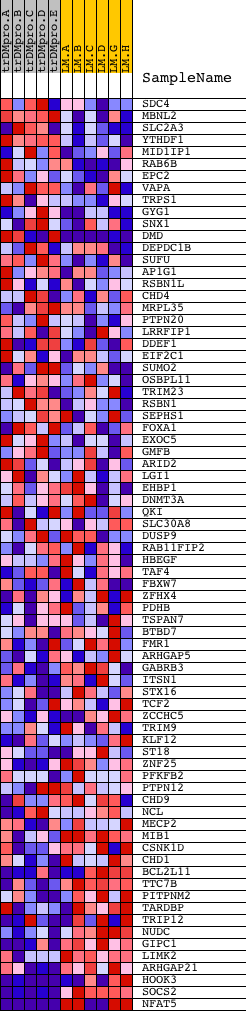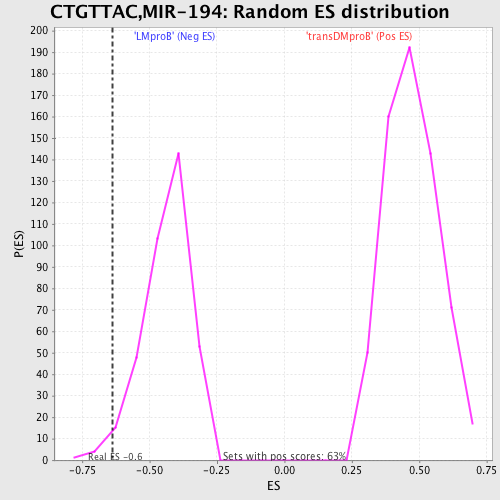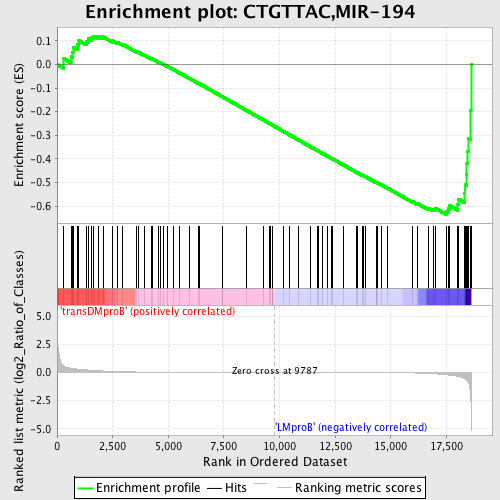
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_LMproB.phenotype_transDMproB_versus_LMproB.cls #transDMproB_versus_LMproB |
| Phenotype | phenotype_transDMproB_versus_LMproB.cls#transDMproB_versus_LMproB |
| Upregulated in class | LMproB |
| GeneSet | CTGTTAC,MIR-194 |
| Enrichment Score (ES) | -0.63569874 |
| Normalized Enrichment Score (NES) | -1.4488603 |
| Nominal p-value | 0.021798365 |
| FDR q-value | 0.5575956 |
| FWER p-Value | 0.991 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SDC4 | 14349 | 283 | 0.591 | 0.0258 | No | ||
| 2 | MBNL2 | 21932 | 626 | 0.363 | 0.0326 | No | ||
| 3 | SLC2A3 | 17010 | 707 | 0.341 | 0.0520 | No | ||
| 4 | YTHDF1 | 14311 | 728 | 0.334 | 0.0741 | No | ||
| 5 | MID1IP1 | 24381 | 907 | 0.294 | 0.0850 | No | ||
| 6 | RAB6B | 11289 | 981 | 0.282 | 0.1006 | No | ||
| 7 | EPC2 | 15015 | 1301 | 0.225 | 0.0991 | No | ||
| 8 | VAPA | 22908 | 1391 | 0.213 | 0.1091 | No | ||
| 9 | TRPS1 | 8195 | 1563 | 0.187 | 0.1129 | No | ||
| 10 | GYG1 | 1820 15371 | 1650 | 0.174 | 0.1204 | No | ||
| 11 | SNX1 | 19076 | 1858 | 0.150 | 0.1196 | No | ||
| 12 | DMD | 24295 2647 | 2070 | 0.125 | 0.1169 | No | ||
| 13 | DEPDC1B | 5757 | 2494 | 0.088 | 0.1003 | No | ||
| 14 | SUFU | 6283 | 2718 | 0.074 | 0.0934 | No | ||
| 15 | AP1G1 | 4392 | 2951 | 0.061 | 0.0851 | No | ||
| 16 | RSBN1L | 16598 | 3562 | 0.038 | 0.0548 | No | ||
| 17 | CHD4 | 8418 17281 4225 | 3663 | 0.035 | 0.0518 | No | ||
| 18 | MRPL35 | 7267 | 3930 | 0.029 | 0.0395 | No | ||
| 19 | PTPN20 | 22049 | 4242 | 0.023 | 0.0243 | No | ||
| 20 | LRRFIP1 | 14191 4007 3965 | 4274 | 0.023 | 0.0242 | No | ||
| 21 | DDEF1 | 22282 2253 | 4574 | 0.019 | 0.0094 | No | ||
| 22 | EIF2C1 | 10672 | 4651 | 0.018 | 0.0066 | No | ||
| 23 | SUMO2 | 1293 9319 | 4792 | 0.017 | 0.0002 | No | ||
| 24 | OSBPL11 | 22781 | 4966 | 0.015 | -0.0080 | No | ||
| 25 | TRIM23 | 8165 3230 | 5228 | 0.014 | -0.0212 | No | ||
| 26 | RSBN1 | 6035 | 5521 | 0.012 | -0.0361 | No | ||
| 27 | SEPHS1 | 15125 | 5953 | 0.010 | -0.0587 | No | ||
| 28 | FOXA1 | 21060 | 6372 | 0.008 | -0.0807 | No | ||
| 29 | EXOC5 | 8370 | 6383 | 0.008 | -0.0806 | No | ||
| 30 | GMFB | 21861 | 7411 | 0.005 | -0.1357 | No | ||
| 31 | ARID2 | 8062 13378 | 7436 | 0.005 | -0.1367 | No | ||
| 32 | LGI1 | 23867 | 8494 | 0.002 | -0.1935 | No | ||
| 33 | EHBP1 | 20516 | 9282 | 0.001 | -0.2359 | No | ||
| 34 | DNMT3A | 2167 21330 | 9548 | 0.000 | -0.2501 | No | ||
| 35 | QKI | 23128 5340 | 9609 | 0.000 | -0.2533 | No | ||
| 36 | SLC30A8 | 10737 22478 | 9690 | 0.000 | -0.2576 | No | ||
| 37 | DUSP9 | 24307 | 9702 | 0.000 | -0.2582 | No | ||
| 38 | RAB11FIP2 | 23635 | 10159 | -0.001 | -0.2828 | No | ||
| 39 | HBEGF | 23457 | 10444 | -0.001 | -0.2980 | No | ||
| 40 | TAF4 | 14319 | 10836 | -0.002 | -0.3189 | No | ||
| 41 | FBXW7 | 1805 11928 | 11408 | -0.003 | -0.3495 | No | ||
| 42 | ZFHX4 | 8158 | 11695 | -0.004 | -0.3647 | No | ||
| 43 | PDHB | 12670 7548 | 11756 | -0.004 | -0.3677 | No | ||
| 44 | TSPAN7 | 24382 | 11911 | -0.004 | -0.3757 | No | ||
| 45 | BTBD7 | 20998 | 12146 | -0.005 | -0.3880 | No | ||
| 46 | FMR1 | 24321 | 12324 | -0.005 | -0.3971 | No | ||
| 47 | ARHGAP5 | 4412 8625 | 12380 | -0.005 | -0.3997 | No | ||
| 48 | GABRB3 | 4747 | 12887 | -0.007 | -0.4266 | No | ||
| 49 | ITSN1 | 9196 | 13452 | -0.009 | -0.4564 | No | ||
| 50 | STX16 | 10464 | 13521 | -0.009 | -0.4594 | No | ||
| 51 | TCF2 | 1395 20727 | 13705 | -0.010 | -0.4686 | No | ||
| 52 | ZCCHC5 | 24081 | 13772 | -0.010 | -0.4715 | No | ||
| 53 | TRIM9 | 2118 8076 13577 8234 | 13841 | -0.010 | -0.4744 | No | ||
| 54 | KLF12 | 4960 2637 21732 9228 | 14343 | -0.013 | -0.5005 | No | ||
| 55 | ST18 | 14298 | 14359 | -0.013 | -0.5004 | No | ||
| 56 | ZNF25 | 17025 | 14421 | -0.014 | -0.5027 | No | ||
| 57 | PFKFB2 | 4116 5242 | 14566 | -0.015 | -0.5094 | No | ||
| 58 | PTPN12 | 16597 3502 3586 3665 | 14871 | -0.018 | -0.5246 | No | ||
| 59 | CHD9 | 6784 11553 18531 | 15951 | -0.043 | -0.5798 | No | ||
| 60 | NCL | 5153 13899 | 16193 | -0.055 | -0.5889 | No | ||
| 61 | MECP2 | 5088 | 16680 | -0.091 | -0.6088 | No | ||
| 62 | MIB1 | 5908 | 16903 | -0.114 | -0.6129 | No | ||
| 63 | CSNK1D | 4181 8352 1387 | 16987 | -0.124 | -0.6087 | No | ||
| 64 | CHD1 | 23377 | 17488 | -0.204 | -0.6215 | Yes | ||
| 65 | BCL2L11 | 2790 14861 | 17594 | -0.224 | -0.6116 | Yes | ||
| 66 | TTC7B | 21006 | 17651 | -0.237 | -0.5981 | Yes | ||
| 67 | PITPNM2 | 16371 | 18012 | -0.345 | -0.5935 | Yes | ||
| 68 | TARDBP | 10520 6066 | 18040 | -0.356 | -0.5703 | Yes | ||
| 69 | TRIP12 | 4813 9055 9054 | 18329 | -0.554 | -0.5473 | Yes | ||
| 70 | NUDC | 9495 | 18334 | -0.558 | -0.5087 | Yes | ||
| 71 | GIPC1 | 3841 3882 18552 | 18416 | -0.681 | -0.4657 | Yes | ||
| 72 | LIMK2 | 9278 1296 | 18422 | -0.690 | -0.4180 | Yes | ||
| 73 | ARHGAP21 | 12929 | 18449 | -0.738 | -0.3681 | Yes | ||
| 74 | HOOK3 | 18903 6734 | 18477 | -0.824 | -0.3122 | Yes | ||
| 75 | SOCS2 | 5694 | 18575 | -1.790 | -0.1930 | Yes | ||
| 76 | NFAT5 | 3921 7037 12036 | 18604 | -2.808 | 0.0006 | Yes |