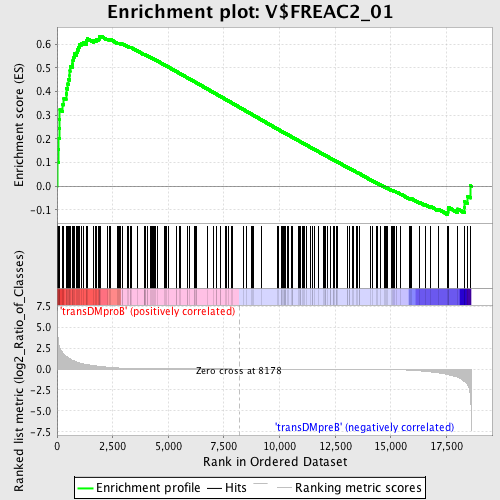
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_transDMpreB.phenotype_transDMproB_versus_transDMpreB.cls #transDMproB_versus_transDMpreB.phenotype_transDMproB_versus_transDMpreB.cls #transDMproB_versus_transDMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_transDMpreB.cls#transDMproB_versus_transDMpreB_repos |
| Upregulated in class | transDMproB |
| GeneSet | V$FREAC2_01 |
| Enrichment Score (ES) | 0.6348514 |
| Normalized Enrichment Score (NES) | 1.5254483 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.08136297 |
| FWER p-Value | 0.689 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | EMP1 | 17260 | 1 | 6.359 | 0.1015 | Yes | ||
| 2 | CDKN1A | 4511 8729 | 49 | 3.556 | 0.1557 | Yes | ||
| 3 | MARCKS | 9331 | 69 | 3.057 | 0.2035 | Yes | ||
| 4 | UHRF2 | 3731 4262 8452 23887 | 99 | 2.667 | 0.2446 | Yes | ||
| 5 | SEPP1 | 22524 | 116 | 2.535 | 0.2842 | Yes | ||
| 6 | TIAM1 | 22547 | 127 | 2.451 | 0.3228 | Yes | ||
| 7 | KLF3 | 4961 | 257 | 1.890 | 0.3460 | Yes | ||
| 8 | NDRG1 | 22276 | 305 | 1.727 | 0.3710 | Yes | ||
| 9 | UBE2H | 5823 5822 | 402 | 1.542 | 0.3904 | Yes | ||
| 10 | DUSP1 | 23061 | 416 | 1.523 | 0.4140 | Yes | ||
| 11 | MBNL2 | 21932 | 484 | 1.376 | 0.4324 | Yes | ||
| 12 | SQSTM1 | 9517 | 505 | 1.326 | 0.4525 | Yes | ||
| 13 | CHD2 | 10847 | 573 | 1.217 | 0.4683 | Yes | ||
| 14 | RAB6IP1 | 17676 | 577 | 1.204 | 0.4873 | Yes | ||
| 15 | NFIX | 9458 | 582 | 1.194 | 0.5062 | Yes | ||
| 16 | ABTB1 | 17075 | 689 | 1.036 | 0.5170 | Yes | ||
| 17 | SMAD5 | 21621 | 710 | 1.010 | 0.5320 | Yes | ||
| 18 | BCL11A | 4691 | 748 | 0.967 | 0.5455 | Yes | ||
| 19 | HOXB4 | 20686 | 767 | 0.952 | 0.5597 | Yes | ||
| 20 | PPAP2B | 7503 16161 | 891 | 0.836 | 0.5664 | Yes | ||
| 21 | PRKCH | 21246 | 909 | 0.818 | 0.5785 | Yes | ||
| 22 | UTX | 10266 2574 | 960 | 0.767 | 0.5880 | Yes | ||
| 23 | EXT1 | 22297 | 997 | 0.734 | 0.5978 | Yes | ||
| 24 | CDKN1C | 17546 | 1079 | 0.663 | 0.6040 | Yes | ||
| 25 | TBCC | 23215 | 1182 | 0.601 | 0.6081 | Yes | ||
| 26 | PPM1D | 20721 | 1315 | 0.540 | 0.6095 | Yes | ||
| 27 | BCAS3 | 5314 | 1330 | 0.533 | 0.6173 | Yes | ||
| 28 | EGLN1 | 8504 18707 4298 | 1367 | 0.516 | 0.6236 | Yes | ||
| 29 | HIBADH | 17144 | 1636 | 0.402 | 0.6155 | Yes | ||
| 30 | KBTBD2 | 17137 | 1731 | 0.368 | 0.6162 | Yes | ||
| 31 | AXL | 17922 | 1779 | 0.353 | 0.6193 | Yes | ||
| 32 | STK3 | 7084 | 1848 | 0.328 | 0.6209 | Yes | ||
| 33 | BRIP1 | 20311 | 1888 | 0.315 | 0.6238 | Yes | ||
| 34 | BCL9 | 15232 | 1891 | 0.315 | 0.6287 | Yes | ||
| 35 | ETV5 | 22630 | 1903 | 0.314 | 0.6331 | Yes | ||
| 36 | RBM4 | 9706 3693 | 1960 | 0.298 | 0.6349 | Yes | ||
| 37 | ZADH2 | 23501 | 2250 | 0.217 | 0.6226 | No | ||
| 38 | DNAJB12 | 7146 | 2333 | 0.196 | 0.6213 | No | ||
| 39 | MBNL1 | 1921 15582 | 2390 | 0.183 | 0.6212 | No | ||
| 40 | NAP1L3 | 24266 | 2732 | 0.125 | 0.6047 | No | ||
| 41 | TCF15 | 14798 | 2767 | 0.118 | 0.6047 | No | ||
| 42 | SH3GL3 | 18201 | 2788 | 0.114 | 0.6055 | No | ||
| 43 | PAK1 | 9527 | 2868 | 0.104 | 0.6028 | No | ||
| 44 | IL6ST | 4920 | 2935 | 0.096 | 0.6008 | No | ||
| 45 | THRA | 1447 10171 1406 | 2947 | 0.096 | 0.6017 | No | ||
| 46 | RPS18 | 1529 5397 1603 | 3143 | 0.077 | 0.5924 | No | ||
| 47 | PITPNC1 | 7759 | 3222 | 0.069 | 0.5893 | No | ||
| 48 | VPS52 | 23287 1622 | 3290 | 0.064 | 0.5866 | No | ||
| 49 | MAP4K5 | 11853 21048 2101 2142 | 3298 | 0.063 | 0.5873 | No | ||
| 50 | HIPK1 | 4851 | 3337 | 0.061 | 0.5862 | No | ||
| 51 | DHRS3 | 15997 | 3601 | 0.044 | 0.5726 | No | ||
| 52 | PHF15 | 20467 8050 | 3911 | 0.032 | 0.5564 | No | ||
| 53 | MRVI1 | 17674 1020 2526 | 3919 | 0.031 | 0.5565 | No | ||
| 54 | DUSP4 | 18632 3820 | 3938 | 0.031 | 0.5560 | No | ||
| 55 | EYA1 | 4695 4061 | 3955 | 0.030 | 0.5556 | No | ||
| 56 | ARID4A | 2080 6215 | 3964 | 0.030 | 0.5557 | No | ||
| 57 | CHCHD7 | 16280 | 4047 | 0.027 | 0.5517 | No | ||
| 58 | HEBP2 | 19812 | 4067 | 0.027 | 0.5511 | No | ||
| 59 | PELI2 | 8217 | 4180 | 0.024 | 0.5454 | No | ||
| 60 | MMP13 | 19573 3007 | 4253 | 0.022 | 0.5418 | No | ||
| 61 | RASD1 | 20424 | 4299 | 0.021 | 0.5397 | No | ||
| 62 | CPNE1 | 11186 | 4352 | 0.020 | 0.5372 | No | ||
| 63 | RORA | 3019 5388 | 4394 | 0.019 | 0.5353 | No | ||
| 64 | FOXB1 | 12254 19069 | 4427 | 0.018 | 0.5338 | No | ||
| 65 | TCF1 | 16416 | 4523 | 0.017 | 0.5290 | No | ||
| 66 | NRAP | 23642 | 4811 | 0.013 | 0.5136 | No | ||
| 67 | IRS4 | 9183 4926 | 4849 | 0.013 | 0.5118 | No | ||
| 68 | NGEF | 13891 | 4894 | 0.013 | 0.5096 | No | ||
| 69 | CALD1 | 4273 8463 | 5005 | 0.012 | 0.5038 | No | ||
| 70 | HOXA11 | 17146 | 5351 | 0.009 | 0.4853 | No | ||
| 71 | MITF | 17349 | 5365 | 0.009 | 0.4847 | No | ||
| 72 | FGF9 | 8966 | 5506 | 0.009 | 0.4773 | No | ||
| 73 | TRPC4 | 15593 | 5540 | 0.008 | 0.4756 | No | ||
| 74 | VIT | 23155 | 5867 | 0.007 | 0.4580 | No | ||
| 75 | ROR1 | 6472 | 5871 | 0.007 | 0.4580 | No | ||
| 76 | BMP5 | 19376 | 5941 | 0.007 | 0.4543 | No | ||
| 77 | PLCB2 | 5262 | 6175 | 0.006 | 0.4418 | No | ||
| 78 | KCTD15 | 17864 | 6178 | 0.006 | 0.4418 | No | ||
| 79 | SLC38A3 | 13319 | 6240 | 0.005 | 0.4386 | No | ||
| 80 | STOML3 | 15594 | 6264 | 0.005 | 0.4374 | No | ||
| 81 | ADAM12 | 2207 8545 | 6278 | 0.005 | 0.4368 | No | ||
| 82 | GREM1 | 14485 | 6777 | 0.004 | 0.4098 | No | ||
| 83 | NFIB | 15855 | 7045 | 0.003 | 0.3954 | No | ||
| 84 | PITX2 | 15424 1878 | 7147 | 0.003 | 0.3899 | No | ||
| 85 | SYT1 | 5565 | 7163 | 0.003 | 0.3892 | No | ||
| 86 | PIK3C2A | 17667 | 7330 | 0.002 | 0.3802 | No | ||
| 87 | HHIP | 4849 | 7361 | 0.002 | 0.3786 | No | ||
| 88 | SCRN3 | 14984 | 7554 | 0.002 | 0.3682 | No | ||
| 89 | BRUNOL6 | 19421 | 7622 | 0.001 | 0.3646 | No | ||
| 90 | GNPNAT1 | 12027 7031 | 7688 | 0.001 | 0.3611 | No | ||
| 91 | HOXD9 | 14982 | 7822 | 0.001 | 0.3539 | No | ||
| 92 | AKT2 | 4365 4366 | 7867 | 0.001 | 0.3515 | No | ||
| 93 | MRGPRF | 17995 | 8366 | -0.000 | 0.3245 | No | ||
| 94 | PCF11 | 121 | 8517 | -0.001 | 0.3164 | No | ||
| 95 | PURA | 9670 | 8758 | -0.001 | 0.3034 | No | ||
| 96 | HAND2 | 18615 | 8794 | -0.001 | 0.3015 | No | ||
| 97 | RBM16 | 11492 8387 | 8822 | -0.001 | 0.3001 | No | ||
| 98 | ACCN1 | 20327 1194 | 9191 | -0.002 | 0.2802 | No | ||
| 99 | ENPP2 | 9548 | 9926 | -0.004 | 0.2404 | No | ||
| 100 | EPHB3 | 22816 | 9969 | -0.004 | 0.2382 | No | ||
| 101 | EN2 | 16898 | 10072 | -0.004 | 0.2328 | No | ||
| 102 | CXXC5 | 12523 | 10097 | -0.004 | 0.2315 | No | ||
| 103 | CLDN8 | 22549 1747 | 10152 | -0.005 | 0.2287 | No | ||
| 104 | RASGEF1B | 11515 16469 | 10157 | -0.005 | 0.2285 | No | ||
| 105 | CD109 | 6174 10656 19369 | 10221 | -0.005 | 0.2252 | No | ||
| 106 | FGF12 | 1723 22621 | 10236 | -0.005 | 0.2245 | No | ||
| 107 | EVI1 | 4689 8926 | 10259 | -0.005 | 0.2234 | No | ||
| 108 | PRDM1 | 19775 3337 | 10270 | -0.005 | 0.2229 | No | ||
| 109 | BMP2 | 14833 | 10344 | -0.005 | 0.2191 | No | ||
| 110 | DTNA | 8870 4647 23611 1991 | 10399 | -0.005 | 0.2162 | No | ||
| 111 | GPR4 | 18364 | 10524 | -0.006 | 0.2096 | No | ||
| 112 | PCSK1 | 21600 | 10560 | -0.006 | 0.2078 | No | ||
| 113 | PLUNC | 9593 | 10576 | -0.006 | 0.2071 | No | ||
| 114 | MAFF | 22423 2243 | 10839 | -0.007 | 0.1929 | No | ||
| 115 | BDNF | 14926 2797 | 10903 | -0.007 | 0.1896 | No | ||
| 116 | RARA | 5358 | 10935 | -0.007 | 0.1881 | No | ||
| 117 | DSTN | 14823 | 10946 | -0.007 | 0.1876 | No | ||
| 118 | SLC34A3 | 14664 | 11050 | -0.007 | 0.1822 | No | ||
| 119 | LRP5 | 23948 9285 | 11079 | -0.007 | 0.1808 | No | ||
| 120 | HOXC12 | 22343 | 11134 | -0.008 | 0.1780 | No | ||
| 121 | PCDH18 | 15348 | 11231 | -0.008 | 0.1729 | No | ||
| 122 | SLITRK5 | 21939 | 11387 | -0.009 | 0.1646 | No | ||
| 123 | EZH1 | 20217 | 11467 | -0.009 | 0.1605 | No | ||
| 124 | PAPPA | 16190 | 11498 | -0.009 | 0.1590 | No | ||
| 125 | SYT6 | 13437 7042 12040 | 11583 | -0.009 | 0.1546 | No | ||
| 126 | WEE1 | 18127 | 11768 | -0.010 | 0.1448 | No | ||
| 127 | GPRC5C | 12864 | 11964 | -0.011 | 0.1344 | No | ||
| 128 | SLC2A4 | 20380 | 12021 | -0.011 | 0.1315 | No | ||
| 129 | CITED2 | 5118 14477 | 12067 | -0.011 | 0.1292 | No | ||
| 130 | OFCC1 | 21486 | 12078 | -0.012 | 0.1289 | No | ||
| 131 | RAD21 | 22298 | 12173 | -0.012 | 0.1240 | No | ||
| 132 | NRG1 | 3885 | 12271 | -0.013 | 0.1189 | No | ||
| 133 | EN1 | 387 14155 | 12307 | -0.013 | 0.1172 | No | ||
| 134 | DOCK3 | 9920 5537 10751 | 12401 | -0.013 | 0.1124 | No | ||
| 135 | FOSL2 | 4733 8978 16878 | 12417 | -0.013 | 0.1118 | No | ||
| 136 | FGF20 | 18892 | 12484 | -0.014 | 0.1084 | No | ||
| 137 | NNT | 3273 5181 9471 3238 | 12554 | -0.014 | 0.1049 | No | ||
| 138 | PLAG1 | 7148 15949 12161 | 12574 | -0.014 | 0.1041 | No | ||
| 139 | MLLT6 | 20678 | 12595 | -0.015 | 0.1033 | No | ||
| 140 | CGA | 16246 | 13058 | -0.018 | 0.0785 | No | ||
| 141 | HR | 3622 9119 | 13059 | -0.018 | 0.0788 | No | ||
| 142 | KLF12 | 4960 2637 21732 9228 | 13156 | -0.019 | 0.0739 | No | ||
| 143 | CD2AP | 22975 | 13164 | -0.019 | 0.0738 | No | ||
| 144 | ERG | 1686 8915 | 13281 | -0.020 | 0.0678 | No | ||
| 145 | POU3F3 | 14261 | 13339 | -0.021 | 0.0651 | No | ||
| 146 | MID1 | 5097 5098 | 13477 | -0.022 | 0.0580 | No | ||
| 147 | TGFB3 | 10161 | 13520 | -0.023 | 0.0561 | No | ||
| 148 | ELMO1 | 3275 8939 3202 3271 3254 3235 3164 3175 3274 3188 3293 3214 3218 3239 | 13585 | -0.024 | 0.0530 | No | ||
| 149 | SMAD1 | 18831 922 | 13589 | -0.024 | 0.0532 | No | ||
| 150 | AP1G2 | 2677 21827 | 14069 | -0.032 | 0.0277 | No | ||
| 151 | DSG4 | 9262 23616 | 14166 | -0.034 | 0.0231 | No | ||
| 152 | FOXP2 | 4312 17523 8523 | 14370 | -0.039 | 0.0127 | No | ||
| 153 | PROX1 | 9623 | 14412 | -0.040 | 0.0111 | No | ||
| 154 | CDX2 | 16289 | 14529 | -0.043 | 0.0055 | No | ||
| 155 | IGF1 | 3352 9156 3409 | 14702 | -0.048 | -0.0031 | No | ||
| 156 | NFRKB | 10642 | 14765 | -0.050 | -0.0056 | No | ||
| 157 | POU3F4 | 347 | 14812 | -0.052 | -0.0073 | No | ||
| 158 | TBX4 | 5633 | 14830 | -0.053 | -0.0074 | No | ||
| 159 | BCL11B | 7190 | 15046 | -0.064 | -0.0180 | No | ||
| 160 | AP4M1 | 3499 16656 | 15074 | -0.065 | -0.0185 | No | ||
| 161 | HABP4 | 12145 | 15136 | -0.069 | -0.0207 | No | ||
| 162 | MCC | 552 | 15167 | -0.071 | -0.0212 | No | ||
| 163 | HOXC8 | 4865 | 15260 | -0.077 | -0.0249 | No | ||
| 164 | HOXC4 | 4864 | 15454 | -0.090 | -0.0339 | No | ||
| 165 | SIN3A | 5442 | 15834 | -0.133 | -0.0524 | No | ||
| 166 | CLPX | 19403 | 15874 | -0.140 | -0.0523 | No | ||
| 167 | DHX8 | 20649 | 15940 | -0.148 | -0.0534 | No | ||
| 168 | TPP2 | 14257 | 16293 | -0.217 | -0.0691 | No | ||
| 169 | STOML2 | 15902 | 16537 | -0.277 | -0.0778 | No | ||
| 170 | PDE7A | 9544 1788 | 16789 | -0.343 | -0.0859 | No | ||
| 171 | E2F3 | 21500 8874 | 17131 | -0.456 | -0.0972 | No | ||
| 172 | ATBF1 | 916 8633 | 17549 | -0.655 | -0.1093 | No | ||
| 173 | TOB1 | 20703 | 17571 | -0.670 | -0.0998 | No | ||
| 174 | MPP6 | 17448 | 17589 | -0.682 | -0.0898 | No | ||
| 175 | LMO4 | 15151 | 18004 | -0.982 | -0.0966 | No | ||
| 176 | FBXW7 | 1805 11928 | 18311 | -1.475 | -0.0896 | No | ||
| 177 | HBP1 | 2048 2162 21086 | 18322 | -1.493 | -0.0663 | No | ||
| 178 | MCM7 | 9372 3568 | 18458 | -1.891 | -0.0434 | No | ||
| 179 | CPN1 | 23667 | 18582 | -3.251 | 0.0018 | No |