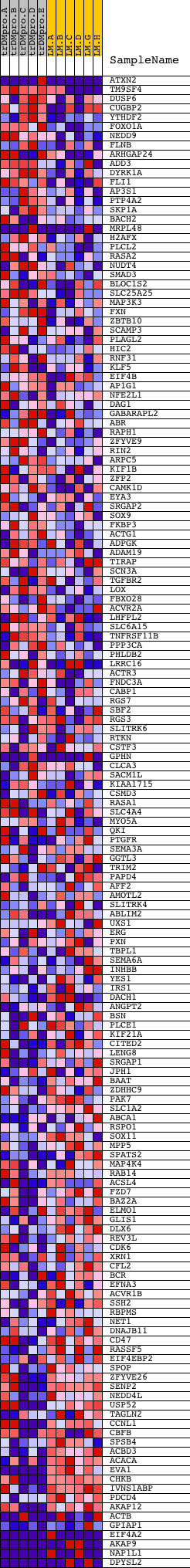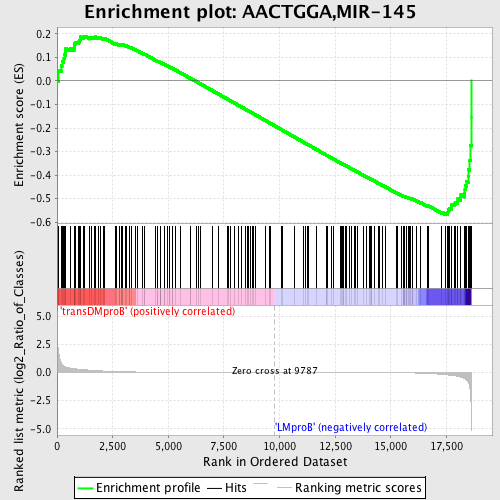
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_LMproB.phenotype_transDMproB_versus_LMproB.cls #transDMproB_versus_LMproB |
| Phenotype | phenotype_transDMproB_versus_LMproB.cls#transDMproB_versus_LMproB |
| Upregulated in class | LMproB |
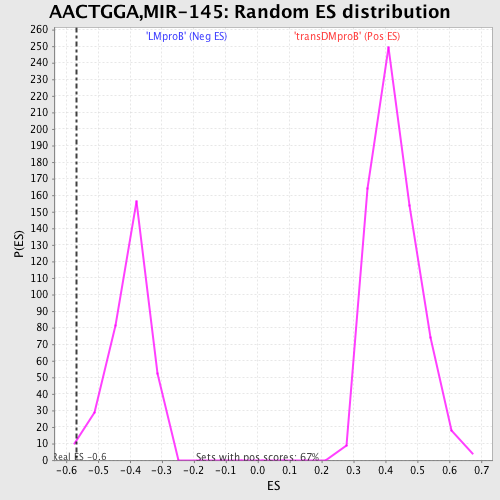
| GeneSet | AACTGGA,MIR-145 |
| Enrichment Score (ES) | -0.5674249 |
| Normalized Enrichment Score (NES) | -1.4117477 |
| Nominal p-value | 0.015243903 |
| FDR q-value | 0.5037006 |
| FWER p-Value | 0.999 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ATXN2 | 9780 3506 | 77 | 1.578 | 0.0438 | No | ||
| 2 | TM9SF4 | 14786 2830 8279 | 182 | 0.869 | 0.0646 | No | ||
| 3 | DUSP6 | 19891 3399 | 221 | 0.712 | 0.0842 | No | ||
| 4 | CUGBP2 | 4687 8923 | 307 | 0.561 | 0.0966 | No | ||
| 5 | YTHDF2 | 5623 5624 | 340 | 0.532 | 0.1111 | No | ||
| 6 | FOXO1A | 7124 | 364 | 0.506 | 0.1252 | No | ||
| 7 | NEDD9 | 3206 21483 | 392 | 0.481 | 0.1384 | No | ||
| 8 | FLNB | 13742 | 596 | 0.372 | 0.1387 | No | ||
| 9 | ARHGAP24 | 16778 3655 | 761 | 0.325 | 0.1397 | No | ||
| 10 | ADD3 | 23812 3694 | 772 | 0.321 | 0.1489 | No | ||
| 11 | DYRK1A | 4649 | 794 | 0.317 | 0.1574 | No | ||
| 12 | FLI1 | 4729 | 841 | 0.307 | 0.1642 | No | ||
| 13 | AP3S1 | 8604 | 976 | 0.283 | 0.1656 | No | ||
| 14 | PTP4A2 | 9659 2463 2518 5324 | 1006 | 0.276 | 0.1724 | No | ||
| 15 | SKP1A | 20889 | 1036 | 0.271 | 0.1791 | No | ||
| 16 | BACH2 | 8648 | 1042 | 0.269 | 0.1870 | No | ||
| 17 | MRPL48 | 3328 17739 2035 2242 3883 | 1182 | 0.244 | 0.1869 | No | ||
| 18 | H2AFX | 19477 | 1247 | 0.232 | 0.1904 | No | ||
| 19 | PLCL2 | 5902 | 1474 | 0.200 | 0.1843 | No | ||
| 20 | RASA2 | 4329 | 1565 | 0.187 | 0.1851 | No | ||
| 21 | NUDT4 | 19639 | 1678 | 0.171 | 0.1842 | No | ||
| 22 | SMAD3 | 19084 | 1706 | 0.168 | 0.1879 | No | ||
| 23 | BLOC1S2 | 23664 | 1843 | 0.152 | 0.1851 | No | ||
| 24 | SLC25A25 | 10431 | 1939 | 0.139 | 0.1842 | No | ||
| 25 | MAP3K3 | 20626 | 2075 | 0.124 | 0.1807 | No | ||
| 26 | FXN | 23709 | 2127 | 0.120 | 0.1816 | No | ||
| 27 | ZBTB10 | 10468 | 2613 | 0.081 | 0.1578 | No | ||
| 28 | SCAMP3 | 15543 | 2662 | 0.077 | 0.1575 | No | ||
| 29 | PLAGL2 | 7055 | 2811 | 0.068 | 0.1516 | No | ||
| 30 | HIC2 | 7185 | 2818 | 0.067 | 0.1533 | No | ||
| 31 | RNF31 | 11206 | 2887 | 0.064 | 0.1515 | No | ||
| 32 | KLF5 | 4456 | 2897 | 0.063 | 0.1530 | No | ||
| 33 | EIF4B | 13279 7979 | 2919 | 0.062 | 0.1537 | No | ||
| 34 | AP1G1 | 4392 | 2951 | 0.061 | 0.1539 | No | ||
| 35 | NFE2L1 | 9457 | 3072 | 0.055 | 0.1491 | No | ||
| 36 | DAG1 | 18996 8837 | 3109 | 0.053 | 0.1487 | No | ||
| 37 | GABARAPL2 | 8211 13552 | 3243 | 0.048 | 0.1430 | No | ||
| 38 | ABR | 8468 20346 1476 1429 | 3264 | 0.047 | 0.1433 | No | ||
| 39 | RAPH1 | 19110 | 3325 | 0.045 | 0.1415 | No | ||
| 40 | ZFYVE9 | 10501 2449 | 3515 | 0.039 | 0.1324 | No | ||
| 41 | RIN2 | 14817 | 3597 | 0.037 | 0.1291 | No | ||
| 42 | ARPC5 | 12580 7487 | 3855 | 0.031 | 0.1161 | No | ||
| 43 | KIF1B | 15671 2476 4950 15673 | 3912 | 0.029 | 0.1140 | No | ||
| 44 | ZFP2 | 10385 1366 1474 | 3935 | 0.029 | 0.1137 | No | ||
| 45 | CAMK1D | 5977 2948 | 4430 | 0.021 | 0.0875 | No | ||
| 46 | EYA3 | 8936 2352 | 4500 | 0.020 | 0.0844 | No | ||
| 47 | SRGAP2 | 13836 17045 6433 17046 | 4524 | 0.020 | 0.0837 | No | ||
| 48 | SOX9 | 20611 | 4634 | 0.019 | 0.0784 | No | ||
| 49 | FKBP3 | 21056 | 4636 | 0.018 | 0.0789 | No | ||
| 50 | ACTG1 | 8535 | 4653 | 0.018 | 0.0786 | No | ||
| 51 | ADPGK | 13003 7788 19422 | 4834 | 0.016 | 0.0693 | No | ||
| 52 | ADAM19 | 8548 | 4848 | 0.016 | 0.0691 | No | ||
| 53 | TIRAP | 3075 19187 | 4972 | 0.015 | 0.0629 | No | ||
| 54 | SCN3A | 14573 | 5036 | 0.015 | 0.0600 | No | ||
| 55 | TGFBR2 | 2994 3001 10167 | 5038 | 0.015 | 0.0604 | No | ||
| 56 | LOX | 23438 | 5182 | 0.014 | 0.0530 | No | ||
| 57 | FBXO28 | 7508 12612 | 5191 | 0.014 | 0.0530 | No | ||
| 58 | ACVR2A | 8542 | 5300 | 0.013 | 0.0476 | No | ||
| 59 | LHFPL2 | 21584 | 5564 | 0.012 | 0.0337 | No | ||
| 60 | SLC6A15 | 8335 3321 | 5973 | 0.010 | 0.0119 | No | ||
| 61 | TNFRSF11B | 22296 | 5998 | 0.010 | 0.0108 | No | ||
| 62 | PPP3CA | 1863 5284 | 6266 | 0.008 | -0.0034 | No | ||
| 63 | PHLDB2 | 22581 | 6352 | 0.008 | -0.0077 | No | ||
| 64 | LRRC16 | 21515 21514 | 6435 | 0.008 | -0.0119 | No | ||
| 65 | ACTR3 | 13166 | 7005 | 0.006 | -0.0426 | No | ||
| 66 | FNDC3A | 6672 21756 | 7242 | 0.005 | -0.0552 | No | ||
| 67 | CABP1 | 16412 | 7636 | 0.004 | -0.0764 | No | ||
| 68 | RGS7 | 13743 | 7690 | 0.004 | -0.0791 | No | ||
| 69 | SBF2 | 11485 | 7794 | 0.004 | -0.0846 | No | ||
| 70 | RGS3 | 2434 2474 6937 11935 2374 2332 2445 2390 | 7971 | 0.003 | -0.0940 | No | ||
| 71 | SLITRK6 | 21724 | 8165 | 0.003 | -0.1044 | No | ||
| 72 | RTKN | 17394 1156 | 8301 | 0.003 | -0.1116 | No | ||
| 73 | CSTF3 | 10449 6002 | 8484 | 0.002 | -0.1214 | No | ||
| 74 | GPHN | 6492 21232 | 8549 | 0.002 | -0.1248 | No | ||
| 75 | CLCA3 | 6224 | 8588 | 0.002 | -0.1268 | No | ||
| 76 | SACM1L | 19253 | 8705 | 0.002 | -0.1330 | No | ||
| 77 | KIAA1715 | 7629 | 8780 | 0.002 | -0.1369 | No | ||
| 78 | CSMD3 | 10735 7938 | 8827 | 0.002 | -0.1394 | No | ||
| 79 | RASA1 | 10174 | 8933 | 0.002 | -0.1450 | No | ||
| 80 | SLC4A4 | 16807 12035 7036 | 9363 | 0.001 | -0.1682 | No | ||
| 81 | MYO5A | 19378 2978 | 9529 | 0.001 | -0.1772 | No | ||
| 82 | QKI | 23128 5340 | 9609 | 0.000 | -0.1814 | No | ||
| 83 | PTGFR | 15138 | 10095 | -0.001 | -0.2077 | No | ||
| 84 | SEMA3A | 16919 | 10151 | -0.001 | -0.2107 | No | ||
| 85 | GGTL3 | 14381 | 10689 | -0.002 | -0.2397 | No | ||
| 86 | TRIM2 | 13485 8157 | 11078 | -0.002 | -0.2606 | No | ||
| 87 | PAPD4 | 8299 8300 4146 | 11168 | -0.003 | -0.2654 | No | ||
| 88 | AFF2 | 8977 | 11260 | -0.003 | -0.2702 | No | ||
| 89 | AMOTL2 | 19339 | 11295 | -0.003 | -0.2720 | No | ||
| 90 | SLITRK4 | 24144 | 11644 | -0.004 | -0.2907 | No | ||
| 91 | ABLIM2 | 16866 3605 | 12096 | -0.005 | -0.3150 | No | ||
| 92 | UXS1 | 13965 | 12121 | -0.005 | -0.3162 | No | ||
| 93 | ERG | 1686 8915 | 12159 | -0.005 | -0.3180 | No | ||
| 94 | PXN | 5339 3573 | 12316 | -0.005 | -0.3263 | No | ||
| 95 | TBPL1 | 19805 | 12407 | -0.005 | -0.3311 | No | ||
| 96 | SEMA6A | 23439 9802 | 12748 | -0.006 | -0.3493 | No | ||
| 97 | INHBB | 13858 | 12799 | -0.006 | -0.3518 | No | ||
| 98 | YES1 | 5930 | 12834 | -0.007 | -0.3534 | No | ||
| 99 | IRS1 | 4925 | 12877 | -0.007 | -0.3555 | No | ||
| 100 | DACH1 | 4595 3371 | 12890 | -0.007 | -0.3560 | No | ||
| 101 | ANGPT2 | 14885 3783 18923 | 12981 | -0.007 | -0.3606 | No | ||
| 102 | BSN | 18997 | 13022 | -0.007 | -0.3626 | No | ||
| 103 | PLCE1 | 13156 | 13158 | -0.008 | -0.3696 | No | ||
| 104 | KIF21A | 4952 | 13236 | -0.008 | -0.3736 | No | ||
| 105 | CITED2 | 5118 14477 | 13352 | -0.008 | -0.3796 | No | ||
| 106 | LENG8 | 6114 | 13421 | -0.009 | -0.3830 | No | ||
| 107 | SRGAP1 | 19615 | 13493 | -0.009 | -0.3866 | No | ||
| 108 | JPH1 | 13994 | 13763 | -0.010 | -0.4008 | No | ||
| 109 | BAAT | 15885 | 13913 | -0.011 | -0.4086 | No | ||
| 110 | ZDHHC9 | 24163 | 14056 | -0.012 | -0.4159 | No | ||
| 111 | PAK7 | 14417 | 14063 | -0.012 | -0.4159 | No | ||
| 112 | SLC1A2 | 5453 | 14145 | -0.012 | -0.4199 | No | ||
| 113 | ABCA1 | 15881 | 14149 | -0.012 | -0.4197 | No | ||
| 114 | RSPO1 | 16087 | 14266 | -0.013 | -0.4256 | No | ||
| 115 | SOX11 | 5477 | 14467 | -0.014 | -0.4360 | No | ||
| 116 | MPP5 | 21230 7078 | 14477 | -0.014 | -0.4361 | No | ||
| 117 | SPATS2 | 13044 | 14621 | -0.015 | -0.4433 | No | ||
| 118 | MAP4K4 | 11241 | 14772 | -0.017 | -0.4510 | No | ||
| 119 | RAB14 | 12685 | 15252 | -0.023 | -0.4762 | No | ||
| 120 | ACSL4 | 24043 11936 24042 2614 6940 | 15277 | -0.024 | -0.4768 | No | ||
| 121 | FZD7 | 14242 | 15480 | -0.028 | -0.4869 | No | ||
| 122 | BAZ2A | 4371 | 15560 | -0.030 | -0.4903 | No | ||
| 123 | ELMO1 | 3275 8939 3202 3271 3254 3235 3164 3175 3274 3188 3293 3214 3218 3239 | 15589 | -0.030 | -0.4909 | No | ||
| 124 | GLIS1 | 16150 | 15626 | -0.032 | -0.4919 | No | ||
| 125 | DLX6 | 17527 | 15629 | -0.032 | -0.4910 | No | ||
| 126 | REV3L | 20050 | 15699 | -0.034 | -0.4937 | No | ||
| 127 | CDK6 | 16600 | 15720 | -0.035 | -0.4937 | No | ||
| 128 | XRN1 | 10797 | 15795 | -0.037 | -0.4966 | No | ||
| 129 | CFL2 | 2155 21069 | 15840 | -0.039 | -0.4978 | No | ||
| 130 | BCR | 8478 3384 19990 | 15891 | -0.041 | -0.4993 | No | ||
| 131 | EFNA3 | 15278 | 15955 | -0.044 | -0.5014 | No | ||
| 132 | ACVR1B | 4335 22353 | 15980 | -0.045 | -0.5013 | No | ||
| 133 | SSH2 | 6202 | 16152 | -0.053 | -0.5089 | No | ||
| 134 | RBPMS | 5369 | 16345 | -0.064 | -0.5174 | No | ||
| 135 | NET1 | 7100 | 16631 | -0.087 | -0.5302 | No | ||
| 136 | DNAJB11 | 22813 | 16698 | -0.092 | -0.5310 | No | ||
| 137 | CD47 | 4933 | 17282 | -0.166 | -0.5575 | No | ||
| 138 | RASSF5 | 13837 | 17466 | -0.199 | -0.5614 | Yes | ||
| 139 | EIF4EBP2 | 4662 | 17561 | -0.218 | -0.5598 | Yes | ||
| 140 | SPOP | 5498 | 17572 | -0.221 | -0.5536 | Yes | ||
| 141 | ZFYVE26 | 9999 | 17576 | -0.221 | -0.5471 | Yes | ||
| 142 | SENP2 | 7990 | 17626 | -0.231 | -0.5427 | Yes | ||
| 143 | NEDD4L | 8192 | 17710 | -0.249 | -0.5396 | Yes | ||
| 144 | USP52 | 3397 19841 | 17721 | -0.251 | -0.5325 | Yes | ||
| 145 | TAGLN2 | 14043 | 17746 | -0.256 | -0.5260 | Yes | ||
| 146 | CCNL1 | 15325 | 17844 | -0.284 | -0.5227 | Yes | ||
| 147 | CBFB | 3769 18499 | 17915 | -0.310 | -0.5170 | Yes | ||
| 148 | SPSB4 | 19035 | 18000 | -0.341 | -0.5112 | Yes | ||
| 149 | ACBD3 | 14025 | 18015 | -0.347 | -0.5014 | Yes | ||
| 150 | ACACA | 309 4209 | 18140 | -0.402 | -0.4959 | Yes | ||
| 151 | EVA1 | 19472 | 18150 | -0.407 | -0.4840 | Yes | ||
| 152 | CHKB | 4518 | 18321 | -0.547 | -0.4766 | Yes | ||
| 153 | IVNS1ABP | 4384 4050 | 18332 | -0.558 | -0.4602 | Yes | ||
| 154 | PDCD4 | 5232 23816 | 18348 | -0.571 | -0.4436 | Yes | ||
| 155 | AKAP12 | 17600 | 18415 | -0.677 | -0.4266 | Yes | ||
| 156 | ACTB | 8534 337 337 338 | 18478 | -0.827 | -0.4048 | Yes | ||
| 157 | GPIAP1 | 2819 7014 | 18504 | -0.970 | -0.3767 | Yes | ||
| 158 | EIF4A2 | 4660 1679 1645 | 18550 | -1.408 | -0.3363 | Yes | ||
| 159 | AKAP9 | 16604 16603 4150 8303 | 18585 | -2.077 | -0.2750 | Yes | ||
| 160 | NAP1L1 | 19882 3427 330 9485 5193 | 18614 | -3.991 | -0.1551 | Yes | ||
| 161 | DPYSL2 | 3122 4561 8787 | 18616 | -5.104 | -0.0000 | Yes |