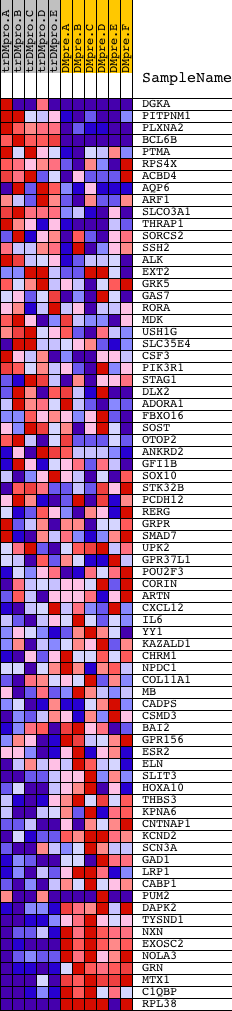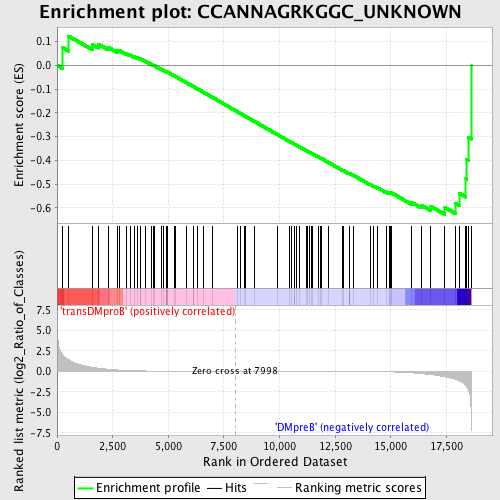
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_DMpreB.cls#transDMproB_versus_DMpreB_repos |
| Upregulated in class | DMpreB |
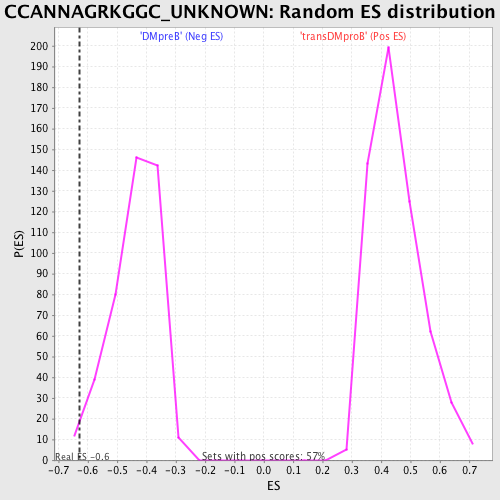
| GeneSet | CCANNAGRKGGC_UNKNOWN |
| Enrichment Score (ES) | -0.6279658 |
| Normalized Enrichment Score (NES) | -1.4312924 |
| Nominal p-value | 0.009302326 |
| FDR q-value | 0.66887456 |
| FWER p-Value | 0.983 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | DGKA | 3359 19589 | 260 | 1.980 | 0.0738 | No | ||
| 2 | PITPNM1 | 23770 | 517 | 1.419 | 0.1229 | No | ||
| 3 | PLXNA2 | 5269 | 1582 | 0.488 | 0.0871 | No | ||
| 4 | BCL6B | 20373 | 1848 | 0.373 | 0.0894 | No | ||
| 5 | PTMA | 9657 | 2295 | 0.243 | 0.0761 | No | ||
| 6 | RPS4X | 9756 | 2704 | 0.154 | 0.0610 | No | ||
| 7 | ACBD4 | 20637 | 2794 | 0.137 | 0.0622 | No | ||
| 8 | AQP6 | 22364 8621 | 3135 | 0.091 | 0.0480 | No | ||
| 9 | ARF1 | 64 | 3283 | 0.077 | 0.0434 | No | ||
| 10 | SLCO3A1 | 17796 4237 8430 | 3472 | 0.062 | 0.0360 | No | ||
| 11 | THRAP1 | 20309 | 3592 | 0.053 | 0.0320 | No | ||
| 12 | SORCS2 | 16553 | 3739 | 0.045 | 0.0261 | No | ||
| 13 | SSH2 | 6202 | 3751 | 0.044 | 0.0274 | No | ||
| 14 | ALK | 8574 | 3977 | 0.034 | 0.0168 | No | ||
| 15 | EXT2 | 14516 | 4237 | 0.025 | 0.0039 | No | ||
| 16 | GRK5 | 9036 23814 | 4329 | 0.023 | 0.0001 | No | ||
| 17 | GAS7 | 20836 1299 | 4386 | 0.022 | -0.0020 | No | ||
| 18 | RORA | 3019 5388 | 4700 | 0.015 | -0.0182 | No | ||
| 19 | MDK | 14523 | 4801 | 0.014 | -0.0230 | No | ||
| 20 | USH1G | 20153 | 4904 | 0.013 | -0.0279 | No | ||
| 21 | SLC35E4 | 20549 | 4917 | 0.013 | -0.0280 | No | ||
| 22 | CSF3 | 1394 20671 | 4924 | 0.013 | -0.0277 | No | ||
| 23 | PIK3R1 | 3170 | 4940 | 0.013 | -0.0280 | No | ||
| 24 | STAG1 | 5520 9905 2987 | 4983 | 0.012 | -0.0297 | No | ||
| 25 | DLX2 | 14561 353 | 5292 | 0.010 | -0.0459 | No | ||
| 26 | ADORA1 | 13831 | 5329 | 0.009 | -0.0474 | No | ||
| 27 | FBXO16 | 11929 | 5800 | 0.007 | -0.0725 | No | ||
| 28 | SOST | 20210 | 6131 | 0.005 | -0.0901 | No | ||
| 29 | OTOP2 | 20600 | 6144 | 0.005 | -0.0905 | No | ||
| 30 | ANKRD2 | 23853 | 6324 | 0.005 | -0.0999 | No | ||
| 31 | GFI1B | 14637 | 6593 | 0.004 | -0.1142 | No | ||
| 32 | SOX10 | 22211 | 6993 | 0.002 | -0.1356 | No | ||
| 33 | STK32B | 12255 | 8085 | -0.000 | -0.1945 | No | ||
| 34 | PCDH12 | 7005 | 8253 | -0.001 | -0.2034 | No | ||
| 35 | RERG | 16949 | 8430 | -0.001 | -0.2129 | No | ||
| 36 | GRPR | 24015 | 8457 | -0.001 | -0.2142 | No | ||
| 37 | SMAD7 | 23513 | 8883 | -0.002 | -0.2371 | No | ||
| 38 | UPK2 | 19146 | 9913 | -0.005 | -0.2924 | No | ||
| 39 | GPR37L1 | 13825 | 10459 | -0.006 | -0.3215 | No | ||
| 40 | POU2F3 | 9602 | 10532 | -0.007 | -0.3251 | No | ||
| 41 | CORIN | 16517 | 10667 | -0.007 | -0.3320 | No | ||
| 42 | ARTN | 15783 2430 2532 | 10678 | -0.007 | -0.3322 | No | ||
| 43 | CXCL12 | 9792 1182 1031 | 10758 | -0.007 | -0.3362 | No | ||
| 44 | IL6 | 16895 | 10903 | -0.008 | -0.3436 | No | ||
| 45 | YY1 | 10371 | 11211 | -0.009 | -0.3597 | No | ||
| 46 | KAZALD1 | 4206 | 11241 | -0.009 | -0.3609 | No | ||
| 47 | CHRM1 | 23943 | 11348 | -0.009 | -0.3662 | No | ||
| 48 | NPDC1 | 9479 | 11445 | -0.010 | -0.3710 | No | ||
| 49 | COL11A1 | 15442 1937 4540 8762 | 11475 | -0.010 | -0.3721 | No | ||
| 50 | MB | 22232 | 11748 | -0.011 | -0.3863 | No | ||
| 51 | CADPS | 11298 | 11843 | -0.012 | -0.3908 | No | ||
| 52 | CSMD3 | 10735 7938 | 11884 | -0.012 | -0.3924 | No | ||
| 53 | BAI2 | 2541 2379 16067 | 12214 | -0.014 | -0.4095 | No | ||
| 54 | GPR156 | 10753 22763 | 12815 | -0.019 | -0.4411 | No | ||
| 55 | ESR2 | 21038 | 12886 | -0.020 | -0.4440 | No | ||
| 56 | ELN | 8896 | 13122 | -0.022 | -0.4557 | No | ||
| 57 | SLIT3 | 20925 | 13141 | -0.022 | -0.4556 | No | ||
| 58 | HOXA10 | 17145 989 | 13144 | -0.023 | -0.4547 | No | ||
| 59 | THBS3 | 15542 1796 | 13329 | -0.025 | -0.4636 | No | ||
| 60 | KPNA6 | 4972 | 14072 | -0.038 | -0.5019 | No | ||
| 61 | CNTNAP1 | 11987 | 14234 | -0.042 | -0.5087 | No | ||
| 62 | KCND2 | 4941 17515 | 14414 | -0.047 | -0.5163 | No | ||
| 63 | SCN3A | 14573 | 14783 | -0.063 | -0.5333 | No | ||
| 64 | GAD1 | 14993 2690 | 14918 | -0.070 | -0.5374 | No | ||
| 65 | LRP1 | 9284 | 14976 | -0.073 | -0.5373 | No | ||
| 66 | CABP1 | 16412 | 15010 | -0.076 | -0.5357 | No | ||
| 67 | PUM2 | 2071 8161 | 15940 | -0.188 | -0.5774 | No | ||
| 68 | DAPK2 | 19395 | 16365 | -0.275 | -0.5881 | No | ||
| 69 | TYSND1 | 20007 | 16800 | -0.397 | -0.5939 | No | ||
| 70 | NXN | 5204 | 17432 | -0.662 | -0.5986 | Yes | ||
| 71 | EXOSC2 | 15044 | 17897 | -0.967 | -0.5807 | Yes | ||
| 72 | NOLA3 | 14914 | 18080 | -1.172 | -0.5386 | Yes | ||
| 73 | GRN | 20640 | 18377 | -1.741 | -0.4774 | Yes | ||
| 74 | MTX1 | 15282 1918 | 18414 | -1.864 | -0.3966 | Yes | ||
| 75 | C1QBP | 20364 | 18478 | -2.188 | -0.3030 | Yes | ||
| 76 | RPL38 | 12562 20606 7475 | 18614 | -7.000 | 0.0001 | Yes |