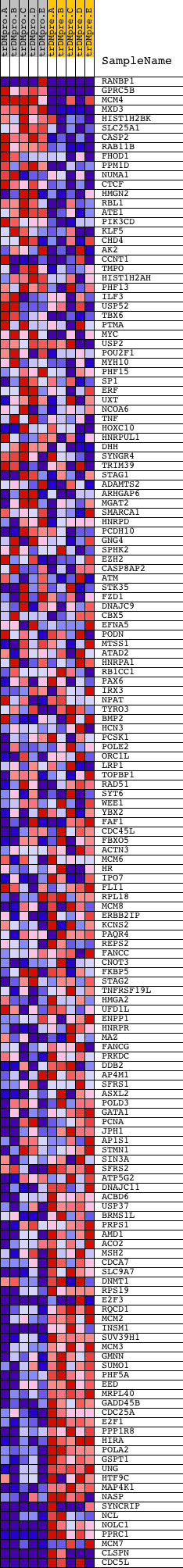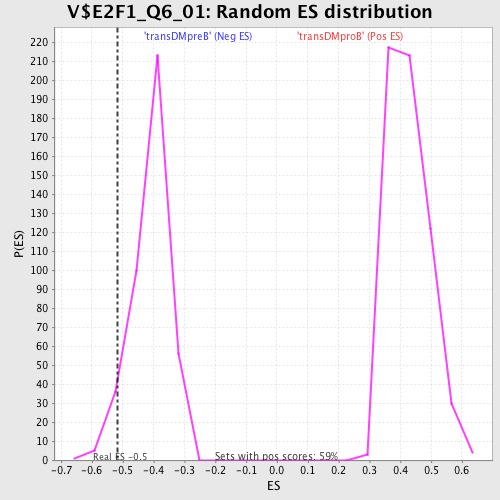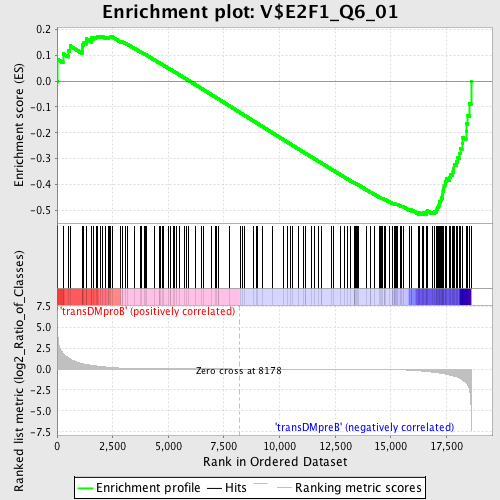
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_transDMpreB.phenotype_transDMproB_versus_transDMpreB.cls #transDMproB_versus_transDMpreB.phenotype_transDMproB_versus_transDMpreB.cls #transDMproB_versus_transDMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_transDMpreB.cls#transDMproB_versus_transDMpreB_repos |
| Upregulated in class | transDMpreB |
| GeneSet | V$E2F1_Q6_01 |
| Enrichment Score (ES) | -0.5157013 |
| Normalized Enrichment Score (NES) | -1.2614106 |
| Nominal p-value | 0.05596107 |
| FDR q-value | 0.72287697 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | RANBP1 | 9692 5357 | 14 | 4.554 | 0.0845 | No | ||
| 2 | GPRC5B | 17662 | 269 | 1.857 | 0.1055 | No | ||
| 3 | MCM4 | 22655 1708 | 497 | 1.345 | 0.1184 | No | ||
| 4 | MXD3 | 21455 | 579 | 1.199 | 0.1365 | No | ||
| 5 | HIST1H2BK | 11423 | 1126 | 0.632 | 0.1187 | No | ||
| 6 | SLC25A1 | 22645 | 1148 | 0.619 | 0.1292 | No | ||
| 7 | CASP2 | 8692 | 1152 | 0.617 | 0.1406 | No | ||
| 8 | RAB11B | 9676 | 1204 | 0.591 | 0.1489 | No | ||
| 9 | FHOD1 | 18772 | 1306 | 0.542 | 0.1536 | No | ||
| 10 | PPM1D | 20721 | 1315 | 0.540 | 0.1633 | No | ||
| 11 | NUMA1 | 18167 | 1544 | 0.441 | 0.1592 | No | ||
| 12 | CTCF | 18490 | 1551 | 0.438 | 0.1670 | No | ||
| 13 | HMGN2 | 9095 | 1653 | 0.396 | 0.1690 | No | ||
| 14 | RBL1 | 14373 | 1752 | 0.362 | 0.1704 | No | ||
| 15 | ATE1 | 388 17602 | 1805 | 0.341 | 0.1740 | No | ||
| 16 | PIK3CD | 9563 | 1930 | 0.306 | 0.1730 | No | ||
| 17 | KLF5 | 4456 | 2037 | 0.274 | 0.1724 | No | ||
| 18 | CHD4 | 8418 17281 4225 | 2169 | 0.239 | 0.1698 | No | ||
| 19 | AK2 | 8563 2479 16073 | 2297 | 0.206 | 0.1668 | No | ||
| 20 | CCNT1 | 22140 11607 | 2331 | 0.197 | 0.1687 | No | ||
| 21 | TMPO | 5778 | 2364 | 0.188 | 0.1704 | No | ||
| 22 | HIST1H2AH | 11414 | 2419 | 0.178 | 0.1709 | No | ||
| 23 | PHF13 | 15659 | 2467 | 0.167 | 0.1714 | No | ||
| 24 | ILF3 | 3110 3030 9176 | 2862 | 0.105 | 0.1521 | No | ||
| 25 | USP52 | 3397 19841 | 2928 | 0.097 | 0.1504 | No | ||
| 26 | TBX6 | 1993 18075 | 2940 | 0.096 | 0.1516 | No | ||
| 27 | PTMA | 9657 | 3085 | 0.082 | 0.1453 | No | ||
| 28 | MYC | 22465 9435 | 3151 | 0.076 | 0.1432 | No | ||
| 29 | USP2 | 19480 3052 3043 | 3457 | 0.052 | 0.1277 | No | ||
| 30 | POU2F1 | 5275 3989 4065 4010 | 3740 | 0.038 | 0.1131 | No | ||
| 31 | MYH10 | 8077 | 3813 | 0.036 | 0.1099 | No | ||
| 32 | PHF15 | 20467 8050 | 3911 | 0.032 | 0.1052 | No | ||
| 33 | SP1 | 9852 | 3958 | 0.030 | 0.1033 | No | ||
| 34 | ERF | 4680 8914 | 4006 | 0.029 | 0.1013 | No | ||
| 35 | UXT | 5837 10267 | 4363 | 0.020 | 0.0823 | No | ||
| 36 | NCOA6 | 2876 14382 | 4596 | 0.016 | 0.0701 | No | ||
| 37 | TNF | 23004 | 4599 | 0.016 | 0.0703 | No | ||
| 38 | HOXC10 | 22342 | 4625 | 0.015 | 0.0692 | No | ||
| 39 | HNRPUL1 | 6118 17923 1608 10577 1345 | 4740 | 0.014 | 0.0633 | No | ||
| 40 | DHH | 4629 | 4785 | 0.014 | 0.0612 | No | ||
| 41 | SYNGR4 | 17825 | 5015 | 0.012 | 0.0490 | No | ||
| 42 | TRIM39 | 22994 | 5082 | 0.011 | 0.0456 | No | ||
| 43 | STAG1 | 5520 9905 2987 | 5091 | 0.011 | 0.0454 | No | ||
| 44 | ADAMTS2 | 5706 20899 1356 | 5235 | 0.010 | 0.0378 | No | ||
| 45 | ARHGAP6 | 24207 | 5279 | 0.010 | 0.0357 | No | ||
| 46 | MGAT2 | 21256 | 5358 | 0.009 | 0.0316 | No | ||
| 47 | SMARCA1 | 24165 | 5509 | 0.009 | 0.0237 | No | ||
| 48 | HNRPD | 8645 | 5733 | 0.007 | 0.0117 | No | ||
| 49 | PCDH10 | 5227 9534 1893 | 5809 | 0.007 | 0.0078 | No | ||
| 50 | GNG4 | 21706 | 5887 | 0.007 | 0.0037 | No | ||
| 51 | SPHK2 | 1217 1806 12155 | 6203 | 0.006 | -0.0132 | No | ||
| 52 | EZH2 | 1092 17163 | 6481 | 0.005 | -0.0281 | No | ||
| 53 | CASP8AP2 | 16253 | 6558 | 0.004 | -0.0322 | No | ||
| 54 | ATM | 2976 19115 | 6932 | 0.003 | -0.0523 | No | ||
| 55 | STK35 | 14851 | 7102 | 0.003 | -0.0614 | No | ||
| 56 | FZD1 | 16923 | 7175 | 0.003 | -0.0653 | No | ||
| 57 | DNAJC9 | 21912 | 7236 | 0.002 | -0.0685 | No | ||
| 58 | CBX5 | 22108 4485 | 7271 | 0.002 | -0.0703 | No | ||
| 59 | EFNA5 | 4655 8884 | 7746 | 0.001 | -0.0959 | No | ||
| 60 | PODN | 15811 | 8257 | -0.000 | -0.1236 | No | ||
| 61 | MTSS1 | 5585 22285 | 8332 | -0.000 | -0.1276 | No | ||
| 62 | ATAD2 | 2268 2256 | 8408 | -0.001 | -0.1316 | No | ||
| 63 | HNRPA1 | 9103 4860 | 8807 | -0.001 | -0.1532 | No | ||
| 64 | RB1CC1 | 4486 14299 | 8977 | -0.002 | -0.1623 | No | ||
| 65 | PAX6 | 5223 | 8999 | -0.002 | -0.1634 | No | ||
| 66 | IRX3 | 18793 | 9249 | -0.002 | -0.1768 | No | ||
| 67 | NPAT | 19452 6356 | 9674 | -0.003 | -0.1997 | No | ||
| 68 | TYRO3 | 5811 | 10170 | -0.005 | -0.2265 | No | ||
| 69 | BMP2 | 14833 | 10344 | -0.005 | -0.2357 | No | ||
| 70 | HCN3 | 4843 1763 | 10495 | -0.006 | -0.2438 | No | ||
| 71 | PCSK1 | 21600 | 10560 | -0.006 | -0.2471 | No | ||
| 72 | POLE2 | 21053 | 10834 | -0.007 | -0.2618 | No | ||
| 73 | ORC1L | 327 16144 | 11070 | -0.007 | -0.2744 | No | ||
| 74 | LRP1 | 9284 | 11148 | -0.008 | -0.2784 | No | ||
| 75 | TOPBP1 | 10659 | 11169 | -0.008 | -0.2794 | No | ||
| 76 | RAD51 | 2897 14903 | 11430 | -0.009 | -0.2933 | No | ||
| 77 | SYT6 | 13437 7042 12040 | 11583 | -0.009 | -0.3013 | No | ||
| 78 | WEE1 | 18127 | 11768 | -0.010 | -0.3111 | No | ||
| 79 | YBX2 | 20818 | 11868 | -0.011 | -0.3163 | No | ||
| 80 | FAF1 | 2508 16137 8951 | 11881 | -0.011 | -0.3167 | No | ||
| 81 | CDC45L | 22642 1752 | 12341 | -0.013 | -0.3414 | No | ||
| 82 | FBXO5 | 19830 | 12414 | -0.013 | -0.3450 | No | ||
| 83 | ACTN3 | 23967 8540 | 12724 | -0.015 | -0.3615 | No | ||
| 84 | MCM6 | 4000 13845 4119 | 12906 | -0.017 | -0.3710 | No | ||
| 85 | HR | 3622 9119 | 13059 | -0.018 | -0.3789 | No | ||
| 86 | IPO7 | 6130 | 13206 | -0.019 | -0.3864 | No | ||
| 87 | FLI1 | 4729 | 13349 | -0.021 | -0.3937 | No | ||
| 88 | RPL18 | 450 5390 | 13376 | -0.021 | -0.3947 | No | ||
| 89 | MCM8 | 14834 | 13377 | -0.021 | -0.3943 | No | ||
| 90 | ERBB2IP | 3232 12234 | 13396 | -0.021 | -0.3949 | No | ||
| 91 | KCNS2 | 4949 2232 | 13468 | -0.022 | -0.3983 | No | ||
| 92 | PAQR4 | 23105 | 13521 | -0.023 | -0.4007 | No | ||
| 93 | REPS2 | 24018 | 13554 | -0.023 | -0.4020 | No | ||
| 94 | FANCC | 4712 | 13902 | -0.029 | -0.4203 | No | ||
| 95 | CNOT3 | 6112 10565 6113 | 14085 | -0.032 | -0.4296 | No | ||
| 96 | FKBP5 | 23051 | 14102 | -0.032 | -0.4298 | No | ||
| 97 | STAG2 | 5521 | 14273 | -0.036 | -0.4384 | No | ||
| 98 | TNFRSF19L | 11498 6727 6726 | 14481 | -0.041 | -0.4488 | No | ||
| 99 | HMGA2 | 4857 9098 3341 3444 | 14519 | -0.043 | -0.4500 | No | ||
| 100 | UFD1L | 22825 | 14588 | -0.045 | -0.4528 | No | ||
| 101 | ENPP1 | 19804 | 14605 | -0.045 | -0.4529 | No | ||
| 102 | HNRPR | 13196 7909 2500 | 14697 | -0.048 | -0.4569 | No | ||
| 103 | MAZ | 1327 17623 | 14733 | -0.049 | -0.4579 | No | ||
| 104 | FANCG | 15904 | 14774 | -0.050 | -0.4591 | No | ||
| 105 | PRKDC | 9617 22847 | 14938 | -0.057 | -0.4669 | No | ||
| 106 | DDB2 | 2721 2878 14525 2661 | 15066 | -0.065 | -0.4725 | No | ||
| 107 | AP4M1 | 3499 16656 | 15074 | -0.065 | -0.4717 | No | ||
| 108 | SFRS1 | 8492 | 15175 | -0.071 | -0.4758 | No | ||
| 109 | ASXL2 | 7957 | 15178 | -0.072 | -0.4745 | No | ||
| 110 | POLD3 | 17742 | 15222 | -0.074 | -0.4755 | No | ||
| 111 | GATA1 | 24196 | 15262 | -0.077 | -0.4761 | No | ||
| 112 | PCNA | 9535 | 15286 | -0.078 | -0.4759 | No | ||
| 113 | JPH1 | 13994 | 15412 | -0.087 | -0.4811 | No | ||
| 114 | AP1S1 | 3500 3453 16335 | 15484 | -0.093 | -0.4832 | No | ||
| 115 | STMN1 | 9261 | 15569 | -0.101 | -0.4858 | No | ||
| 116 | SIN3A | 5442 | 15834 | -0.133 | -0.4976 | No | ||
| 117 | SFRS2 | 9807 20136 | 15923 | -0.146 | -0.4997 | No | ||
| 118 | ATP5G2 | 12610 | 15943 | -0.148 | -0.4979 | No | ||
| 119 | DNAJC11 | 2486 6068 | 16244 | -0.205 | -0.5103 | No | ||
| 120 | ACBD6 | 14089 | 16283 | -0.214 | -0.5084 | No | ||
| 121 | USP37 | 13923 | 16419 | -0.248 | -0.5111 | Yes | ||
| 122 | BRMS1L | 21268 | 16460 | -0.256 | -0.5084 | Yes | ||
| 123 | PRPS1 | 24233 | 16592 | -0.291 | -0.5101 | Yes | ||
| 124 | AMD1 | 8583 | 16617 | -0.297 | -0.5058 | Yes | ||
| 125 | ACO2 | 8527 | 16630 | -0.302 | -0.5008 | Yes | ||
| 126 | MSH2 | 23138 | 16871 | -0.368 | -0.5069 | Yes | ||
| 127 | CDCA7 | 12437 | 16957 | -0.393 | -0.5042 | Yes | ||
| 128 | SLC9A7 | 24177 | 17037 | -0.424 | -0.5005 | Yes | ||
| 129 | DNMT1 | 19217 | 17074 | -0.435 | -0.4943 | Yes | ||
| 130 | RPS19 | 5398 | 17101 | -0.445 | -0.4874 | Yes | ||
| 131 | E2F3 | 21500 8874 | 17131 | -0.456 | -0.4804 | Yes | ||
| 132 | RQCD1 | 12205 | 17178 | -0.469 | -0.4741 | Yes | ||
| 133 | MCM2 | 17074 | 17185 | -0.472 | -0.4656 | Yes | ||
| 134 | INSM1 | 14815 | 17253 | -0.507 | -0.4598 | Yes | ||
| 135 | SUV39H1 | 24194 | 17262 | -0.510 | -0.4506 | Yes | ||
| 136 | MCM3 | 13991 | 17329 | -0.542 | -0.4441 | Yes | ||
| 137 | GMNN | 21513 | 17342 | -0.547 | -0.4345 | Yes | ||
| 138 | SUMO1 | 5826 3943 10247 | 17344 | -0.548 | -0.4243 | Yes | ||
| 139 | PHF5A | 22194 | 17362 | -0.559 | -0.4147 | Yes | ||
| 140 | EED | 17759 | 17389 | -0.572 | -0.4054 | Yes | ||
| 141 | MRPL40 | 22641 | 17436 | -0.591 | -0.3968 | Yes | ||
| 142 | GADD45B | 19935 | 17459 | -0.605 | -0.3867 | Yes | ||
| 143 | CDC25A | 8721 | 17488 | -0.624 | -0.3766 | Yes | ||
| 144 | E2F1 | 14384 | 17648 | -0.711 | -0.3719 | Yes | ||
| 145 | PPP1R8 | 15730 | 17687 | -0.735 | -0.3602 | Yes | ||
| 146 | HIRA | 4852 9090 | 17792 | -0.810 | -0.3506 | Yes | ||
| 147 | POLA2 | 23988 | 17824 | -0.833 | -0.3367 | Yes | ||
| 148 | GSPT1 | 9046 | 17839 | -0.845 | -0.3216 | Yes | ||
| 149 | UNG | 10257 16744 | 17940 | -0.920 | -0.3098 | Yes | ||
| 150 | HTF9C | 4886 1678 | 18012 | -0.992 | -0.2951 | Yes | ||
| 151 | MAP4K1 | 18313 | 18098 | -1.095 | -0.2792 | Yes | ||
| 152 | NASP | 2383 2387 6955 | 18123 | -1.135 | -0.2592 | Yes | ||
| 153 | SYNCRIP | 3078 3035 3107 | 18203 | -1.259 | -0.2399 | Yes | ||
| 154 | NCL | 5153 13899 | 18230 | -1.310 | -0.2168 | Yes | ||
| 155 | NOLC1 | 7704 | 18381 | -1.612 | -0.1948 | Yes | ||
| 156 | PPRC1 | 23808 | 18408 | -1.703 | -0.1643 | Yes | ||
| 157 | MCM7 | 9372 3568 | 18458 | -1.891 | -0.1315 | Yes | ||
| 158 | CLSPN | 16079 11256 6530 | 18543 | -2.572 | -0.0879 | Yes | ||
| 159 | CDC5L | 7746 7745 7744 | 18609 | -4.902 | 0.0004 | Yes |