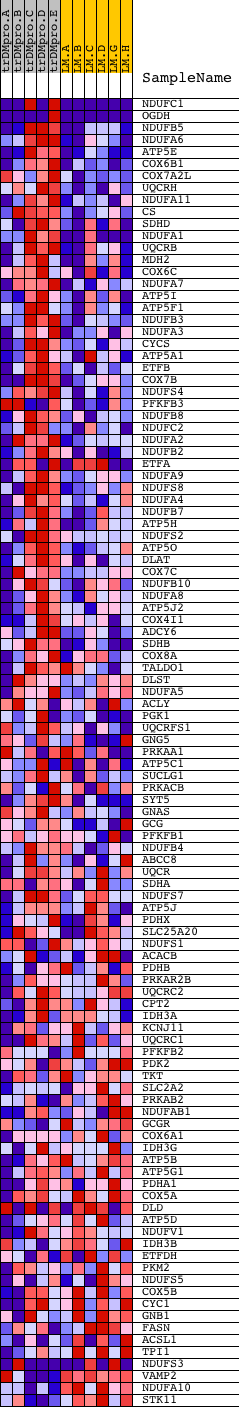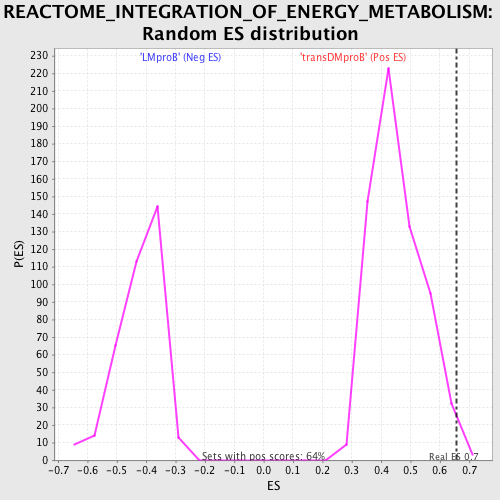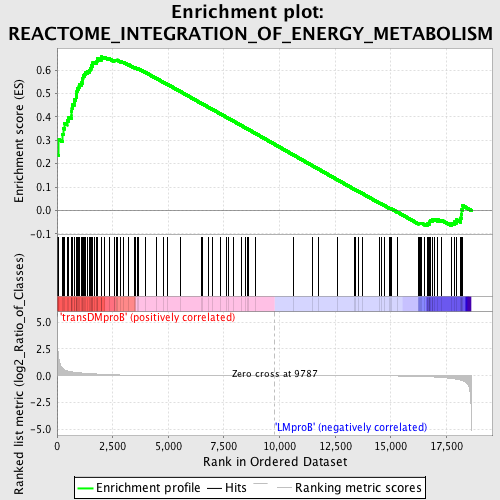
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_LMproB.phenotype_transDMproB_versus_LMproB.cls #transDMproB_versus_LMproB |
| Phenotype | phenotype_transDMproB_versus_LMproB.cls#transDMproB_versus_LMproB |
| Upregulated in class | transDMproB |
| GeneSet | REACTOME_INTEGRATION_OF_ENERGY_METABOLISM |
| Enrichment Score (ES) | 0.65691906 |
| Normalized Enrichment Score (NES) | 1.4494188 |
| Nominal p-value | 0.0062305294 |
| FDR q-value | 0.7736309 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | NDUFC1 | 7287 12339 | 0 | 5.436 | 0.2384 | Yes | ||
| 2 | OGDH | 9502 1412 | 76 | 1.582 | 0.3037 | Yes | ||
| 3 | NDUFB5 | 12277 | 229 | 0.698 | 0.3261 | Yes | ||
| 4 | NDUFA6 | 12473 | 275 | 0.620 | 0.3509 | Yes | ||
| 5 | ATP5E | 14321 | 326 | 0.541 | 0.3719 | Yes | ||
| 6 | COX6B1 | 4283 8479 | 445 | 0.445 | 0.3850 | Yes | ||
| 7 | COX7A2L | 9819 | 525 | 0.401 | 0.3984 | Yes | ||
| 8 | UQCRH | 12378 | 654 | 0.356 | 0.4070 | Yes | ||
| 9 | NDUFA11 | 12832 | 657 | 0.354 | 0.4225 | Yes | ||
| 10 | CS | 19839 | 668 | 0.349 | 0.4373 | Yes | ||
| 11 | SDHD | 19125 | 687 | 0.345 | 0.4514 | Yes | ||
| 12 | NDUFA1 | 24170 | 765 | 0.323 | 0.4614 | Yes | ||
| 13 | UQCRB | 12545 | 779 | 0.320 | 0.4747 | Yes | ||
| 14 | MDH2 | 9399 | 849 | 0.305 | 0.4844 | Yes | ||
| 15 | COX6C | 8775 | 873 | 0.300 | 0.4963 | Yes | ||
| 16 | NDUFA7 | 12347 | 888 | 0.298 | 0.5086 | Yes | ||
| 17 | ATP5I | 8637 | 905 | 0.294 | 0.5206 | Yes | ||
| 18 | ATP5F1 | 15212 | 982 | 0.282 | 0.5289 | Yes | ||
| 19 | NDUFB3 | 12362 | 1009 | 0.275 | 0.5395 | Yes | ||
| 20 | NDUFA3 | 18410 | 1107 | 0.257 | 0.5456 | Yes | ||
| 21 | CYCS | 8821 | 1127 | 0.254 | 0.5557 | Yes | ||
| 22 | ATP5A1 | 23505 | 1134 | 0.252 | 0.5664 | Yes | ||
| 23 | ETFB | 18279 | 1164 | 0.248 | 0.5757 | Yes | ||
| 24 | COX7B | 7260 | 1233 | 0.233 | 0.5822 | Yes | ||
| 25 | NDUFS4 | 9452 5157 3173 | 1272 | 0.228 | 0.5902 | Yes | ||
| 26 | PFKFB3 | 2748 | 1365 | 0.216 | 0.5947 | Yes | ||
| 27 | NDUFB8 | 7418 | 1441 | 0.205 | 0.5997 | Yes | ||
| 28 | NDUFC2 | 18183 | 1482 | 0.199 | 0.6062 | Yes | ||
| 29 | NDUFA2 | 9450 | 1526 | 0.192 | 0.6123 | Yes | ||
| 30 | NDUFB2 | 7542 | 1536 | 0.191 | 0.6202 | Yes | ||
| 31 | ETFA | 4292 8495 3155 | 1598 | 0.181 | 0.6248 | Yes | ||
| 32 | NDUFA9 | 12288 | 1608 | 0.179 | 0.6322 | Yes | ||
| 33 | NDUFS8 | 23950 | 1700 | 0.169 | 0.6347 | Yes | ||
| 34 | NDUFA4 | 9451 | 1786 | 0.159 | 0.6370 | Yes | ||
| 35 | NDUFB7 | 12429 | 1797 | 0.158 | 0.6434 | Yes | ||
| 36 | ATP5H | 12948 | 1804 | 0.157 | 0.6499 | Yes | ||
| 37 | NDUFS2 | 4089 13760 | 1978 | 0.135 | 0.6465 | Yes | ||
| 38 | ATP5O | 22539 | 1996 | 0.133 | 0.6514 | Yes | ||
| 39 | DLAT | 19123 | 2003 | 0.132 | 0.6569 | Yes | ||
| 40 | COX7C | 8776 | 2137 | 0.118 | 0.6549 | No | ||
| 41 | NDUFB10 | 23084 | 2337 | 0.100 | 0.6486 | No | ||
| 42 | NDUFA8 | 14605 | 2559 | 0.085 | 0.6403 | No | ||
| 43 | ATP5J2 | 12186 | 2580 | 0.084 | 0.6429 | No | ||
| 44 | COX4I1 | 18444 | 2649 | 0.079 | 0.6427 | No | ||
| 45 | ADCY6 | 22139 2283 8551 | 2696 | 0.075 | 0.6435 | No | ||
| 46 | SDHB | 2348 12566 14880 | 2862 | 0.065 | 0.6374 | No | ||
| 47 | COX8A | 8777 | 2965 | 0.060 | 0.6346 | No | ||
| 48 | TALDO1 | 10030 | 3223 | 0.049 | 0.6228 | No | ||
| 49 | DLST | 8131 | 3479 | 0.040 | 0.6108 | No | ||
| 50 | NDUFA5 | 1076 7544 | 3509 | 0.039 | 0.6109 | No | ||
| 51 | ACLY | 8348 | 3634 | 0.035 | 0.6058 | No | ||
| 52 | PGK1 | 9557 5244 | 3674 | 0.034 | 0.6052 | No | ||
| 53 | UQCRFS1 | 21499 | 3972 | 0.028 | 0.5904 | No | ||
| 54 | GNG5 | 9030 15393 483 | 4484 | 0.020 | 0.5636 | No | ||
| 55 | PRKAA1 | 22520 | 4782 | 0.017 | 0.5483 | No | ||
| 56 | ATP5C1 | 8635 | 4941 | 0.016 | 0.5405 | No | ||
| 57 | SUCLG1 | 17409 1059 | 5527 | 0.012 | 0.5094 | No | ||
| 58 | PRKACB | 15140 | 6469 | 0.008 | 0.4589 | No | ||
| 59 | SYT5 | 17986 | 6539 | 0.007 | 0.4555 | No | ||
| 60 | GNAS | 9025 2963 2752 | 6792 | 0.007 | 0.4421 | No | ||
| 61 | GCG | 14578 | 6990 | 0.006 | 0.4318 | No | ||
| 62 | PFKFB1 | 9552 | 7348 | 0.005 | 0.4127 | No | ||
| 63 | NDUFB4 | 22596 7541 | 7629 | 0.004 | 0.3978 | No | ||
| 64 | ABCC8 | 9941 11649 | 7713 | 0.004 | 0.3935 | No | ||
| 65 | UQCR | 12382 | 7936 | 0.004 | 0.3816 | No | ||
| 66 | SDHA | 12436 | 8271 | 0.003 | 0.3637 | No | ||
| 67 | NDUFS7 | 19946 | 8476 | 0.002 | 0.3528 | No | ||
| 68 | ATP5J | 880 22555 | 8543 | 0.002 | 0.3493 | No | ||
| 69 | PDHX | 14510 | 8609 | 0.002 | 0.3459 | No | ||
| 70 | SLC25A20 | 19307 | 8907 | 0.002 | 0.3299 | No | ||
| 71 | NDUFS1 | 13940 | 10638 | -0.002 | 0.2365 | No | ||
| 72 | ACACB | 16745 | 11472 | -0.003 | 0.1917 | No | ||
| 73 | PDHB | 12670 7548 | 11756 | -0.004 | 0.1765 | No | ||
| 74 | PRKAR2B | 5288 2107 | 12592 | -0.006 | 0.1317 | No | ||
| 75 | UQCRC2 | 18102 | 13373 | -0.008 | 0.0899 | No | ||
| 76 | CPT2 | 15812 900 | 13404 | -0.009 | 0.0887 | No | ||
| 77 | IDH3A | 19447 | 13566 | -0.009 | 0.0804 | No | ||
| 78 | KCNJ11 | 17822 | 13725 | -0.010 | 0.0723 | No | ||
| 79 | UQCRC1 | 10259 | 14484 | -0.014 | 0.0319 | No | ||
| 80 | PFKFB2 | 4116 5242 | 14566 | -0.015 | 0.0282 | No | ||
| 81 | PDK2 | 20285 | 14717 | -0.016 | 0.0208 | No | ||
| 82 | TKT | 10184 5764 | 14942 | -0.019 | 0.0095 | No | ||
| 83 | SLC2A2 | 15623 1880 1855 | 14974 | -0.019 | 0.0087 | No | ||
| 84 | PRKAB2 | 15488 | 15047 | -0.020 | 0.0057 | No | ||
| 85 | NDUFAB1 | 7667 | 15305 | -0.024 | -0.0071 | No | ||
| 86 | GCGR | 20566 | 16246 | -0.058 | -0.0554 | No | ||
| 87 | COX6A1 | 8774 4553 | 16268 | -0.060 | -0.0539 | No | ||
| 88 | IDH3G | 24132 | 16333 | -0.064 | -0.0546 | No | ||
| 89 | ATP5B | 19846 | 16374 | -0.067 | -0.0538 | No | ||
| 90 | ATP5G1 | 8636 | 16531 | -0.077 | -0.0588 | No | ||
| 91 | PDHA1 | 24020 | 16635 | -0.087 | -0.0606 | No | ||
| 92 | COX5A | 19431 | 16647 | -0.088 | -0.0573 | No | ||
| 93 | DLD | 2097 21090 | 16671 | -0.090 | -0.0546 | No | ||
| 94 | ATP5D | 19949 | 16740 | -0.097 | -0.0540 | No | ||
| 95 | NDUFV1 | 23953 | 16742 | -0.097 | -0.0498 | No | ||
| 96 | IDH3B | 9299 | 16756 | -0.099 | -0.0462 | No | ||
| 97 | ETFDH | 15313 | 16787 | -0.102 | -0.0433 | No | ||
| 98 | PKM2 | 3642 9573 | 16872 | -0.110 | -0.0431 | No | ||
| 99 | NDUFS5 | 9296 | 16878 | -0.110 | -0.0385 | No | ||
| 100 | COX5B | 4552 | 16962 | -0.121 | -0.0377 | No | ||
| 101 | CYC1 | 12348 | 17113 | -0.141 | -0.0396 | No | ||
| 102 | GNB1 | 15967 | 17272 | -0.165 | -0.0410 | No | ||
| 103 | FASN | 8953 | 17740 | -0.255 | -0.0550 | No | ||
| 104 | ACSL1 | 8948 | 17860 | -0.289 | -0.0487 | No | ||
| 105 | TPI1 | 5795 10212 | 17955 | -0.325 | -0.0396 | No | ||
| 106 | NDUFS3 | 7553 12683 | 18145 | -0.404 | -0.0321 | No | ||
| 107 | VAMP2 | 20826 | 18175 | -0.423 | -0.0151 | No | ||
| 108 | NDUFA10 | 7420 | 18192 | -0.435 | 0.0031 | No | ||
| 109 | STK11 | 19950 | 18219 | -0.450 | 0.0214 | No |