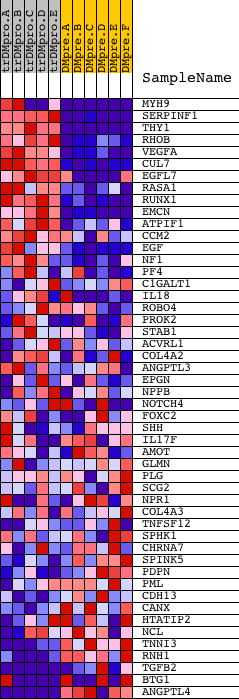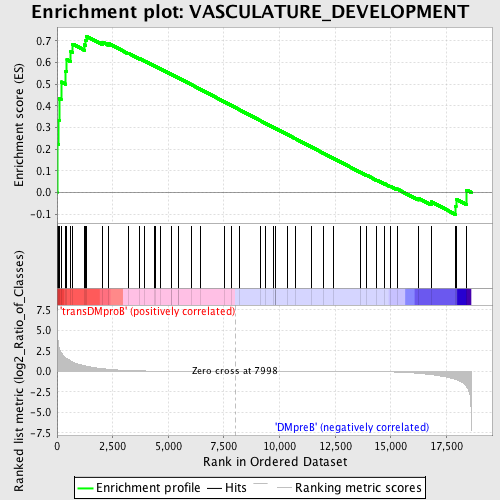
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_DMpreB.cls#transDMproB_versus_DMpreB_repos |
| Upregulated in class | transDMproB |
| GeneSet | VASCULATURE_DEVELOPMENT |
| Enrichment Score (ES) | 0.72224414 |
| Normalized Enrichment Score (NES) | 1.5183156 |
| Nominal p-value | 0.010733453 |
| FDR q-value | 0.26203603 |
| FWER p-Value | 0.978 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | MYH9 | 2252 2244 | 4 | 6.155 | 0.2240 | Yes | ||
| 2 | SERPINF1 | 9793 | 69 | 3.106 | 0.3337 | Yes | ||
| 3 | THY1 | 19481 | 93 | 2.773 | 0.4335 | Yes | ||
| 4 | RHOB | 4411 | 181 | 2.245 | 0.5106 | Yes | ||
| 5 | VEGFA | 22969 | 368 | 1.660 | 0.5610 | Yes | ||
| 6 | CUL7 | 23217 | 442 | 1.547 | 0.6135 | Yes | ||
| 7 | EGFL7 | 87 11687 2706 | 615 | 1.245 | 0.6496 | Yes | ||
| 8 | RASA1 | 10174 | 698 | 1.106 | 0.6854 | Yes | ||
| 9 | RUNX1 | 4481 | 1221 | 0.667 | 0.6816 | Yes | ||
| 10 | EMCN | 7211 | 1286 | 0.627 | 0.7010 | Yes | ||
| 11 | ATPIF1 | 4429 | 1310 | 0.617 | 0.7222 | Yes | ||
| 12 | CCM2 | 20950 | 2040 | 0.312 | 0.6944 | No | ||
| 13 | EGF | 15169 | 2310 | 0.241 | 0.6886 | No | ||
| 14 | NF1 | 5165 | 3219 | 0.082 | 0.6427 | No | ||
| 15 | PF4 | 16800 | 3714 | 0.046 | 0.6178 | No | ||
| 16 | C1GALT1 | 8246 | 3933 | 0.036 | 0.6074 | No | ||
| 17 | IL18 | 9172 | 4373 | 0.022 | 0.5845 | No | ||
| 18 | ROBO4 | 7895 3064 | 4440 | 0.020 | 0.5817 | No | ||
| 19 | PROK2 | 17057 | 4625 | 0.017 | 0.5724 | No | ||
| 20 | STAB1 | 21892 | 5143 | 0.011 | 0.5449 | No | ||
| 21 | ACVRL1 | 22354 | 5159 | 0.011 | 0.5445 | No | ||
| 22 | COL4A2 | 18688 | 5467 | 0.008 | 0.5283 | No | ||
| 23 | ANGPTL3 | 16171 | 5473 | 0.008 | 0.5283 | No | ||
| 24 | EPGN | 16798 | 6039 | 0.006 | 0.4981 | No | ||
| 25 | NPPB | 15994 | 6431 | 0.004 | 0.4772 | No | ||
| 26 | NOTCH4 | 23277 9475 | 7543 | 0.001 | 0.4174 | No | ||
| 27 | FOXC2 | 18442 | 7842 | 0.000 | 0.4013 | No | ||
| 28 | SHH | 16581 | 8203 | -0.001 | 0.3820 | No | ||
| 29 | IL17F | 13992 | 9150 | -0.003 | 0.3311 | No | ||
| 30 | AMOT | 11341 24037 6587 | 9351 | -0.003 | 0.3205 | No | ||
| 31 | GLMN | 16452 3562 | 9712 | -0.004 | 0.3012 | No | ||
| 32 | PLG | 1593 23383 | 9797 | -0.004 | 0.2969 | No | ||
| 33 | SCG2 | 5410 14425 | 9812 | -0.004 | 0.2963 | No | ||
| 34 | NPR1 | 9480 | 9833 | -0.005 | 0.2954 | No | ||
| 35 | COL4A3 | 8766 | 10340 | -0.006 | 0.2683 | No | ||
| 36 | TNFSF12 | 10209 | 10724 | -0.007 | 0.2480 | No | ||
| 37 | SPHK1 | 5484 1266 1402 | 11415 | -0.010 | 0.2112 | No | ||
| 38 | CHRNA7 | 17808 | 11963 | -0.012 | 0.1821 | No | ||
| 39 | SPINK5 | 23569 | 12405 | -0.016 | 0.1590 | No | ||
| 40 | PDPN | 15685 | 13644 | -0.030 | 0.0934 | No | ||
| 41 | PML | 3015 3074 3020 5270 | 13898 | -0.034 | 0.0810 | No | ||
| 42 | CDH13 | 4506 3826 | 14375 | -0.046 | 0.0570 | No | ||
| 43 | CANX | 4474 | 14714 | -0.060 | 0.0410 | No | ||
| 44 | HTATIP2 | 18232 | 14987 | -0.074 | 0.0291 | No | ||
| 45 | NCL | 5153 13899 | 15284 | -0.096 | 0.0166 | No | ||
| 46 | TNNI3 | 89 | 16231 | -0.246 | -0.0254 | No | ||
| 47 | RNH1 | 17562 | 16813 | -0.402 | -0.0420 | No | ||
| 48 | TGFB2 | 5743 | 17912 | -0.982 | -0.0654 | No | ||
| 49 | BTG1 | 4458 4457 8663 | 17932 | -0.993 | -0.0302 | No | ||
| 50 | ANGPTL4 | 1504 7178 | 18408 | -1.840 | 0.0112 | No |