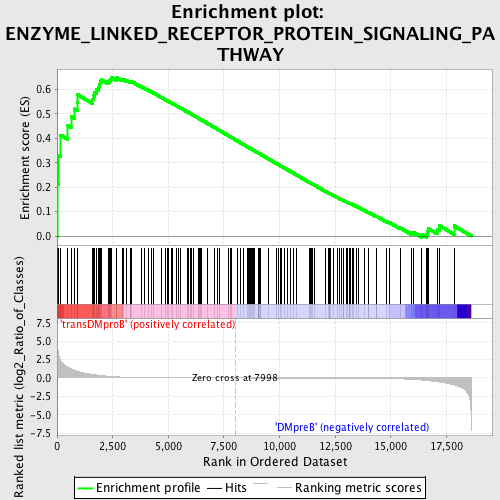
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_DMpreB.cls#transDMproB_versus_DMpreB_repos |
| Upregulated in class | transDMproB |
| GeneSet | ENZYME_LINKED_RECEPTOR_PROTEIN_SIGNALING_PATHWAY |
| Enrichment Score (ES) | 0.6489376 |
| Normalized Enrichment Score (NES) | 1.5210515 |
| Nominal p-value | 0.0033670033 |
| FDR q-value | 0.28651854 |
| FWER p-Value | 0.973 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | GRB7 | 20673 | 5 | 5.748 | 0.2151 | Yes | ||
| 2 | GAB1 | 18828 | 64 | 3.185 | 0.3312 | Yes | ||
| 3 | ENG | 4667 | 160 | 2.332 | 0.4134 | Yes | ||
| 4 | TGFB1 | 18332 | 479 | 1.483 | 0.4518 | Yes | ||
| 5 | SMAD5 | 21621 | 632 | 1.221 | 0.4893 | Yes | ||
| 6 | SMAD3 | 19084 | 770 | 1.030 | 0.5205 | Yes | ||
| 7 | PDPK1 | 23097 | 914 | 0.896 | 0.5463 | Yes | ||
| 8 | PICK1 | 9560 22424 | 931 | 0.881 | 0.5784 | Yes | ||
| 9 | ABI1 | 4300 | 1590 | 0.485 | 0.5610 | Yes | ||
| 10 | NCK2 | 9448 | 1655 | 0.455 | 0.5746 | Yes | ||
| 11 | SMAD4 | 5058 | 1687 | 0.440 | 0.5894 | Yes | ||
| 12 | LEFTY1 | 8877 | 1781 | 0.398 | 0.5993 | Yes | ||
| 13 | FIBP | 7195 | 1872 | 0.364 | 0.6081 | Yes | ||
| 14 | PTK2B | 21776 | 1904 | 0.355 | 0.6197 | Yes | ||
| 15 | GRB2 | 20149 | 1927 | 0.347 | 0.6315 | Yes | ||
| 16 | ACVR1 | 4334 | 1996 | 0.324 | 0.6400 | Yes | ||
| 17 | EGF | 15169 | 2310 | 0.241 | 0.6321 | Yes | ||
| 18 | CDKN1C | 17546 | 2365 | 0.230 | 0.6378 | Yes | ||
| 19 | FNTA | 18904 | 2418 | 0.218 | 0.6431 | Yes | ||
| 20 | AP3S1 | 8604 | 2457 | 0.210 | 0.6489 | Yes | ||
| 21 | IGF1R | 9157 | 2687 | 0.158 | 0.6425 | No | ||
| 22 | HIPK2 | 17180 | 2688 | 0.158 | 0.6484 | No | ||
| 23 | ACVR1B | 4335 22353 | 2930 | 0.114 | 0.6396 | No | ||
| 24 | MAP3K7 | 16255 | 3003 | 0.105 | 0.6397 | No | ||
| 25 | ACVR2B | 19276 | 3128 | 0.092 | 0.6364 | No | ||
| 26 | GFRA2 | 21965 | 3282 | 0.077 | 0.6310 | No | ||
| 27 | ZFYVE9 | 10501 2449 | 3299 | 0.076 | 0.6330 | No | ||
| 28 | SNF1LK2 | 6168 10649 | 3322 | 0.073 | 0.6346 | No | ||
| 29 | SMAD2 | 23511 | 3778 | 0.043 | 0.6116 | No | ||
| 30 | GDF9 | 20887 | 3940 | 0.035 | 0.6042 | No | ||
| 31 | ACVR2A | 8542 | 4102 | 0.029 | 0.5966 | No | ||
| 32 | BMPR2 | 14241 4081 | 4228 | 0.026 | 0.5908 | No | ||
| 33 | CIDEA | 23529 | 4322 | 0.023 | 0.5866 | No | ||
| 34 | ERBB2IP | 3232 12234 | 4704 | 0.015 | 0.5666 | No | ||
| 35 | PXN | 5339 3573 | 4852 | 0.013 | 0.5592 | No | ||
| 36 | PIK3R1 | 3170 | 4940 | 0.013 | 0.5549 | No | ||
| 37 | SHC3 | 21465 | 5022 | 0.012 | 0.5510 | No | ||
| 38 | ACVRL1 | 22354 | 5159 | 0.011 | 0.5440 | No | ||
| 39 | RAPGEF1 | 4218 2860 | 5189 | 0.010 | 0.5428 | No | ||
| 40 | TGFBR1 | 16213 10165 5745 | 5364 | 0.009 | 0.5338 | No | ||
| 41 | PTPRF | 5331 2545 | 5451 | 0.008 | 0.5294 | No | ||
| 42 | TGFBR3 | 16453 | 5541 | 0.008 | 0.5249 | No | ||
| 43 | KLF10 | 22308 | 5839 | 0.006 | 0.5091 | No | ||
| 44 | BDKRB2 | 21163 | 5912 | 0.006 | 0.5054 | No | ||
| 45 | DOK1 | 17104 1018 1177 | 5995 | 0.006 | 0.5012 | No | ||
| 46 | EPGN | 16798 | 6039 | 0.006 | 0.4991 | No | ||
| 47 | SOST | 20210 | 6131 | 0.005 | 0.4944 | No | ||
| 48 | ERBB3 | 19593 | 6364 | 0.004 | 0.4820 | No | ||
| 49 | PTN | 5323 | 6400 | 0.004 | 0.4803 | No | ||
| 50 | BAIAP2 | 449 4236 | 6433 | 0.004 | 0.4787 | No | ||
| 51 | GP6 | 17990 | 6489 | 0.004 | 0.4759 | No | ||
| 52 | GUCA1B | 23211 | 6748 | 0.003 | 0.4620 | No | ||
| 53 | EREG | 4679 16797 | 7087 | 0.002 | 0.4438 | No | ||
| 54 | MUSK | 5199 | 7220 | 0.002 | 0.4368 | No | ||
| 55 | PTPRT | 5333 | 7295 | 0.002 | 0.4328 | No | ||
| 56 | PTPRU | 15738 | 7724 | 0.001 | 0.4097 | No | ||
| 57 | MTSS1 | 5585 22285 | 7799 | 0.000 | 0.4057 | No | ||
| 58 | FOXC2 | 18442 | 7842 | 0.000 | 0.4034 | No | ||
| 59 | FGF5 | 16785 | 8104 | -0.000 | 0.3893 | No | ||
| 60 | GUCY2F | 24045 | 8251 | -0.001 | 0.3815 | No | ||
| 61 | CNKSR1 | 15719 20331 | 8386 | -0.001 | 0.3742 | No | ||
| 62 | RPS6KA5 | 2076 21005 | 8560 | -0.001 | 0.3649 | No | ||
| 63 | ROR1 | 6472 | 8593 | -0.001 | 0.3633 | No | ||
| 64 | OTX2 | 9519 | 8660 | -0.002 | 0.3597 | No | ||
| 65 | MPZL1 | 13777 | 8702 | -0.002 | 0.3576 | No | ||
| 66 | FMOD | 14133 | 8740 | -0.002 | 0.3557 | No | ||
| 67 | NTRK2 | 9491 | 8794 | -0.002 | 0.3529 | No | ||
| 68 | CD3E | 8714 | 8819 | -0.002 | 0.3516 | No | ||
| 69 | SMAD7 | 23513 | 8883 | -0.002 | 0.3483 | No | ||
| 70 | EDN2 | 16102 | 9046 | -0.003 | 0.3396 | No | ||
| 71 | RGMB | 12736 7585 | 9058 | -0.003 | 0.3391 | No | ||
| 72 | CBLC | 17938 | 9067 | -0.003 | 0.3388 | No | ||
| 73 | GUCY2C | 1140 16954 | 9096 | -0.003 | 0.3374 | No | ||
| 74 | SHC1 | 9813 9812 5430 | 9137 | -0.003 | 0.3353 | No | ||
| 75 | IL17F | 13992 | 9150 | -0.003 | 0.3348 | No | ||
| 76 | FRS3 | 23209 | 9495 | -0.004 | 0.3163 | No | ||
| 77 | TDGF1 | 5704 10108 | 9872 | -0.005 | 0.2962 | No | ||
| 78 | GDF5 | 4768 | 9948 | -0.005 | 0.2923 | No | ||
| 79 | IRS1 | 4925 | 10028 | -0.005 | 0.2882 | No | ||
| 80 | KDR | 16509 | 10086 | -0.005 | 0.2853 | No | ||
| 81 | TGFA | 17381 | 10236 | -0.006 | 0.2775 | No | ||
| 82 | LEFTY2 | 6735 | 10355 | -0.006 | 0.2713 | No | ||
| 83 | FOXH1 | 22238 | 10486 | -0.006 | 0.2645 | No | ||
| 84 | CITED2 | 5118 14477 | 10610 | -0.007 | 0.2581 | No | ||
| 85 | REPS2 | 24018 | 10738 | -0.007 | 0.2515 | No | ||
| 86 | SRC | 5507 | 11341 | -0.009 | 0.2193 | No | ||
| 87 | SMURF1 | 13289 7989 | 11404 | -0.010 | 0.2163 | No | ||
| 88 | BMPR1A | 8660 4450 | 11430 | -0.010 | 0.2153 | No | ||
| 89 | PTPRD | 11554 5330 | 11467 | -0.010 | 0.2138 | No | ||
| 90 | GRB10 | 4799 | 11584 | -0.010 | 0.2079 | No | ||
| 91 | FGFR3 | 8969 3566 | 12054 | -0.013 | 0.1830 | No | ||
| 92 | TEK | 16175 | 12179 | -0.014 | 0.1768 | No | ||
| 93 | IGF2 | 17550 | 12243 | -0.014 | 0.1739 | No | ||
| 94 | EFNB3 | 20391 | 12254 | -0.014 | 0.1739 | No | ||
| 95 | PTPRG | 22097 3151 | 12299 | -0.015 | 0.1721 | No | ||
| 96 | FLT3 | 16288 | 12429 | -0.016 | 0.1657 | No | ||
| 97 | EGFR | 1329 20944 | 12610 | -0.017 | 0.1566 | No | ||
| 98 | FGFR4 | 3226 | 12709 | -0.018 | 0.1520 | No | ||
| 99 | GDF10 | 4767 | 12762 | -0.018 | 0.1499 | No | ||
| 100 | MAPK8IP2 | 77 | 12876 | -0.020 | 0.1445 | No | ||
| 101 | NTRK3 | 2069 3652 3705 9492 | 12880 | -0.020 | 0.1451 | No | ||
| 102 | GDF15 | 10721 | 13009 | -0.021 | 0.1389 | No | ||
| 103 | ERBB2 | 8913 | 13044 | -0.021 | 0.1379 | No | ||
| 104 | SNX6 | 21070 | 13120 | -0.022 | 0.1347 | No | ||
| 105 | FLT4 | 20904 1461 | 13136 | -0.022 | 0.1347 | No | ||
| 106 | AKT1 | 8568 | 13176 | -0.023 | 0.1335 | No | ||
| 107 | EID2 | 11815 | 13290 | -0.024 | 0.1283 | No | ||
| 108 | NRTN | 22923 | 13336 | -0.025 | 0.1268 | No | ||
| 109 | HPGD | 4867 | 13478 | -0.027 | 0.1201 | No | ||
| 110 | PTPRJ | 9664 | 13565 | -0.028 | 0.1166 | No | ||
| 111 | CBL | 19154 | 13815 | -0.033 | 0.1043 | No | ||
| 112 | INSR | 18950 | 13984 | -0.036 | 0.0966 | No | ||
| 113 | NTRK1 | 15299 | 14008 | -0.037 | 0.0967 | No | ||
| 114 | LCP2 | 4988 9268 | 14343 | -0.045 | 0.0804 | No | ||
| 115 | CDH13 | 4506 3826 | 14375 | -0.046 | 0.0804 | No | ||
| 116 | SMAD1 | 18831 922 | 14826 | -0.065 | 0.0585 | No | ||
| 117 | PRLR | 9620 5291 | 14948 | -0.072 | 0.0546 | No | ||
| 118 | STUB1 | 23070 | 15416 | -0.112 | 0.0336 | No | ||
| 119 | EPS15 | 4677 | 15917 | -0.184 | 0.0134 | No | ||
| 120 | ADRB2 | 23422 | 16024 | -0.203 | 0.0153 | No | ||
| 121 | EPS8 | 16948 | 16398 | -0.284 | 0.0058 | No | ||
| 122 | CDKN2B | 15840 | 16616 | -0.344 | 0.0069 | No | ||
| 123 | LTBP2 | 5007 | 16654 | -0.358 | 0.0183 | No | ||
| 124 | SOCS1 | 4522 | 16690 | -0.364 | 0.0300 | No | ||
| 125 | BCAR1 | 18741 | 17079 | -0.500 | 0.0278 | No | ||
| 126 | AGRN | 4361 | 17174 | -0.539 | 0.0429 | No | ||
| 127 | PIK3R3 | 5248 | 17847 | -0.935 | 0.0416 | No |

