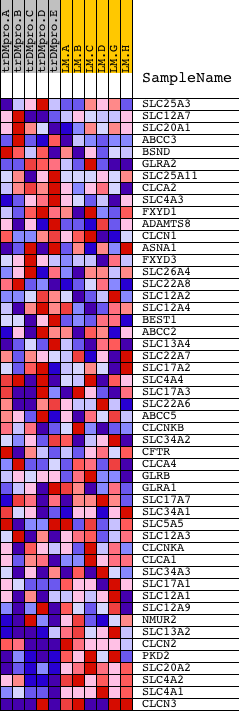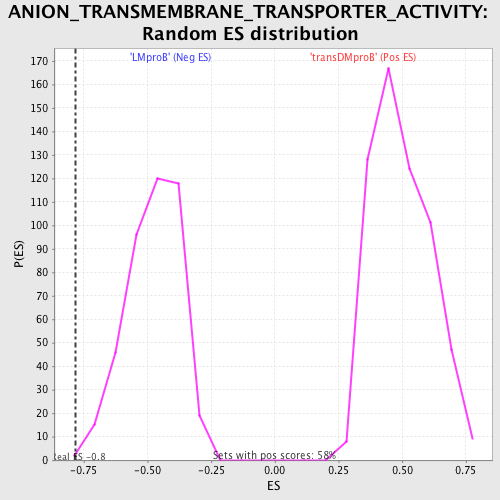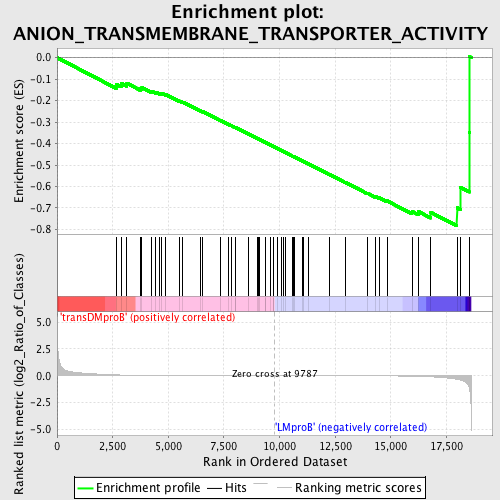
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_LMproB.phenotype_transDMproB_versus_LMproB.cls #transDMproB_versus_LMproB |
| Phenotype | phenotype_transDMproB_versus_LMproB.cls#transDMproB_versus_LMproB |
| Upregulated in class | LMproB |
| GeneSet | ANION_TRANSMEMBRANE_TRANSPORTER_ACTIVITY |
| Enrichment Score (ES) | -0.7838449 |
| Normalized Enrichment Score (NES) | -1.6456748 |
| Nominal p-value | 0.0024038462 |
| FDR q-value | 0.42028695 |
| FWER p-Value | 0.575 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SLC25A3 | 19644 | 2673 | 0.076 | -0.1244 | No | ||
| 2 | SLC12A7 | 21603 | 2882 | 0.064 | -0.1191 | No | ||
| 3 | SLC20A1 | 14855 | 3133 | 0.052 | -0.1191 | No | ||
| 4 | ABCC3 | 20293 | 3740 | 0.033 | -0.1432 | No | ||
| 5 | BSND | 15821 | 3809 | 0.032 | -0.1387 | No | ||
| 6 | GLRA2 | 24008 | 4241 | 0.023 | -0.1560 | No | ||
| 7 | SLC25A11 | 20366 1368 | 4438 | 0.020 | -0.1613 | No | ||
| 8 | CLCA2 | 15148 | 4611 | 0.019 | -0.1657 | No | ||
| 9 | SLC4A3 | 5454 14213 9829 | 4686 | 0.018 | -0.1651 | No | ||
| 10 | FXYD1 | 17874 85 | 4888 | 0.016 | -0.1718 | No | ||
| 11 | ADAMTS8 | 6626 | 5504 | 0.012 | -0.2018 | No | ||
| 12 | CLCN1 | 8744 | 5648 | 0.011 | -0.2067 | No | ||
| 13 | ASNA1 | 18810 | 6426 | 0.008 | -0.2465 | No | ||
| 14 | FXYD3 | 9370 | 6554 | 0.007 | -0.2514 | No | ||
| 15 | SLC26A4 | 21089 | 7326 | 0.005 | -0.2917 | No | ||
| 16 | SLC22A8 | 23944 | 7685 | 0.004 | -0.3099 | No | ||
| 17 | SLC12A2 | 23547 2026 | 7833 | 0.004 | -0.3168 | No | ||
| 18 | SLC12A4 | 18763 | 7996 | 0.003 | -0.3246 | No | ||
| 19 | BEST1 | 23747 | 8024 | 0.003 | -0.3252 | No | ||
| 20 | ABCC2 | 8755 23844 | 8587 | 0.002 | -0.3549 | No | ||
| 21 | SLC13A4 | 17185 | 8991 | 0.002 | -0.3762 | No | ||
| 22 | SLC22A7 | 22964 | 9044 | 0.001 | -0.3786 | No | ||
| 23 | SLC17A2 | 10163 5744 | 9092 | 0.001 | -0.3808 | No | ||
| 24 | SLC4A4 | 16807 12035 7036 | 9363 | 0.001 | -0.3952 | No | ||
| 25 | SLC17A3 | 3165 21694 | 9376 | 0.001 | -0.3956 | No | ||
| 26 | SLC22A6 | 5213 | 9584 | 0.000 | -0.4067 | No | ||
| 27 | ABCC5 | 22639 351 | 9727 | 0.000 | -0.4143 | No | ||
| 28 | CLCNKB | 7105 | 9730 | 0.000 | -0.4143 | No | ||
| 29 | SLC34A2 | 16845 | 9923 | -0.000 | -0.4246 | No | ||
| 30 | CFTR | 17518 | 10072 | -0.001 | -0.4324 | No | ||
| 31 | CLCA4 | 6041 | 10178 | -0.001 | -0.4379 | No | ||
| 32 | GLRB | 15312 1846 | 10280 | -0.001 | -0.4431 | No | ||
| 33 | GLRA1 | 9021 | 10565 | -0.001 | -0.4580 | No | ||
| 34 | SLC17A7 | 13081 | 10619 | -0.002 | -0.4605 | No | ||
| 35 | SLC34A1 | 9824 | 10653 | -0.002 | -0.4618 | No | ||
| 36 | SLC5A5 | 4322 | 11034 | -0.002 | -0.4817 | No | ||
| 37 | SLC12A3 | 3799 5451 | 11089 | -0.002 | -0.4840 | No | ||
| 38 | CLCNKA | 15690 | 11299 | -0.003 | -0.4945 | No | ||
| 39 | CLCA1 | 15149 | 12256 | -0.005 | -0.5447 | No | ||
| 40 | SLC34A3 | 14664 | 12945 | -0.007 | -0.5800 | No | ||
| 41 | SLC17A1 | 21693 | 13948 | -0.011 | -0.6311 | No | ||
| 42 | SLC12A1 | 14874 2772 2943 | 14290 | -0.013 | -0.6462 | No | ||
| 43 | SLC12A9 | 16332 3654 | 14481 | -0.014 | -0.6527 | No | ||
| 44 | NMUR2 | 20443 | 14828 | -0.017 | -0.6669 | No | ||
| 45 | SLC13A2 | 20336 | 15952 | -0.044 | -0.7162 | No | ||
| 46 | CLCN2 | 22637 | 16235 | -0.058 | -0.7166 | No | ||
| 47 | PKD2 | 16772 | 16797 | -0.102 | -0.7205 | Yes | ||
| 48 | SLC20A2 | 18654 | 17974 | -0.331 | -0.6988 | Yes | ||
| 49 | SLC4A2 | 9828 | 18130 | -0.398 | -0.6048 | Yes | ||
| 50 | SLC4A1 | 20204 | 18519 | -1.080 | -0.3481 | Yes | ||
| 51 | CLCN3 | 8745 3782 3893 4524 | 18547 | -1.374 | 0.0037 | Yes |