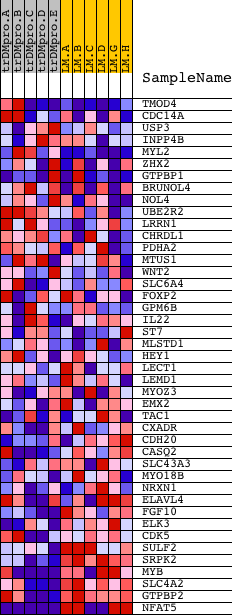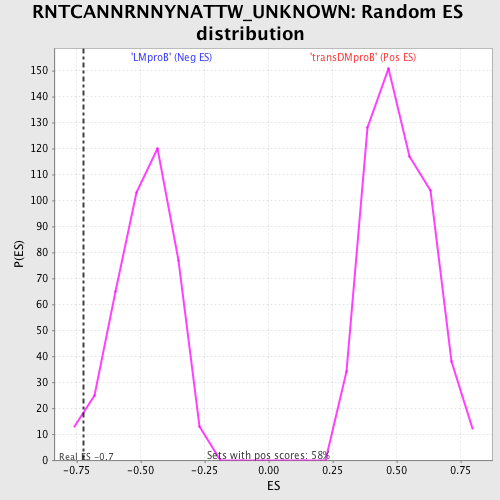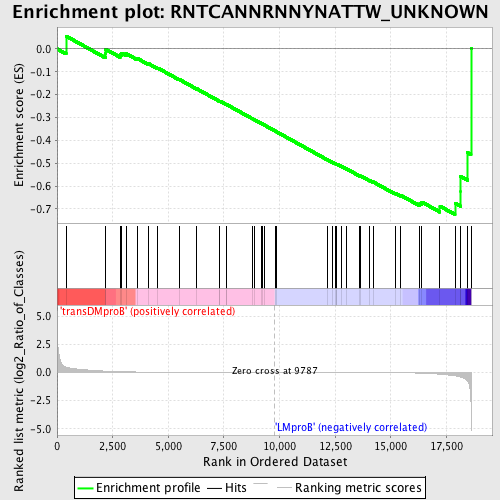
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_LMproB.phenotype_transDMproB_versus_LMproB.cls #transDMproB_versus_LMproB |
| Phenotype | phenotype_transDMproB_versus_LMproB.cls#transDMproB_versus_LMproB |
| Upregulated in class | LMproB |
| GeneSet | RNTCANNRNNYNATTW_UNKNOWN |
| Enrichment Score (ES) | -0.7244794 |
| Normalized Enrichment Score (NES) | -1.4925107 |
| Nominal p-value | 0.03125 |
| FDR q-value | 0.5313427 |
| FWER p-Value | 0.948 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | TMOD4 | 1816 1884 15509 | 404 | 0.471 | 0.0560 | No | ||
| 2 | CDC14A | 15184 | 2165 | 0.116 | -0.0195 | No | ||
| 3 | USP3 | 19074 | 2175 | 0.115 | -0.0011 | No | ||
| 4 | INPP4B | 18557 | 2829 | 0.067 | -0.0252 | No | ||
| 5 | MYL2 | 16713 | 2900 | 0.063 | -0.0185 | No | ||
| 6 | ZHX2 | 11842 | 3117 | 0.053 | -0.0214 | No | ||
| 7 | GTPBP1 | 4814 2255 | 3590 | 0.037 | -0.0408 | No | ||
| 8 | BRUNOL4 | 2010 4229 | 4102 | 0.026 | -0.0641 | No | ||
| 9 | NOL4 | 2028 11432 | 4532 | 0.020 | -0.0840 | No | ||
| 10 | UBE2R2 | 16240 | 5486 | 0.012 | -0.1333 | No | ||
| 11 | LRRN1 | 17343 | 6285 | 0.008 | -0.1749 | No | ||
| 12 | CHRDL1 | 8180 13504 | 7299 | 0.005 | -0.2286 | No | ||
| 13 | PDHA2 | 15153 | 7316 | 0.005 | -0.2286 | No | ||
| 14 | MTUS1 | 4162 158 | 7598 | 0.004 | -0.2430 | No | ||
| 15 | WNT2 | 17212 | 7606 | 0.004 | -0.2426 | No | ||
| 16 | SLC6A4 | 20770 | 8766 | 0.002 | -0.3047 | No | ||
| 17 | FOXP2 | 4312 17523 8523 | 8879 | 0.002 | -0.3104 | No | ||
| 18 | GPM6B | 9034 | 9175 | 0.001 | -0.3261 | No | ||
| 19 | IL22 | 11948 | 9232 | 0.001 | -0.3290 | No | ||
| 20 | ST7 | 1054 17519 | 9318 | 0.001 | -0.3334 | No | ||
| 21 | MLSTD1 | 17234 | 9815 | -0.000 | -0.3601 | No | ||
| 22 | HEY1 | 15378 | 9852 | -0.000 | -0.3620 | No | ||
| 23 | LECT1 | 21740 | 12132 | -0.005 | -0.4839 | No | ||
| 24 | LEMD1 | 10023 537 | 12361 | -0.005 | -0.4953 | No | ||
| 25 | MYOZ3 | 23431 | 12509 | -0.006 | -0.5023 | No | ||
| 26 | EMX2 | 23806 | 12568 | -0.006 | -0.5045 | No | ||
| 27 | TAC1 | 17526 19853 | 12571 | -0.006 | -0.5036 | No | ||
| 28 | CXADR | 22720 | 12767 | -0.006 | -0.5131 | No | ||
| 29 | CDH20 | 14168 6222 | 13013 | -0.007 | -0.5251 | No | ||
| 30 | CASQ2 | 15477 | 13610 | -0.009 | -0.5556 | No | ||
| 31 | SLC43A3 | 2824 14973 | 13657 | -0.010 | -0.5565 | No | ||
| 32 | MYO18B | 16434 | 14058 | -0.012 | -0.5761 | No | ||
| 33 | NRXN1 | 1581 1575 22875 | 14207 | -0.012 | -0.5821 | No | ||
| 34 | ELAVL4 | 15805 4889 9137 | 15191 | -0.022 | -0.6313 | No | ||
| 35 | FGF10 | 4719 8962 | 15428 | -0.027 | -0.6396 | No | ||
| 36 | ELK3 | 4663 94 3388 | 16307 | -0.062 | -0.6766 | No | ||
| 37 | CDK5 | 16591 | 16392 | -0.068 | -0.6699 | No | ||
| 38 | SULF2 | 12988 | 17209 | -0.156 | -0.6881 | Yes | ||
| 39 | SRPK2 | 5513 | 17885 | -0.299 | -0.6752 | Yes | ||
| 40 | MYB | 3344 3355 19806 | 18127 | -0.395 | -0.6230 | Yes | ||
| 41 | SLC4A2 | 9828 | 18130 | -0.398 | -0.5574 | Yes | ||
| 42 | GTPBP2 | 23221 | 18447 | -0.729 | -0.4541 | Yes | ||
| 43 | NFAT5 | 3921 7037 12036 | 18604 | -2.808 | 0.0006 | Yes |