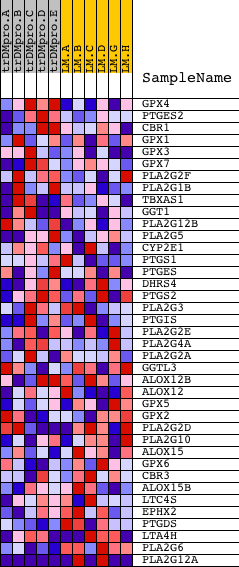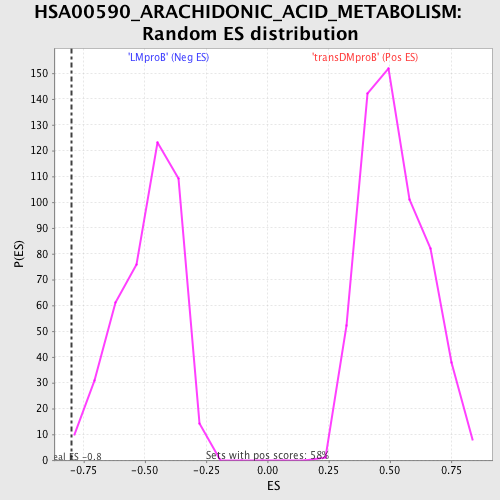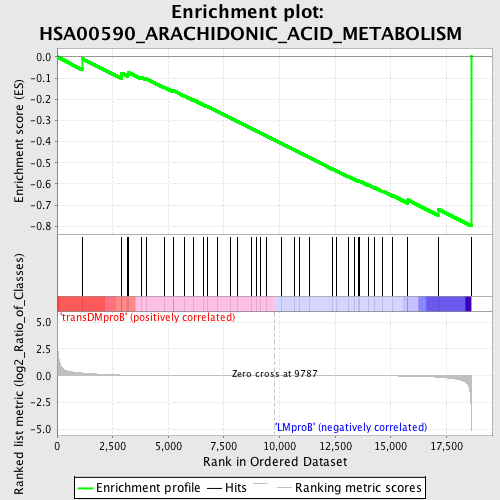
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_LMproB.phenotype_transDMproB_versus_LMproB.cls #transDMproB_versus_LMproB |
| Phenotype | phenotype_transDMproB_versus_LMproB.cls#transDMproB_versus_LMproB |
| Upregulated in class | LMproB |
| GeneSet | HSA00590_ARACHIDONIC_ACID_METABOLISM |
| Enrichment Score (ES) | -0.7985528 |
| Normalized Enrichment Score (NES) | -1.6467687 |
| Nominal p-value | 0.009433962 |
| FDR q-value | 0.38572314 |
| FWER p-Value | 0.321 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | GPX4 | 9038 | 1142 | 0.251 | -0.0088 | No | ||
| 2 | PTGES2 | 15038 | 2905 | 0.063 | -0.0904 | No | ||
| 3 | CBR1 | 22697 | 2908 | 0.063 | -0.0773 | No | ||
| 4 | GPX1 | 19310 | 3178 | 0.050 | -0.0812 | No | ||
| 5 | GPX3 | 20880 | 3221 | 0.049 | -0.0732 | No | ||
| 6 | GPX7 | 15809 | 3794 | 0.032 | -0.0973 | No | ||
| 7 | PLA2G2F | 15700 | 4012 | 0.027 | -0.1033 | No | ||
| 8 | PLA2G1B | 9581 | 4820 | 0.017 | -0.1432 | No | ||
| 9 | TBXAS1 | 17480 1105 | 5210 | 0.014 | -0.1613 | No | ||
| 10 | GGT1 | 19987 | 5239 | 0.013 | -0.1600 | No | ||
| 11 | PLA2G12B | 20016 | 5735 | 0.011 | -0.1844 | No | ||
| 12 | PLA2G5 | 9583 | 6122 | 0.009 | -0.2033 | No | ||
| 13 | CYP2E1 | 18025 | 6593 | 0.007 | -0.2270 | No | ||
| 14 | PTGS1 | 2726 5316 15028 | 6766 | 0.007 | -0.2349 | No | ||
| 15 | PTGES | 7224 | 7194 | 0.006 | -0.2567 | No | ||
| 16 | DHRS4 | 6609 | 7787 | 0.004 | -0.2877 | No | ||
| 17 | PTGS2 | 5317 9655 | 8127 | 0.003 | -0.3053 | No | ||
| 18 | PLA2G3 | 20968 | 8718 | 0.002 | -0.3367 | No | ||
| 19 | PTGIS | 9653 | 8943 | 0.002 | -0.3484 | No | ||
| 20 | PLA2G2E | 16016 | 9132 | 0.001 | -0.3582 | No | ||
| 21 | PLA2G4A | 13809 | 9405 | 0.001 | -0.3727 | No | ||
| 22 | PLA2G2A | 16017 | 10085 | -0.001 | -0.4091 | No | ||
| 23 | GGTL3 | 14381 | 10689 | -0.002 | -0.4413 | No | ||
| 24 | ALOX12B | 20825 | 10893 | -0.002 | -0.4517 | No | ||
| 25 | ALOX12 | 20372 | 11354 | -0.003 | -0.4759 | No | ||
| 26 | GPX5 | 21532 3242 | 12358 | -0.005 | -0.5288 | No | ||
| 27 | GPX2 | 9037 | 12567 | -0.006 | -0.5387 | No | ||
| 28 | PLA2G2D | 16018 | 13076 | -0.007 | -0.5645 | No | ||
| 29 | PLA2G10 | 22658 | 13349 | -0.008 | -0.5774 | No | ||
| 30 | ALOX15 | 20370 | 13531 | -0.009 | -0.5853 | No | ||
| 31 | GPX6 | 21698 | 13588 | -0.009 | -0.5863 | No | ||
| 32 | CBR3 | 22696 | 13974 | -0.011 | -0.6047 | No | ||
| 33 | ALOX15B | 20397 | 14274 | -0.013 | -0.6181 | No | ||
| 34 | LTC4S | 20478 | 14641 | -0.015 | -0.6346 | No | ||
| 35 | EPHX2 | 21778 | 15084 | -0.021 | -0.6540 | No | ||
| 36 | PTGDS | 14658 | 15766 | -0.036 | -0.6831 | No | ||
| 37 | LTA4H | 19906 3374 | 15771 | -0.036 | -0.6757 | No | ||
| 38 | PLA2G6 | 22209 | 17123 | -0.143 | -0.7184 | Yes | ||
| 39 | PLA2G12A | 1782 7281 1839 | 18613 | -3.804 | 0.0002 | Yes |