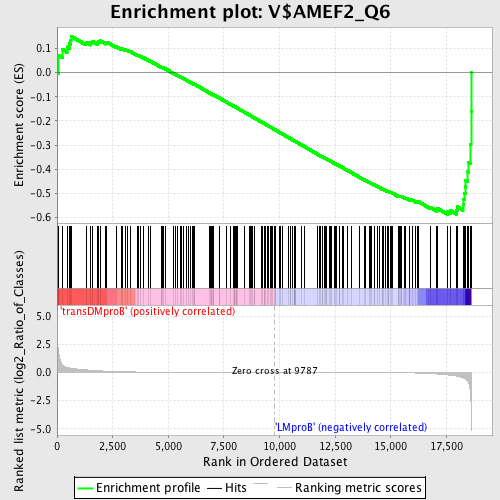
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_LMproB.phenotype_transDMproB_versus_LMproB.cls #transDMproB_versus_LMproB |
| Phenotype | phenotype_transDMproB_versus_LMproB.cls#transDMproB_versus_LMproB |
| Upregulated in class | LMproB |
| GeneSet | V$AMEF2_Q6 |
| Enrichment Score (ES) | -0.58789873 |
| Normalized Enrichment Score (NES) | -1.4799086 |
| Nominal p-value | 0.002793296 |
| FDR q-value | 0.47945237 |
| FWER p-Value | 0.968 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | FGF13 | 4720 8963 2627 | 81 | 1.558 | 0.0715 | No | ||
| 2 | HMGN2 | 9095 | 238 | 0.678 | 0.0961 | No | ||
| 3 | CDC42EP3 | 6436 | 447 | 0.444 | 0.1065 | No | ||
| 4 | ZFP36L1 | 4453 4454 | 557 | 0.389 | 0.1195 | No | ||
| 5 | CLIC1 | 23262 | 601 | 0.369 | 0.1352 | No | ||
| 6 | MBNL2 | 21932 | 626 | 0.363 | 0.1515 | No | ||
| 7 | ARL6IP2 | 1496 7090 12093 | 1298 | 0.226 | 0.1262 | No | ||
| 8 | HLX1 | 9093 | 1511 | 0.194 | 0.1241 | No | ||
| 9 | GPSN2 | 18823 3868 | 1568 | 0.187 | 0.1302 | No | ||
| 10 | ATP5H | 12948 | 1804 | 0.157 | 0.1251 | No | ||
| 11 | BIRC2 | 4397 4398 | 1839 | 0.152 | 0.1307 | No | ||
| 12 | PHKA2 | 8471 24216 | 1942 | 0.139 | 0.1319 | No | ||
| 13 | TMEM24 | 19150 | 2185 | 0.115 | 0.1244 | No | ||
| 14 | KPNA3 | 21797 | 2238 | 0.110 | 0.1269 | No | ||
| 15 | B4GALT1 | 15918 | 2660 | 0.078 | 0.1079 | No | ||
| 16 | MYL2 | 16713 | 2900 | 0.063 | 0.0980 | No | ||
| 17 | CDKL5 | 24019 | 2917 | 0.063 | 0.1002 | No | ||
| 18 | P2RX5 | 1370 20790 | 3062 | 0.055 | 0.0951 | No | ||
| 19 | CPNE1 | 11186 | 3146 | 0.052 | 0.0931 | No | ||
| 20 | PPP1R16B | 6014 | 3283 | 0.046 | 0.0880 | No | ||
| 21 | SOX5 | 9850 1044 5480 16937 | 3612 | 0.036 | 0.0719 | No | ||
| 22 | USP47 | 6794 | 3646 | 0.035 | 0.0719 | No | ||
| 23 | FBXW11 | 20926 | 3738 | 0.033 | 0.0686 | No | ||
| 24 | SMYD1 | 4452 | 3893 | 0.030 | 0.0617 | No | ||
| 25 | CHD2 | 10847 | 4086 | 0.026 | 0.0525 | No | ||
| 26 | USP2 | 19480 3052 3043 | 4193 | 0.024 | 0.0479 | No | ||
| 27 | HOXC6 | 22339 2302 | 4688 | 0.018 | 0.0220 | No | ||
| 28 | DKK1 | 23700 | 4692 | 0.018 | 0.0227 | No | ||
| 29 | RORB | 10356 | 4724 | 0.018 | 0.0219 | No | ||
| 30 | LMCD1 | 17337 1117 | 4760 | 0.017 | 0.0209 | No | ||
| 31 | CTBP2 | 17591 1137 | 4870 | 0.016 | 0.0157 | No | ||
| 32 | GSC | 20990 | 5247 | 0.013 | -0.0040 | No | ||
| 33 | GJB6 | 21805 | 5323 | 0.013 | -0.0074 | No | ||
| 34 | TBX4 | 5633 | 5412 | 0.012 | -0.0116 | No | ||
| 35 | SV2A | 15498 12250 | 5548 | 0.012 | -0.0183 | No | ||
| 36 | JMJD1C | 11531 19996 | 5574 | 0.012 | -0.0191 | No | ||
| 37 | BNC2 | 15850 | 5694 | 0.011 | -0.0250 | No | ||
| 38 | NFIB | 15855 | 5832 | 0.010 | -0.0320 | No | ||
| 39 | HAS2 | 22293 | 5921 | 0.010 | -0.0363 | No | ||
| 40 | KBTBD5 | 13020 | 6016 | 0.009 | -0.0409 | No | ||
| 41 | PRRX1 | 9595 5271 | 6101 | 0.009 | -0.0450 | No | ||
| 42 | PPARGC1A | 16533 | 6148 | 0.009 | -0.0471 | No | ||
| 43 | TACR1 | 1082 17403 | 6172 | 0.009 | -0.0479 | No | ||
| 44 | HOXB6 | 20688 | 6175 | 0.009 | -0.0476 | No | ||
| 45 | ITGA7 | 19835 | 6838 | 0.007 | -0.0831 | No | ||
| 46 | CAPN1 | 23989 | 6909 | 0.006 | -0.0866 | No | ||
| 47 | MAS1 | 23122 | 6941 | 0.006 | -0.0880 | No | ||
| 48 | LUZP1 | 6533 11257 2503 | 6958 | 0.006 | -0.0886 | No | ||
| 49 | CACNG3 | 12029 | 6984 | 0.006 | -0.0896 | No | ||
| 50 | TLE3 | 5767 19416 | 7040 | 0.006 | -0.0923 | No | ||
| 51 | FST | 8985 | 7050 | 0.006 | -0.0925 | No | ||
| 52 | CHRDL1 | 8180 13504 | 7299 | 0.005 | -0.1057 | No | ||
| 53 | LYN | 16281 | 7301 | 0.005 | -0.1055 | No | ||
| 54 | MTUS1 | 4162 158 | 7598 | 0.004 | -0.1213 | No | ||
| 55 | SOX2 | 9849 15612 | 7808 | 0.004 | -0.1325 | No | ||
| 56 | NR0B2 | 16050 | 7916 | 0.004 | -0.1381 | No | ||
| 57 | RGS3 | 2434 2474 6937 11935 2374 2332 2445 2390 | 7971 | 0.003 | -0.1409 | No | ||
| 58 | NR4A1 | 9099 | 7973 | 0.003 | -0.1407 | No | ||
| 59 | SNCAIP | 12589 | 8013 | 0.003 | -0.1427 | No | ||
| 60 | BHLHB5 | 15629 | 8041 | 0.003 | -0.1440 | No | ||
| 61 | CKMT2 | 13350 | 8076 | 0.003 | -0.1457 | No | ||
| 62 | SLC24A2 | 8020 13326 8021 | 8129 | 0.003 | -0.1483 | No | ||
| 63 | GNAO1 | 4785 3829 | 8406 | 0.003 | -0.1632 | No | ||
| 64 | HAPLN1 | 21592 | 8651 | 0.002 | -0.1763 | No | ||
| 65 | ELAVL2 | 15839 | 8687 | 0.002 | -0.1781 | No | ||
| 66 | NOG | 13663 | 8755 | 0.002 | -0.1816 | No | ||
| 67 | NOX3 | 10322 | 8801 | 0.002 | -0.1840 | No | ||
| 68 | TNNI3K | 11495 | 8887 | 0.002 | -0.1885 | No | ||
| 69 | COL13A1 | 8763 | 8891 | 0.002 | -0.1886 | No | ||
| 70 | ACCN1 | 20327 1194 | 9194 | 0.001 | -0.2049 | No | ||
| 71 | BCL9L | 19475 | 9246 | 0.001 | -0.2076 | No | ||
| 72 | HOXA10 | 17145 989 | 9306 | 0.001 | -0.2108 | No | ||
| 73 | MYL1 | 4015 | 9319 | 0.001 | -0.2114 | No | ||
| 74 | SLC4A4 | 16807 12035 7036 | 9363 | 0.001 | -0.2137 | No | ||
| 75 | RBM8A | 12239 7212 | 9437 | 0.001 | -0.2176 | No | ||
| 76 | HIST1H1E | 21520 | 9490 | 0.001 | -0.2204 | No | ||
| 77 | SLC3A1 | 23145 | 9577 | 0.000 | -0.2250 | No | ||
| 78 | DNAJA2 | 12133 18806 | 9629 | 0.000 | -0.2278 | No | ||
| 79 | MYO9A | 11325 | 9662 | 0.000 | -0.2295 | No | ||
| 80 | MYH6 | 9436 | 9792 | -0.000 | -0.2365 | No | ||
| 81 | PYY | 20207 | 9798 | -0.000 | -0.2367 | No | ||
| 82 | HOXA2 | 17151 | 9807 | -0.000 | -0.2372 | No | ||
| 83 | TTN | 14552 2743 14553 | 9975 | -0.000 | -0.2462 | No | ||
| 84 | UBXD3 | 15701 | 10042 | -0.001 | -0.2498 | No | ||
| 85 | SEMA3A | 16919 | 10151 | -0.001 | -0.2556 | No | ||
| 86 | GRIN2B | 16957 | 10419 | -0.001 | -0.2700 | No | ||
| 87 | HOXB8 | 9111 | 10475 | -0.001 | -0.2729 | No | ||
| 88 | PACSIN3 | 14948 8149 2776 2665 | 10476 | -0.001 | -0.2728 | No | ||
| 89 | USP13 | 13047 | 10586 | -0.001 | -0.2787 | No | ||
| 90 | CMYA1 | 18966 | 10659 | -0.002 | -0.2825 | No | ||
| 91 | SALL1 | 18796 12207 | 10678 | -0.002 | -0.2834 | No | ||
| 92 | NEDD4 | 19384 3101 | 10684 | -0.002 | -0.2836 | No | ||
| 93 | GDF8 | 9411 | 10719 | -0.002 | -0.2853 | No | ||
| 94 | EDG1 | 15187 | 10722 | -0.002 | -0.2854 | No | ||
| 95 | SMARCA1 | 24165 | 10996 | -0.002 | -0.3001 | No | ||
| 96 | EDN1 | 21658 | 11116 | -0.003 | -0.3064 | No | ||
| 97 | GPR27 | 13703 | 11690 | -0.004 | -0.3373 | No | ||
| 98 | IGJ | 4903 | 11802 | -0.004 | -0.3431 | No | ||
| 99 | GTF2A1 | 2064 8187 8188 13512 | 11847 | -0.004 | -0.3453 | No | ||
| 100 | FGF9 | 8966 | 11912 | -0.004 | -0.3485 | No | ||
| 101 | CNTN1 | 4538 | 11938 | -0.004 | -0.3497 | No | ||
| 102 | GABRA6 | 20491 | 12023 | -0.004 | -0.3540 | No | ||
| 103 | NRP2 | 5194 9486 5195 | 12077 | -0.005 | -0.3567 | No | ||
| 104 | BDNF | 14926 2797 | 12108 | -0.005 | -0.3581 | No | ||
| 105 | UXS1 | 13965 | 12121 | -0.005 | -0.3585 | No | ||
| 106 | ASB18 | 13886 | 12228 | -0.005 | -0.3640 | No | ||
| 107 | TNNC1 | 22058 | 12269 | -0.005 | -0.3659 | No | ||
| 108 | FGF16 | 24270 | 12311 | -0.005 | -0.3679 | No | ||
| 109 | SSH3 | 3709 10890 | 12472 | -0.006 | -0.3763 | No | ||
| 110 | MID1 | 5097 5098 | 12523 | -0.006 | -0.3787 | No | ||
| 111 | HDAC9 | 21084 2131 | 12544 | -0.006 | -0.3795 | No | ||
| 112 | SSPN | 9246 | 12670 | -0.006 | -0.3860 | No | ||
| 113 | HFE2 | 1777 15495 | 12672 | -0.006 | -0.3858 | No | ||
| 114 | RTN1 | 4177 2099 | 12844 | -0.007 | -0.3947 | No | ||
| 115 | RUNX2 | 4480 8700 | 12851 | -0.007 | -0.3947 | No | ||
| 116 | ABCA6 | 1470 20165 | 13050 | -0.007 | -0.4051 | No | ||
| 117 | CREB5 | 10551 | 13237 | -0.008 | -0.4148 | No | ||
| 118 | CASQ2 | 15477 | 13610 | -0.009 | -0.4345 | No | ||
| 119 | COL9A1 | 3950 4549 | 13837 | -0.010 | -0.4462 | No | ||
| 120 | DLGAP4 | 2950 10461 | 13861 | -0.011 | -0.4470 | No | ||
| 121 | OLIG3 | 20084 | 14020 | -0.011 | -0.4550 | No | ||
| 122 | GPC4 | 24153 | 14100 | -0.012 | -0.4587 | No | ||
| 123 | ASB16 | 20644 | 14122 | -0.012 | -0.4592 | No | ||
| 124 | ESRRG | 6447 | 14246 | -0.013 | -0.4653 | No | ||
| 125 | PURA | 9670 | 14263 | -0.013 | -0.4655 | No | ||
| 126 | WIF1 | 19866 | 14396 | -0.014 | -0.4720 | No | ||
| 127 | SOX15 | 9848 | 14482 | -0.014 | -0.4760 | No | ||
| 128 | CKM | 18362 | 14614 | -0.015 | -0.4823 | No | ||
| 129 | HTR7 | 23690 | 14623 | -0.015 | -0.4820 | No | ||
| 130 | FLT1 | 3483 16287 | 14676 | -0.016 | -0.4841 | No | ||
| 131 | SDC1 | 5560 9953 | 14755 | -0.016 | -0.4875 | No | ||
| 132 | SLC26A9 | 11560 980 4067 | 14859 | -0.018 | -0.4922 | No | ||
| 133 | CACNB3 | 22373 8679 | 14881 | -0.018 | -0.4925 | No | ||
| 134 | ACTA1 | 18715 | 14894 | -0.018 | -0.4923 | No | ||
| 135 | ARHGEF2 | 4987 | 14989 | -0.019 | -0.4964 | No | ||
| 136 | CYP26B1 | 17093 | 15018 | -0.020 | -0.4970 | No | ||
| 137 | RFX3 | 9722 5377 3712 3762 | 15090 | -0.021 | -0.4998 | No | ||
| 138 | SLC35A2 | 24389 | 15359 | -0.025 | -0.5131 | No | ||
| 139 | HOXB5 | 20687 | 15360 | -0.025 | -0.5119 | No | ||
| 140 | PCYT1B | 24289 | 15389 | -0.026 | -0.5121 | No | ||
| 141 | TGFB3 | 10161 | 15414 | -0.026 | -0.5121 | No | ||
| 142 | MRPS18B | 7365 | 15471 | -0.028 | -0.5138 | No | ||
| 143 | STARD7 | 14866 2722 | 15495 | -0.028 | -0.5137 | No | ||
| 144 | TPM2 | 2367 2437 | 15616 | -0.031 | -0.5187 | No | ||
| 145 | BACE2 | 1738 22687 1695 | 15665 | -0.033 | -0.5197 | No | ||
| 146 | CLASP1 | 14159 | 15839 | -0.039 | -0.5272 | No | ||
| 147 | LIN28 | 15723 | 15852 | -0.039 | -0.5259 | No | ||
| 148 | ELMO3 | 10629 18495 | 15855 | -0.039 | -0.5241 | No | ||
| 149 | ETV1 | 4688 8925 8924 | 15981 | -0.045 | -0.5287 | No | ||
| 150 | RHOBTB1 | 12789 | 16130 | -0.052 | -0.5342 | No | ||
| 151 | HIVEP3 | 4973 | 16207 | -0.056 | -0.5356 | No | ||
| 152 | EYA1 | 4695 4061 | 16241 | -0.058 | -0.5346 | No | ||
| 153 | CRTAP | 18979 | 16789 | -0.102 | -0.5593 | No | ||
| 154 | RBBP6 | 5365 | 17059 | -0.133 | -0.5673 | No | ||
| 155 | PACS1 | 23975 | 17105 | -0.140 | -0.5630 | No | ||
| 156 | ENO3 | 8905 | 17566 | -0.219 | -0.5772 | Yes | ||
| 157 | ANKMY2 | 5721 21288 | 17680 | -0.243 | -0.5715 | Yes | ||
| 158 | BZW2 | 21081 | 17960 | -0.326 | -0.5707 | Yes | ||
| 159 | PDLIM1 | 7023 | 17983 | -0.336 | -0.5555 | Yes | ||
| 160 | JUN | 15832 | 18245 | -0.472 | -0.5467 | Yes | ||
| 161 | ZDHHC8 | 11359 | 18276 | -0.498 | -0.5240 | Yes | ||
| 162 | EHD4 | 8275 | 18322 | -0.548 | -0.4998 | Yes | ||
| 163 | NLK | 5179 5178 | 18353 | -0.578 | -0.4733 | Yes | ||
| 164 | INPPL1 | 17733 3775 | 18372 | -0.598 | -0.4451 | Yes | ||
| 165 | DCTN1 | 1100 17392 | 18464 | -0.790 | -0.4116 | Yes | ||
| 166 | PPP1R10 | 6972 11960 6971 | 18483 | -0.855 | -0.3709 | Yes | ||
| 167 | CRHBP | 8785 | 18570 | -1.639 | -0.2957 | Yes | ||
| 168 | NFAT5 | 3921 7037 12036 | 18604 | -2.808 | -0.1607 | Yes | ||
| 169 | PABPC4 | 10510 2418 2548 | 18611 | -3.311 | 0.0003 | Yes |

