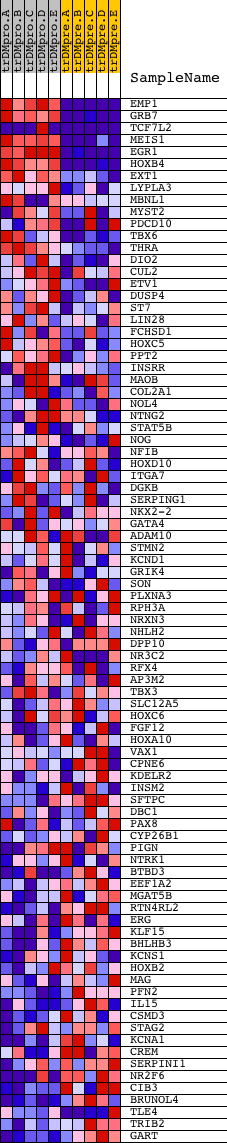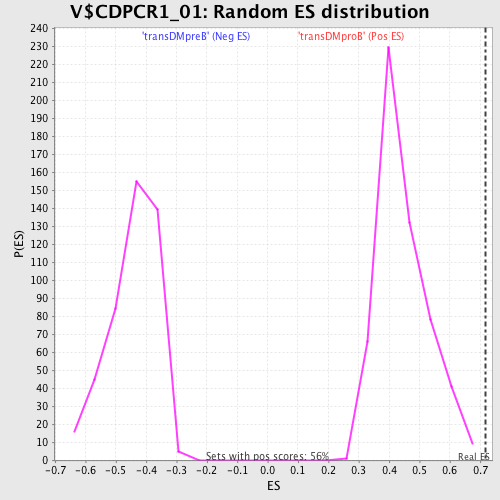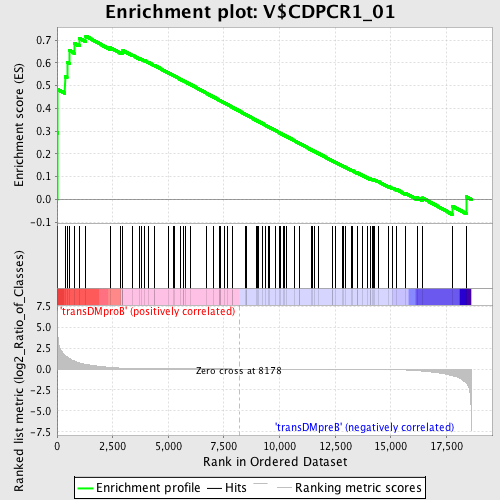
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_transDMpreB.phenotype_transDMproB_versus_transDMpreB.cls #transDMproB_versus_transDMpreB.phenotype_transDMproB_versus_transDMpreB.cls #transDMproB_versus_transDMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_transDMpreB.cls#transDMproB_versus_transDMpreB_repos |
| Upregulated in class | transDMproB |
| GeneSet | V$CDPCR1_01 |
| Enrichment Score (ES) | 0.7179821 |
| Normalized Enrichment Score (NES) | 1.607379 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.034103 |
| FWER p-Value | 0.204 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | EMP1 | 17260 | 1 | 6.359 | 0.2929 | Yes | ||
| 2 | GRB7 | 20673 | 25 | 4.169 | 0.4837 | Yes | ||
| 3 | TCF7L2 | 10048 5646 | 354 | 1.605 | 0.5400 | Yes | ||
| 4 | MEIS1 | 20524 | 446 | 1.464 | 0.6025 | Yes | ||
| 5 | EGR1 | 23598 | 542 | 1.256 | 0.6553 | Yes | ||
| 6 | HOXB4 | 20686 | 767 | 0.952 | 0.6870 | Yes | ||
| 7 | EXT1 | 22297 | 997 | 0.734 | 0.7085 | Yes | ||
| 8 | LYPLA3 | 18482 | 1290 | 0.548 | 0.7180 | Yes | ||
| 9 | MBNL1 | 1921 15582 | 2390 | 0.183 | 0.6671 | No | ||
| 10 | MYST2 | 20283 | 2842 | 0.108 | 0.6477 | No | ||
| 11 | PDCD10 | 15314 | 2939 | 0.096 | 0.6469 | No | ||
| 12 | TBX6 | 1993 18075 | 2940 | 0.096 | 0.6514 | No | ||
| 13 | THRA | 1447 10171 1406 | 2947 | 0.096 | 0.6555 | No | ||
| 14 | DIO2 | 21014 2159 | 3384 | 0.057 | 0.6345 | No | ||
| 15 | CUL2 | 23630 | 3714 | 0.039 | 0.6186 | No | ||
| 16 | ETV1 | 4688 8925 8924 | 3792 | 0.036 | 0.6161 | No | ||
| 17 | DUSP4 | 18632 3820 | 3938 | 0.031 | 0.6097 | No | ||
| 18 | ST7 | 1054 17519 | 4090 | 0.026 | 0.6028 | No | ||
| 19 | LIN28 | 15723 | 4361 | 0.020 | 0.5891 | No | ||
| 20 | FCHSD1 | 6661 11435 | 4368 | 0.019 | 0.5897 | No | ||
| 21 | HOXC5 | 22340 | 5008 | 0.012 | 0.5557 | No | ||
| 22 | PPT2 | 1617 12034 | 5226 | 0.010 | 0.5445 | No | ||
| 23 | INSRR | 15557 | 5273 | 0.010 | 0.5425 | No | ||
| 24 | MAOB | 24182 2556 | 5558 | 0.008 | 0.5275 | No | ||
| 25 | COL2A1 | 22145 4542 | 5678 | 0.008 | 0.5215 | No | ||
| 26 | NOL4 | 2028 11432 | 5695 | 0.008 | 0.5209 | No | ||
| 27 | NTNG2 | 2731 9330 | 5782 | 0.007 | 0.5166 | No | ||
| 28 | STAT5B | 20222 | 5994 | 0.006 | 0.5055 | No | ||
| 29 | NOG | 13663 | 6721 | 0.004 | 0.4665 | No | ||
| 30 | NFIB | 15855 | 7045 | 0.003 | 0.4492 | No | ||
| 31 | HOXD10 | 14983 | 7290 | 0.002 | 0.4362 | No | ||
| 32 | ITGA7 | 19835 | 7306 | 0.002 | 0.4355 | No | ||
| 33 | DGKB | 5722 | 7326 | 0.002 | 0.4345 | No | ||
| 34 | SERPING1 | 14546 | 7359 | 0.002 | 0.4329 | No | ||
| 35 | NKX2-2 | 5176 | 7544 | 0.002 | 0.4231 | No | ||
| 36 | GATA4 | 21792 4755 | 7652 | 0.001 | 0.4174 | No | ||
| 37 | ADAM10 | 4336 | 7875 | 0.001 | 0.4054 | No | ||
| 38 | STMN2 | 15638 | 7898 | 0.001 | 0.4043 | No | ||
| 39 | KCND1 | 4940 | 8477 | -0.001 | 0.3731 | No | ||
| 40 | GRIK4 | 19157 | 8497 | -0.001 | 0.3721 | No | ||
| 41 | SON | 5473 1657 1684 | 8508 | -0.001 | 0.3716 | No | ||
| 42 | PLXNA3 | 24299 | 8946 | -0.002 | 0.3481 | No | ||
| 43 | RPH3A | 13628 | 9008 | -0.002 | 0.3449 | No | ||
| 44 | NRXN3 | 2153 5196 | 9052 | -0.002 | 0.3427 | No | ||
| 45 | NHLH2 | 9462 | 9248 | -0.002 | 0.3323 | No | ||
| 46 | DPP10 | 13850 | 9374 | -0.003 | 0.3256 | No | ||
| 47 | NR3C2 | 18562 13798 | 9488 | -0.003 | 0.3197 | No | ||
| 48 | RFX4 | 7721 3410 | 9565 | -0.003 | 0.3157 | No | ||
| 49 | AP3M2 | 11 | 9822 | -0.004 | 0.3021 | No | ||
| 50 | TBX3 | 16723 | 10008 | -0.004 | 0.2923 | No | ||
| 51 | SLC12A5 | 14731 | 10060 | -0.004 | 0.2897 | No | ||
| 52 | HOXC6 | 22339 2302 | 10191 | -0.005 | 0.2829 | No | ||
| 53 | FGF12 | 1723 22621 | 10236 | -0.005 | 0.2808 | No | ||
| 54 | HOXA10 | 17145 989 | 10319 | -0.005 | 0.2766 | No | ||
| 55 | VAX1 | 23636 | 10658 | -0.006 | 0.2586 | No | ||
| 56 | CPNE6 | 22010 | 10878 | -0.007 | 0.2471 | No | ||
| 57 | KDELR2 | 16634 12428 | 10885 | -0.007 | 0.2471 | No | ||
| 58 | INSM2 | 19821 | 11443 | -0.009 | 0.2175 | No | ||
| 59 | SFTPC | 21759 | 11484 | -0.009 | 0.2157 | No | ||
| 60 | DBC1 | 7147 | 11582 | -0.009 | 0.2109 | No | ||
| 61 | PAX8 | 2791 14671 | 11751 | -0.010 | 0.2023 | No | ||
| 62 | CYP26B1 | 17093 | 12357 | -0.013 | 0.1702 | No | ||
| 63 | PIGN | 13865 | 12396 | -0.013 | 0.1688 | No | ||
| 64 | NTRK1 | 15299 | 12524 | -0.014 | 0.1626 | No | ||
| 65 | BTBD3 | 10457 2975 6007 6006 | 12821 | -0.016 | 0.1474 | No | ||
| 66 | EEF1A2 | 8880 14309 | 12854 | -0.016 | 0.1464 | No | ||
| 67 | MGAT5B | 20584 11197 | 12941 | -0.017 | 0.1425 | No | ||
| 68 | RTN4RL2 | 14544 | 13216 | -0.019 | 0.1286 | No | ||
| 69 | ERG | 1686 8915 | 13281 | -0.020 | 0.1261 | No | ||
| 70 | KLF15 | 17359 | 13488 | -0.022 | 0.1160 | No | ||
| 71 | BHLHB3 | 16935 | 13490 | -0.022 | 0.1170 | No | ||
| 72 | KCNS1 | 14358 | 13715 | -0.026 | 0.1061 | No | ||
| 73 | HOXB2 | 8346 | 13938 | -0.029 | 0.0955 | No | ||
| 74 | MAG | 17880 | 14081 | -0.032 | 0.0893 | No | ||
| 75 | PFN2 | 15337 1818 | 14090 | -0.032 | 0.0903 | No | ||
| 76 | IL15 | 18826 3801 | 14186 | -0.034 | 0.0868 | No | ||
| 77 | CSMD3 | 10735 7938 | 14203 | -0.034 | 0.0875 | No | ||
| 78 | STAG2 | 5521 | 14273 | -0.036 | 0.0854 | No | ||
| 79 | KCNA1 | 9202 | 14430 | -0.040 | 0.0789 | No | ||
| 80 | CREM | 2005 1965 23496 1951 | 14902 | -0.056 | 0.0560 | No | ||
| 81 | SERPINI1 | 340 | 15086 | -0.066 | 0.0492 | No | ||
| 82 | NR2F6 | 4651 8876 18848 8912 | 15272 | -0.078 | 0.0428 | No | ||
| 83 | CIB3 | 18840 | 15658 | -0.112 | 0.0272 | No | ||
| 84 | BRUNOL4 | 2010 4229 | 16200 | -0.195 | 0.0069 | No | ||
| 85 | TLE4 | 23720 3697 | 16421 | -0.248 | 0.0065 | No | ||
| 86 | TRIB2 | 21102 | 17771 | -0.796 | -0.0296 | No | ||
| 87 | GART | 22543 1754 | 18389 | -1.632 | 0.0123 | No |