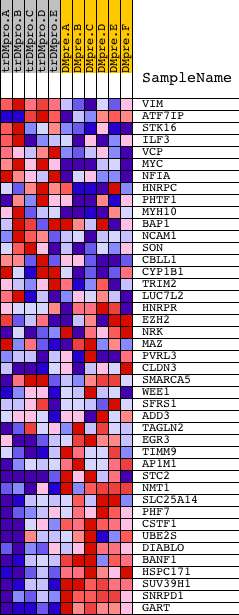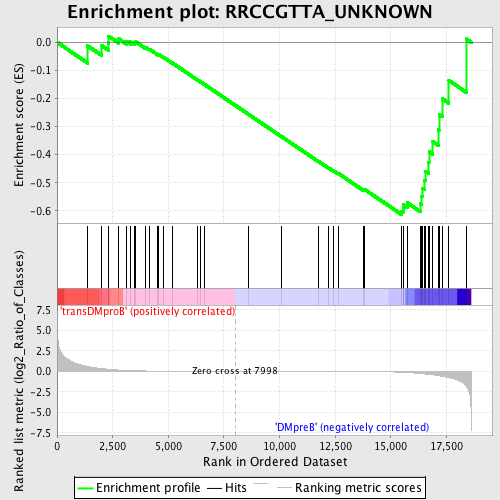
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_DMpreB.cls#transDMproB_versus_DMpreB_repos |
| Upregulated in class | DMpreB |
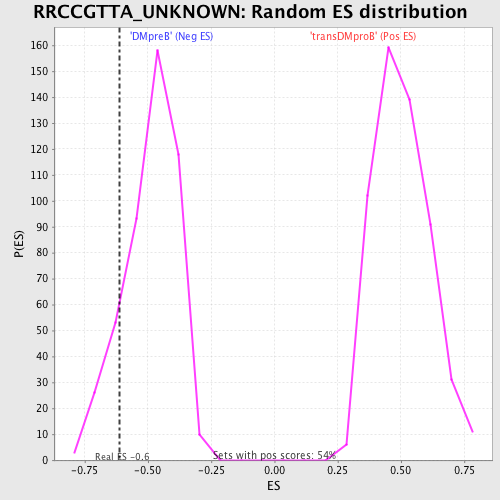
| GeneSet | RRCCGTTA_UNKNOWN |
| Enrichment Score (ES) | -0.6126868 |
| Normalized Enrichment Score (NES) | -1.2585087 |
| Nominal p-value | 0.123644255 |
| FDR q-value | 0.6409869 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | VIM | 80 | 1380 | 0.582 | -0.0131 | No | ||
| 2 | ATF7IP | 12028 13038 | 1986 | 0.329 | -0.0111 | No | ||
| 3 | STK16 | 14216 | 2300 | 0.243 | -0.0024 | No | ||
| 4 | ILF3 | 3110 3030 9176 | 2318 | 0.239 | 0.0219 | No | ||
| 5 | VCP | 11254 | 2778 | 0.140 | 0.0119 | No | ||
| 6 | MYC | 22465 9435 | 3120 | 0.093 | 0.0033 | No | ||
| 7 | NFIA | 16172 5170 | 3302 | 0.075 | 0.0015 | No | ||
| 8 | HNRPC | 9102 4859 | 3463 | 0.062 | -0.0006 | No | ||
| 9 | PHTF1 | 15469 | 3537 | 0.057 | 0.0014 | No | ||
| 10 | MYH10 | 8077 | 3952 | 0.035 | -0.0172 | No | ||
| 11 | BAP1 | 22057 | 4139 | 0.028 | -0.0242 | No | ||
| 12 | NCAM1 | 5149 | 4533 | 0.018 | -0.0434 | No | ||
| 13 | SON | 5473 1657 1684 | 4567 | 0.018 | -0.0434 | No | ||
| 14 | CBLL1 | 4188 | 4782 | 0.014 | -0.0534 | No | ||
| 15 | CYP1B1 | 4581 | 5172 | 0.011 | -0.0732 | No | ||
| 16 | TRIM2 | 13485 8157 | 6310 | 0.005 | -0.1340 | No | ||
| 17 | LUC7L2 | 5313 5312 | 6453 | 0.004 | -0.1412 | No | ||
| 18 | HNRPR | 13196 7909 2500 | 6621 | 0.003 | -0.1498 | No | ||
| 19 | EZH2 | 1092 17163 | 8583 | -0.001 | -0.2552 | No | ||
| 20 | NRK | 6559 | 10105 | -0.005 | -0.3366 | No | ||
| 21 | MAZ | 1327 17623 | 11736 | -0.011 | -0.4232 | No | ||
| 22 | PVRL3 | 1674 22579 1675 | 12205 | -0.014 | -0.4469 | No | ||
| 23 | CLDN3 | 16680 | 12421 | -0.016 | -0.4568 | No | ||
| 24 | SMARCA5 | 8215 | 12634 | -0.017 | -0.4664 | No | ||
| 25 | WEE1 | 18127 | 13773 | -0.032 | -0.5243 | No | ||
| 26 | SFRS1 | 8492 | 13823 | -0.033 | -0.5234 | No | ||
| 27 | ADD3 | 23812 3694 | 15482 | -0.118 | -0.6003 | Yes | ||
| 28 | TAGLN2 | 14043 | 15551 | -0.127 | -0.5906 | Yes | ||
| 29 | EGR3 | 4656 | 15573 | -0.129 | -0.5782 | Yes | ||
| 30 | TIMM9 | 19686 | 15736 | -0.152 | -0.5709 | Yes | ||
| 31 | AP1M1 | 18571 | 16333 | -0.269 | -0.5747 | Yes | ||
| 32 | STC2 | 5526 | 16391 | -0.282 | -0.5480 | Yes | ||
| 33 | NMT1 | 20638 | 16432 | -0.292 | -0.5195 | Yes | ||
| 34 | SLC25A14 | 24344 | 16522 | -0.316 | -0.4910 | Yes | ||
| 35 | PHF7 | 21890 | 16553 | -0.325 | -0.4584 | Yes | ||
| 36 | CSTF1 | 12515 | 16705 | -0.367 | -0.4279 | Yes | ||
| 37 | UBE2S | 17979 | 16717 | -0.370 | -0.3896 | Yes | ||
| 38 | DIABLO | 16375 | 16890 | -0.433 | -0.3533 | Yes | ||
| 39 | BANF1 | 6216 10706 3760 | 17134 | -0.523 | -0.3114 | Yes | ||
| 40 | HSPC171 | 18494 | 17168 | -0.536 | -0.2568 | Yes | ||
| 41 | SUV39H1 | 24194 | 17319 | -0.607 | -0.2010 | Yes | ||
| 42 | SNRPD1 | 23622 | 17611 | -0.767 | -0.1359 | Yes | ||
| 43 | GART | 22543 1754 | 18397 | -1.806 | 0.0118 | Yes |