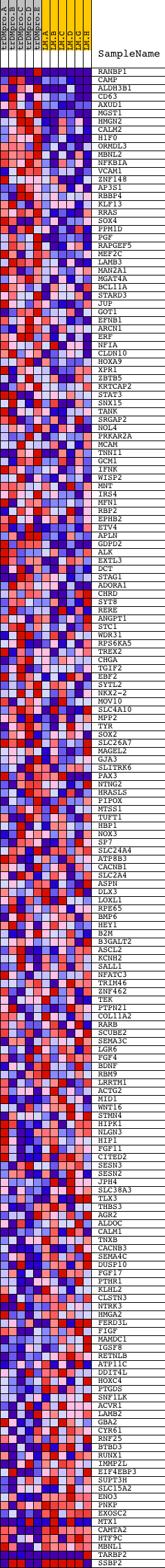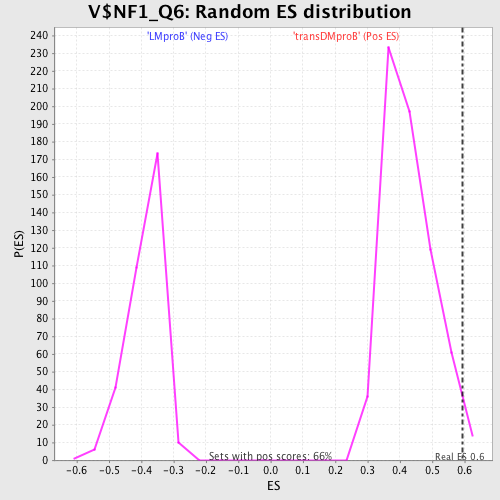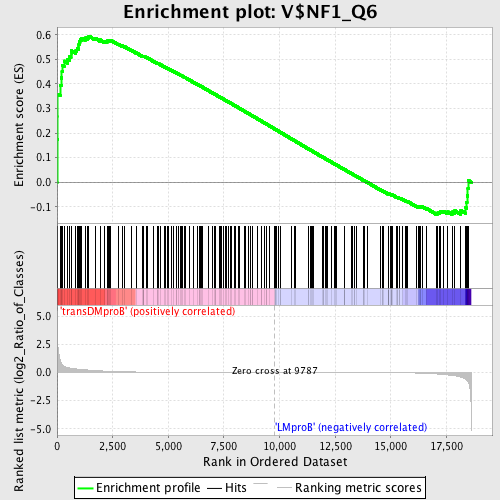
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_LMproB.phenotype_transDMproB_versus_LMproB.cls #transDMproB_versus_LMproB |
| Phenotype | phenotype_transDMproB_versus_LMproB.cls#transDMproB_versus_LMproB |
| Upregulated in class | transDMproB |
| GeneSet | V$NF1_Q6 |
| Enrichment Score (ES) | 0.59481627 |
| Normalized Enrichment Score (NES) | 1.3931361 |
| Nominal p-value | 0.018181818 |
| FDR q-value | 0.6059103 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | RANBP1 | 9692 5357 | 3 | 4.557 | 0.1756 | Yes | ||
| 2 | CAMP | 18990 | 22 | 2.405 | 0.2673 | Yes | ||
| 3 | ALDH3B1 | 12569 23949 | 24 | 2.342 | 0.3576 | Yes | ||
| 4 | CD63 | 8716 | 137 | 1.127 | 0.3950 | Yes | ||
| 5 | AXUD1 | 18967 | 187 | 0.847 | 0.4250 | Yes | ||
| 6 | MGST1 | 17254 | 218 | 0.726 | 0.4513 | Yes | ||
| 7 | HMGN2 | 9095 | 238 | 0.678 | 0.4765 | Yes | ||
| 8 | CALM2 | 8681 | 316 | 0.552 | 0.4936 | Yes | ||
| 9 | H1F0 | 22428 | 478 | 0.426 | 0.5013 | Yes | ||
| 10 | ORMDL3 | 12386 | 568 | 0.384 | 0.5113 | Yes | ||
| 11 | MBNL2 | 21932 | 626 | 0.363 | 0.5222 | Yes | ||
| 12 | NFKBIA | 21065 | 629 | 0.362 | 0.5360 | Yes | ||
| 13 | VCAM1 | 5851 | 846 | 0.306 | 0.5361 | Yes | ||
| 14 | ZNF148 | 5949 | 893 | 0.297 | 0.5450 | Yes | ||
| 15 | AP3S1 | 8604 | 976 | 0.283 | 0.5515 | Yes | ||
| 16 | RBBP4 | 5364 | 978 | 0.283 | 0.5623 | Yes | ||
| 17 | KLF13 | 17807 | 992 | 0.279 | 0.5724 | Yes | ||
| 18 | RRAS | 18256 | 1029 | 0.273 | 0.5810 | Yes | ||
| 19 | SOX4 | 5479 | 1106 | 0.257 | 0.5868 | Yes | ||
| 20 | PPM1D | 20721 | 1259 | 0.230 | 0.5874 | Yes | ||
| 21 | PGF | 21020 | 1354 | 0.217 | 0.5907 | Yes | ||
| 22 | RAPGEF5 | 5739 10155 21123 | 1426 | 0.208 | 0.5948 | Yes | ||
| 23 | MEF2C | 3204 9378 | 1704 | 0.168 | 0.5863 | No | ||
| 24 | LAMB3 | 14007 | 1931 | 0.141 | 0.5794 | No | ||
| 25 | MAN2A1 | 23170 | 2141 | 0.118 | 0.5727 | No | ||
| 26 | MGAT4A | 13977 | 2245 | 0.109 | 0.5713 | No | ||
| 27 | BCL11A | 4691 | 2248 | 0.109 | 0.5754 | No | ||
| 28 | STARD3 | 20675 | 2327 | 0.101 | 0.5750 | No | ||
| 29 | JUP | 4937 | 2364 | 0.098 | 0.5769 | No | ||
| 30 | GOT1 | 23670 | 2415 | 0.095 | 0.5778 | No | ||
| 31 | EFNB1 | 24285 | 2748 | 0.072 | 0.5626 | No | ||
| 32 | ARCN1 | 19143 | 2926 | 0.062 | 0.5554 | No | ||
| 33 | ERF | 4680 8914 | 3047 | 0.056 | 0.5510 | No | ||
| 34 | NFIA | 16172 5170 | 3326 | 0.045 | 0.5377 | No | ||
| 35 | CLDN10 | 21935 2785 2601 2720 7188 | 3560 | 0.038 | 0.5265 | No | ||
| 36 | HOXA9 | 17147 9109 1024 | 3829 | 0.031 | 0.5132 | No | ||
| 37 | XPR1 | 5383 | 3876 | 0.030 | 0.5118 | No | ||
| 38 | ZBTB5 | 15891 | 3882 | 0.030 | 0.5127 | No | ||
| 39 | KRTCAP2 | 15541 | 3888 | 0.030 | 0.5136 | No | ||
| 40 | STAT3 | 5525 9906 | 4016 | 0.027 | 0.5078 | No | ||
| 41 | SNX15 | 23993 3761 | 4074 | 0.026 | 0.5057 | No | ||
| 42 | TANK | 2839 15006 2729 | 4318 | 0.022 | 0.4933 | No | ||
| 43 | SRGAP2 | 13836 17045 6433 17046 | 4524 | 0.020 | 0.4830 | No | ||
| 44 | NOL4 | 2028 11432 | 4532 | 0.020 | 0.4833 | No | ||
| 45 | PRKAR2A | 5287 | 4540 | 0.019 | 0.4837 | No | ||
| 46 | MCAM | 3109 19479 | 4665 | 0.018 | 0.4777 | No | ||
| 47 | TNNI1 | 14116 | 4837 | 0.016 | 0.4690 | No | ||
| 48 | GCM1 | 19372 | 4856 | 0.016 | 0.4687 | No | ||
| 49 | IFNK | 11837 | 4951 | 0.016 | 0.4642 | No | ||
| 50 | WISP2 | 14745 | 5010 | 0.015 | 0.4616 | No | ||
| 51 | MNT | 20784 | 5154 | 0.014 | 0.4544 | No | ||
| 52 | IRS4 | 9183 4926 | 5237 | 0.014 | 0.4505 | No | ||
| 53 | MFN1 | 12529 1823 | 5356 | 0.013 | 0.4446 | No | ||
| 54 | RBP2 | 19347 | 5357 | 0.013 | 0.4451 | No | ||
| 55 | EPHB2 | 4675 2440 8910 | 5459 | 0.012 | 0.4401 | No | ||
| 56 | ETV4 | 20212 | 5562 | 0.012 | 0.4350 | No | ||
| 57 | APLN | 24164 | 5596 | 0.011 | 0.4336 | No | ||
| 58 | GDPD2 | 24280 | 5641 | 0.011 | 0.4317 | No | ||
| 59 | ALK | 8574 | 5746 | 0.011 | 0.4265 | No | ||
| 60 | EXTL3 | 21785 21786 3088 | 5752 | 0.011 | 0.4266 | No | ||
| 61 | DCT | 4603 21723 | 5934 | 0.010 | 0.4172 | No | ||
| 62 | STAG1 | 5520 9905 2987 | 6144 | 0.009 | 0.4062 | No | ||
| 63 | ADORA1 | 13831 | 6296 | 0.008 | 0.3983 | No | ||
| 64 | CHRD | 22817 | 6408 | 0.008 | 0.3926 | No | ||
| 65 | SYT8 | 18006 | 6421 | 0.008 | 0.3922 | No | ||
| 66 | RERE | 12722 | 6454 | 0.008 | 0.3908 | No | ||
| 67 | ANGPT1 | 8588 2260 22303 | 6479 | 0.008 | 0.3898 | No | ||
| 68 | STC1 | 21974 | 6521 | 0.008 | 0.3878 | No | ||
| 69 | WDR31 | 2446 15869 | 6805 | 0.007 | 0.3727 | No | ||
| 70 | RPS6KA5 | 2076 21005 | 6999 | 0.006 | 0.3625 | No | ||
| 71 | TREX2 | 24137 | 7076 | 0.006 | 0.3586 | No | ||
| 72 | CHGA | 21178 2151 | 7107 | 0.006 | 0.3572 | No | ||
| 73 | TGIF2 | 6013 6012 | 7128 | 0.006 | 0.3563 | No | ||
| 74 | EBF2 | 21975 | 7286 | 0.005 | 0.3480 | No | ||
| 75 | SYTL2 | 18190 1539 3730 | 7347 | 0.005 | 0.3450 | No | ||
| 76 | NKX2-2 | 5176 | 7396 | 0.005 | 0.3426 | No | ||
| 77 | MOV10 | 9403 9402 5114 | 7491 | 0.005 | 0.3376 | No | ||
| 78 | SLC4A10 | 15004 | 7549 | 0.005 | 0.3347 | No | ||
| 79 | MPP2 | 20209 | 7596 | 0.004 | 0.3324 | No | ||
| 80 | TYR | 17763 | 7704 | 0.004 | 0.3268 | No | ||
| 81 | SOX2 | 9849 15612 | 7808 | 0.004 | 0.3213 | No | ||
| 82 | SLC26A7 | 5538 2372 | 7832 | 0.004 | 0.3202 | No | ||
| 83 | MAGEL2 | 18221 | 7981 | 0.003 | 0.3123 | No | ||
| 84 | GJA3 | 21807 | 8022 | 0.003 | 0.3103 | No | ||
| 85 | SLITRK6 | 21724 | 8165 | 0.003 | 0.3027 | No | ||
| 86 | PAX3 | 13910 | 8195 | 0.003 | 0.3012 | No | ||
| 87 | NTNG2 | 2731 9330 | 8428 | 0.003 | 0.2888 | No | ||
| 88 | HRASLS | 22801 | 8477 | 0.002 | 0.2863 | No | ||
| 89 | PIPOX | 20340 | 8618 | 0.002 | 0.2787 | No | ||
| 90 | MTSS1 | 5585 22285 | 8677 | 0.002 | 0.2757 | No | ||
| 91 | TUFT1 | 1847 15253 | 8683 | 0.002 | 0.2755 | No | ||
| 92 | HBP1 | 2048 2162 21086 | 8765 | 0.002 | 0.2712 | No | ||
| 93 | NOX3 | 10322 | 8801 | 0.002 | 0.2693 | No | ||
| 94 | SP7 | 22110 | 9020 | 0.001 | 0.2576 | No | ||
| 95 | SLC24A4 | 6223 | 9191 | 0.001 | 0.2484 | No | ||
| 96 | ATP8B3 | 12513 7426 19693 | 9207 | 0.001 | 0.2476 | No | ||
| 97 | CACNB1 | 1443 1304 | 9321 | 0.001 | 0.2415 | No | ||
| 98 | SLC2A4 | 20380 | 9412 | 0.001 | 0.2367 | No | ||
| 99 | ASPN | 21644 | 9424 | 0.001 | 0.2361 | No | ||
| 100 | DLX3 | 20697 | 9537 | 0.001 | 0.2301 | No | ||
| 101 | LOXL1 | 5002 | 9779 | 0.000 | 0.2170 | No | ||
| 102 | RPE65 | 15382 | 9801 | -0.000 | 0.2158 | No | ||
| 103 | BMP6 | 8659 | 9809 | -0.000 | 0.2155 | No | ||
| 104 | HEY1 | 15378 | 9852 | -0.000 | 0.2132 | No | ||
| 105 | B2M | 72 14882 | 9954 | -0.000 | 0.2077 | No | ||
| 106 | B3GALT2 | 6498 | 10026 | -0.001 | 0.2039 | No | ||
| 107 | ASCL2 | 5079 | 10044 | -0.001 | 0.2030 | No | ||
| 108 | KCNH2 | 16592 | 10523 | -0.001 | 0.1771 | No | ||
| 109 | SALL1 | 18796 12207 | 10678 | -0.002 | 0.1688 | No | ||
| 110 | NFATC3 | 5169 | 10711 | -0.002 | 0.1672 | No | ||
| 111 | TRIM46 | 1779 15281 | 11305 | -0.003 | 0.1351 | No | ||
| 112 | ZNF462 | 6321 | 11320 | -0.003 | 0.1345 | No | ||
| 113 | TEK | 16175 | 11405 | -0.003 | 0.1300 | No | ||
| 114 | PTPN21 | 21009 | 11432 | -0.003 | 0.1287 | No | ||
| 115 | COL11A2 | 23286 | 11494 | -0.003 | 0.1255 | No | ||
| 116 | RARB | 3011 21915 | 11529 | -0.003 | 0.1238 | No | ||
| 117 | SCUBE2 | 1879 17678 | 11917 | -0.004 | 0.1030 | No | ||
| 118 | SEMA3C | 16914 | 11963 | -0.004 | 0.1007 | No | ||
| 119 | LGR6 | 11623 977 | 11984 | -0.004 | 0.0998 | No | ||
| 120 | FGF4 | 8964 | 12064 | -0.005 | 0.0957 | No | ||
| 121 | BDNF | 14926 2797 | 12108 | -0.005 | 0.0936 | No | ||
| 122 | RBM9 | 2276 13528 | 12165 | -0.005 | 0.0907 | No | ||
| 123 | LRRTM1 | 17408 | 12345 | -0.005 | 0.0812 | No | ||
| 124 | ACTG2 | 17098 | 12447 | -0.005 | 0.0759 | No | ||
| 125 | MID1 | 5097 5098 | 12523 | -0.006 | 0.0721 | No | ||
| 126 | WNT16 | 17512 1152 | 12569 | -0.006 | 0.0699 | No | ||
| 127 | STMN4 | 21977 | 12907 | -0.007 | 0.0518 | No | ||
| 128 | HIPK1 | 4851 | 12913 | -0.007 | 0.0518 | No | ||
| 129 | NLGN3 | 10878 | 12924 | -0.007 | 0.0516 | No | ||
| 130 | HIP1 | 5676 | 13246 | -0.008 | 0.0345 | No | ||
| 131 | FGF11 | 20383 | 13286 | -0.008 | 0.0327 | No | ||
| 132 | CITED2 | 5118 14477 | 13352 | -0.008 | 0.0294 | No | ||
| 133 | SESN3 | 19561 | 13371 | -0.008 | 0.0288 | No | ||
| 134 | SESN2 | 15733 | 13462 | -0.009 | 0.0243 | No | ||
| 135 | JPH4 | 21826 | 13765 | -0.010 | 0.0083 | No | ||
| 136 | SLC38A3 | 13319 | 13788 | -0.010 | 0.0075 | No | ||
| 137 | TLX3 | 20501 | 13800 | -0.010 | 0.0073 | No | ||
| 138 | THBS3 | 15542 1796 | 13973 | -0.011 | -0.0016 | No | ||
| 139 | AGR2 | 21289 | 14525 | -0.014 | -0.0310 | No | ||
| 140 | ALDOC | 20759 311 | 14615 | -0.015 | -0.0352 | No | ||
| 141 | CALM1 | 21184 | 14639 | -0.015 | -0.0359 | No | ||
| 142 | TNXB | 1557 8174 878 8175 | 14663 | -0.016 | -0.0365 | No | ||
| 143 | CACNB3 | 22373 8679 | 14881 | -0.018 | -0.0476 | No | ||
| 144 | SEMA4C | 9799 | 14897 | -0.018 | -0.0477 | No | ||
| 145 | DUSP10 | 4003 14016 | 14905 | -0.018 | -0.0474 | No | ||
| 146 | FGF17 | 21757 | 14914 | -0.018 | -0.0471 | No | ||
| 147 | PTHR1 | 5318 18985 | 14995 | -0.019 | -0.0507 | No | ||
| 148 | KLHL2 | 13381 | 15023 | -0.020 | -0.0514 | No | ||
| 149 | CLSTN3 | 17006 | 15026 | -0.020 | -0.0507 | No | ||
| 150 | NTRK3 | 2069 3652 3705 9492 | 15029 | -0.020 | -0.0501 | No | ||
| 151 | HMGA2 | 4857 9098 3341 3444 | 15088 | -0.021 | -0.0524 | No | ||
| 152 | FERD3L | 21292 | 15256 | -0.023 | -0.0606 | No | ||
| 153 | FIGF | 24212 | 15296 | -0.024 | -0.0618 | No | ||
| 154 | MAMDC1 | 21054 6798 | 15299 | -0.024 | -0.0610 | No | ||
| 155 | IGSF8 | 14048 | 15391 | -0.026 | -0.0649 | No | ||
| 156 | RETNLB | 12178 | 15407 | -0.026 | -0.0647 | No | ||
| 157 | ATP11C | 6813 24146 | 15522 | -0.029 | -0.0698 | No | ||
| 158 | DDIT4L | 15410 | 15654 | -0.032 | -0.0756 | No | ||
| 159 | HOXC4 | 4864 | 15715 | -0.035 | -0.0775 | No | ||
| 160 | PTGDS | 14658 | 15766 | -0.036 | -0.0788 | No | ||
| 161 | SNF1LK | 23033 | 16131 | -0.052 | -0.0966 | No | ||
| 162 | ACVR1 | 4334 | 16227 | -0.057 | -0.0995 | No | ||
| 163 | LAMB2 | 9265 | 16275 | -0.060 | -0.0998 | No | ||
| 164 | GBA2 | 15898 | 16319 | -0.063 | -0.0997 | No | ||
| 165 | CYR61 | 9159 15145 9311 5024 | 16341 | -0.064 | -0.0983 | No | ||
| 166 | RNF25 | 13922 3929 | 16406 | -0.069 | -0.0991 | No | ||
| 167 | BTBD3 | 10457 2975 6007 6006 | 16598 | -0.083 | -0.1063 | No | ||
| 168 | RUNX1 | 4481 | 17055 | -0.133 | -0.1259 | No | ||
| 169 | IMMP2L | 21283 | 17119 | -0.142 | -0.1239 | No | ||
| 170 | EIF4EBP3 | 8429 | 17190 | -0.152 | -0.1218 | No | ||
| 171 | SUPT3H | 23228 | 17235 | -0.160 | -0.1180 | No | ||
| 172 | SLC15A2 | 22600 | 17347 | -0.178 | -0.1172 | No | ||
| 173 | ENO3 | 8905 | 17566 | -0.219 | -0.1206 | No | ||
| 174 | PNKP | 4052 18258 | 17753 | -0.258 | -0.1207 | No | ||
| 175 | EXOSC2 | 15044 | 17879 | -0.297 | -0.1160 | No | ||
| 176 | MTX1 | 15282 1918 | 18148 | -0.407 | -0.1149 | No | ||
| 177 | CAMTA2 | 10119 | 18370 | -0.597 | -0.1039 | No | ||
| 178 | HTF9C | 4886 1678 | 18409 | -0.659 | -0.0805 | No | ||
| 179 | MBNL1 | 1921 15582 | 18427 | -0.703 | -0.0544 | No | ||
| 180 | TARBP2 | 10033 10034 5626 | 18465 | -0.792 | -0.0258 | No | ||
| 181 | SSBP2 | 3171 7364 3252 | 18488 | -0.881 | 0.0069 | No |