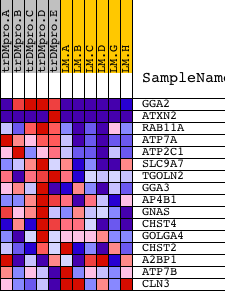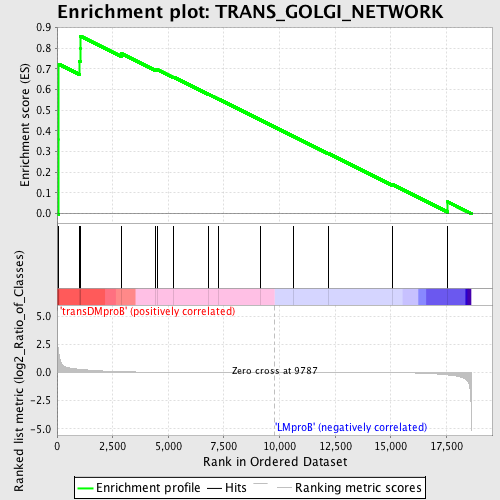
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_LMproB.phenotype_transDMproB_versus_LMproB.cls #transDMproB_versus_LMproB |
| Phenotype | phenotype_transDMproB_versus_LMproB.cls#transDMproB_versus_LMproB |
| Upregulated in class | transDMproB |
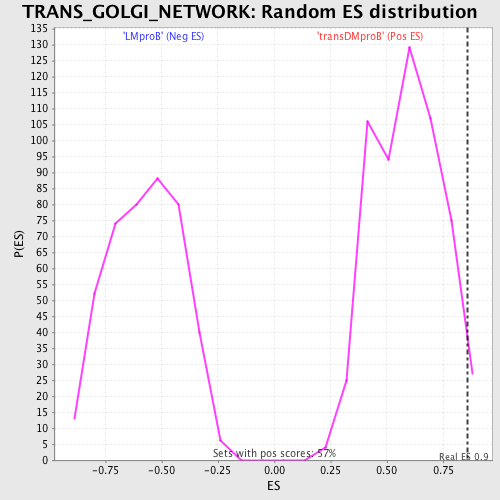
| GeneSet | TRANS_GOLGI_NETWORK |
| Enrichment Score (ES) | 0.85810435 |
| Normalized Enrichment Score (NES) | 1.452363 |
| Nominal p-value | 0.02292769 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | GGA2 | 1164 17652 13163 | 75 | 1.585 | 0.3599 | Yes | ||
| 2 | ATXN2 | 9780 3506 | 77 | 1.578 | 0.7222 | Yes | ||
| 3 | RAB11A | 7013 | 1003 | 0.277 | 0.7362 | Yes | ||
| 4 | ATP7A | 4425 | 1037 | 0.271 | 0.7966 | Yes | ||
| 5 | ATP2C1 | 10660 6178 | 1043 | 0.269 | 0.8581 | Yes | ||
| 6 | SLC9A7 | 24177 | 2892 | 0.063 | 0.7733 | No | ||
| 7 | TGOLN2 | 10230 | 4409 | 0.021 | 0.6966 | No | ||
| 8 | GGA3 | 1206 6435 | 4511 | 0.020 | 0.6957 | No | ||
| 9 | AP4B1 | 4395 | 5244 | 0.013 | 0.6594 | No | ||
| 10 | GNAS | 9025 2963 2752 | 6792 | 0.007 | 0.5778 | No | ||
| 11 | CHST4 | 18749 | 7275 | 0.005 | 0.5531 | No | ||
| 12 | GOLGA4 | 12022 | 9134 | 0.001 | 0.4535 | No | ||
| 13 | CHST2 | 7032 | 10646 | -0.002 | 0.3726 | No | ||
| 14 | A2BP1 | 11212 1652 1689 | 12186 | -0.005 | 0.2910 | No | ||
| 15 | ATP7B | 18918 4426 4427 8641 | 15054 | -0.020 | 0.1415 | No | ||
| 16 | CLN3 | 17635 | 17556 | -0.217 | 0.0570 | No |