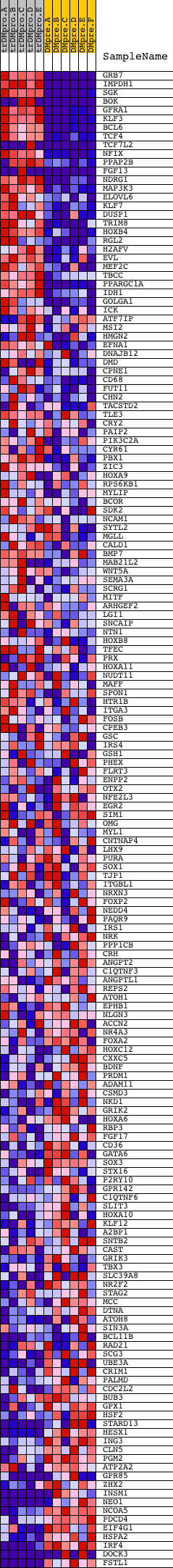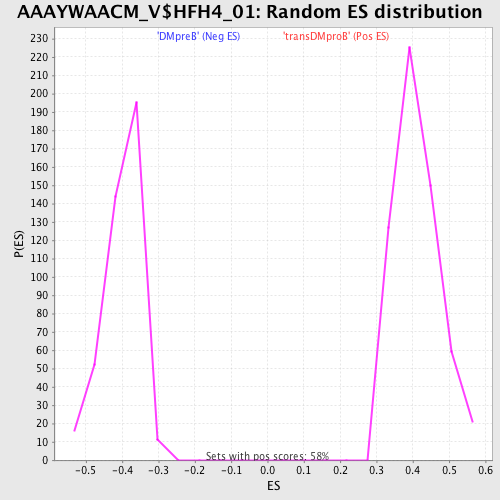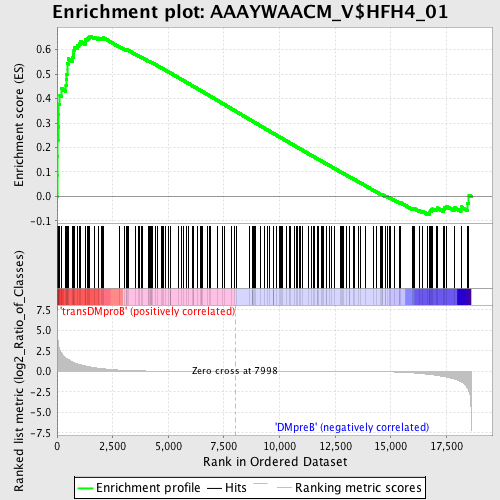
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_DMpreB.cls#transDMproB_versus_DMpreB_repos |
| Upregulated in class | transDMproB |
| GeneSet | AAAYWAACM_V$HFH4_01 |
| Enrichment Score (ES) | 0.6553381 |
| Normalized Enrichment Score (NES) | 1.5902967 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.041649014 |
| FWER p-Value | 0.289 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | GRB7 | 20673 | 5 | 5.748 | 0.0851 | Yes | ||
| 2 | IMPDH1 | 17197 1131 | 10 | 5.254 | 0.1630 | Yes | ||
| 3 | SGK | 9809 3419 | 18 | 4.634 | 0.2314 | Yes | ||
| 4 | BOK | 11953 14174 | 43 | 3.629 | 0.2840 | Yes | ||
| 5 | GFRA1 | 9015 | 56 | 3.423 | 0.3342 | Yes | ||
| 6 | KLF3 | 4961 | 82 | 2.952 | 0.3767 | Yes | ||
| 7 | BCL6 | 22624 | 126 | 2.547 | 0.4122 | Yes | ||
| 8 | TCF4 | 5642 | 203 | 2.182 | 0.4405 | Yes | ||
| 9 | TCF7L2 | 10048 5646 | 397 | 1.602 | 0.4539 | Yes | ||
| 10 | NFIX | 9458 | 409 | 1.590 | 0.4769 | Yes | ||
| 11 | PPAP2B | 7503 16161 | 425 | 1.564 | 0.4993 | Yes | ||
| 12 | FGF13 | 4720 8963 2627 | 450 | 1.532 | 0.5208 | Yes | ||
| 13 | NDRG1 | 22276 | 451 | 1.528 | 0.5435 | Yes | ||
| 14 | MAP3K3 | 20626 | 496 | 1.448 | 0.5626 | Yes | ||
| 15 | ELOVL6 | 5009 | 695 | 1.111 | 0.5684 | Yes | ||
| 16 | KLF7 | 8205 | 727 | 1.080 | 0.5827 | Yes | ||
| 17 | DUSP1 | 23061 | 755 | 1.046 | 0.5968 | Yes | ||
| 18 | TRIM8 | 23824 | 782 | 1.015 | 0.6104 | Yes | ||
| 19 | HOXB4 | 20686 | 912 | 0.898 | 0.6168 | Yes | ||
| 20 | RGL2 | 23289 | 984 | 0.843 | 0.6255 | Yes | ||
| 21 | H2AFV | 1273 20531 | 1054 | 0.795 | 0.6335 | Yes | ||
| 22 | EVL | 21156 | 1262 | 0.644 | 0.6319 | Yes | ||
| 23 | MEF2C | 3204 9378 | 1283 | 0.629 | 0.6401 | Yes | ||
| 24 | TBCC | 23215 | 1352 | 0.597 | 0.6453 | Yes | ||
| 25 | PPARGC1A | 16533 | 1402 | 0.572 | 0.6512 | Yes | ||
| 26 | IDH1 | 4894 | 1474 | 0.540 | 0.6553 | Yes | ||
| 27 | GOLGA1 | 14595 | 1676 | 0.448 | 0.6511 | No | ||
| 28 | ICK | 7135 12146 | 1843 | 0.374 | 0.6476 | No | ||
| 29 | ATF7IP | 12028 13038 | 1986 | 0.329 | 0.6448 | No | ||
| 30 | MSI2 | 8030 1268 | 2030 | 0.314 | 0.6472 | No | ||
| 31 | HMGN2 | 9095 | 2085 | 0.298 | 0.6487 | No | ||
| 32 | EFNA1 | 15279 | 2793 | 0.137 | 0.6124 | No | ||
| 33 | DNAJB12 | 7146 | 3047 | 0.100 | 0.6001 | No | ||
| 34 | DMD | 24295 2647 | 3129 | 0.092 | 0.5971 | No | ||
| 35 | CPNE1 | 11186 | 3138 | 0.091 | 0.5980 | No | ||
| 36 | CD68 | 4501 | 3144 | 0.091 | 0.5991 | No | ||
| 37 | FUT11 | 13084 | 3188 | 0.086 | 0.5981 | No | ||
| 38 | CHN2 | 7652 | 3536 | 0.057 | 0.5801 | No | ||
| 39 | TACSTD2 | 17126 | 3640 | 0.050 | 0.5752 | No | ||
| 40 | TLE3 | 5767 19416 | 3711 | 0.046 | 0.5721 | No | ||
| 41 | CRY2 | 14518 | 3798 | 0.042 | 0.5681 | No | ||
| 42 | PAIP2 | 12593 | 3816 | 0.041 | 0.5678 | No | ||
| 43 | PIK3C2A | 17667 | 4095 | 0.029 | 0.5531 | No | ||
| 44 | CYR61 | 9159 15145 9311 5024 | 4127 | 0.029 | 0.5519 | No | ||
| 45 | PBX1 | 5224 | 4148 | 0.028 | 0.5512 | No | ||
| 46 | ZIC3 | 10430 | 4200 | 0.026 | 0.5489 | No | ||
| 47 | HOXA9 | 17147 9109 1024 | 4211 | 0.026 | 0.5487 | No | ||
| 48 | RPS6KB1 | 7815 1207 13040 | 4229 | 0.026 | 0.5482 | No | ||
| 49 | MYLIP | 21652 3189 | 4242 | 0.025 | 0.5479 | No | ||
| 50 | BCOR | 12934 2561 | 4302 | 0.024 | 0.5450 | No | ||
| 51 | SDK2 | 6208 20162 | 4431 | 0.021 | 0.5384 | No | ||
| 52 | NCAM1 | 5149 | 4533 | 0.018 | 0.5332 | No | ||
| 53 | SYTL2 | 18190 1539 3730 | 4694 | 0.015 | 0.5248 | No | ||
| 54 | MGLL | 17368 1145 | 4733 | 0.015 | 0.5229 | No | ||
| 55 | CALD1 | 4273 8463 | 4763 | 0.015 | 0.5216 | No | ||
| 56 | BMP7 | 14324 | 4885 | 0.013 | 0.5152 | No | ||
| 57 | MAB21L2 | 15305 | 4985 | 0.012 | 0.5100 | No | ||
| 58 | WNT5A | 22066 5880 | 5078 | 0.011 | 0.5052 | No | ||
| 59 | SEMA3A | 16919 | 5083 | 0.011 | 0.5051 | No | ||
| 60 | SCRG1 | 18614 | 5109 | 0.011 | 0.5039 | No | ||
| 61 | MITF | 17349 | 5449 | 0.008 | 0.4857 | No | ||
| 62 | ARHGEF2 | 4987 | 5572 | 0.008 | 0.4792 | No | ||
| 63 | LGI1 | 23867 | 5681 | 0.007 | 0.4734 | No | ||
| 64 | SNCAIP | 12589 | 5818 | 0.007 | 0.4662 | No | ||
| 65 | NTN1 | 20400 | 5889 | 0.006 | 0.4625 | No | ||
| 66 | HOXB8 | 9111 | 6080 | 0.005 | 0.4522 | No | ||
| 67 | TFEC | 17213 | 6136 | 0.005 | 0.4493 | No | ||
| 68 | PRX | 3729 9628 | 6150 | 0.005 | 0.4487 | No | ||
| 69 | HOXA11 | 17146 | 6304 | 0.005 | 0.4405 | No | ||
| 70 | NUDT11 | 24387 | 6429 | 0.004 | 0.4338 | No | ||
| 71 | MAFF | 22423 2243 | 6487 | 0.004 | 0.4308 | No | ||
| 72 | SPON1 | 10600 | 6505 | 0.004 | 0.4299 | No | ||
| 73 | HTR1B | 19051 | 6550 | 0.004 | 0.4276 | No | ||
| 74 | ITGA3 | 20284 | 6741 | 0.003 | 0.4173 | No | ||
| 75 | FOSB | 17945 | 6837 | 0.003 | 0.4122 | No | ||
| 76 | CPEB3 | 5539 | 6900 | 0.003 | 0.4089 | No | ||
| 77 | GSC | 20990 | 7205 | 0.002 | 0.3925 | No | ||
| 78 | IRS4 | 9183 4926 | 7412 | 0.001 | 0.3813 | No | ||
| 79 | GSH1 | 16622 | 7535 | 0.001 | 0.3747 | No | ||
| 80 | PHEX | 24024 | 7827 | 0.000 | 0.3589 | No | ||
| 81 | FLRT3 | 7730 12930 2673 | 7954 | 0.000 | 0.3521 | No | ||
| 82 | ENPP2 | 9548 | 8048 | -0.000 | 0.3471 | No | ||
| 83 | OTX2 | 9519 | 8660 | -0.002 | 0.3140 | No | ||
| 84 | NFE2L3 | 17446 | 8770 | -0.002 | 0.3081 | No | ||
| 85 | EGR2 | 8886 | 8783 | -0.002 | 0.3075 | No | ||
| 86 | SIM1 | 20031 | 8796 | -0.002 | 0.3068 | No | ||
| 87 | OMG | 5209 | 8822 | -0.002 | 0.3055 | No | ||
| 88 | MYL1 | 4015 | 8866 | -0.002 | 0.3032 | No | ||
| 89 | CNTNAP4 | 18462 3857 | 8921 | -0.002 | 0.3003 | No | ||
| 90 | LHX9 | 9276 | 9156 | -0.003 | 0.2877 | No | ||
| 91 | PURA | 9670 | 9307 | -0.003 | 0.2796 | No | ||
| 92 | SOX1 | 18685 | 9464 | -0.004 | 0.2712 | No | ||
| 93 | TJP1 | 17803 | 9546 | -0.004 | 0.2669 | No | ||
| 94 | ITGBL1 | 5850 | 9706 | -0.004 | 0.2583 | No | ||
| 95 | NRXN3 | 2153 5196 | 9732 | -0.004 | 0.2570 | No | ||
| 96 | FOXP2 | 4312 17523 8523 | 9740 | -0.004 | 0.2567 | No | ||
| 97 | NEDD4 | 19384 3101 | 9855 | -0.005 | 0.2506 | No | ||
| 98 | PAQR9 | 19351 | 10000 | -0.005 | 0.2428 | No | ||
| 99 | IRS1 | 4925 | 10028 | -0.005 | 0.2415 | No | ||
| 100 | NRK | 6559 | 10105 | -0.005 | 0.2374 | No | ||
| 101 | PPP1CB | 5282 3529 3504 | 10127 | -0.005 | 0.2364 | No | ||
| 102 | CRH | 8784 | 10289 | -0.006 | 0.2277 | No | ||
| 103 | ANGPT2 | 14885 3783 18923 | 10425 | -0.006 | 0.2205 | No | ||
| 104 | C1QTNF3 | 2224 22505 | 10503 | -0.006 | 0.2164 | No | ||
| 105 | ANGPTL1 | 10195 8589 13064 | 10669 | -0.007 | 0.2076 | No | ||
| 106 | REPS2 | 24018 | 10738 | -0.007 | 0.2040 | No | ||
| 107 | ATOH1 | 4419 | 10818 | -0.007 | 0.1998 | No | ||
| 108 | EPHB1 | 19024 | 10882 | -0.008 | 0.1965 | No | ||
| 109 | NLGN3 | 10878 | 10956 | -0.008 | 0.1927 | No | ||
| 110 | ACCN2 | 22363 | 11008 | -0.008 | 0.1900 | No | ||
| 111 | NR4A3 | 9473 16212 5183 | 11303 | -0.009 | 0.1742 | No | ||
| 112 | FOXA2 | 14405 2889 | 11453 | -0.010 | 0.1663 | No | ||
| 113 | HOXC12 | 22343 | 11528 | -0.010 | 0.1624 | No | ||
| 114 | CXXC5 | 12523 | 11551 | -0.010 | 0.1614 | No | ||
| 115 | BDNF | 14926 2797 | 11684 | -0.011 | 0.1544 | No | ||
| 116 | PRDM1 | 19775 3337 | 11687 | -0.011 | 0.1544 | No | ||
| 117 | ADAM11 | 4340 1432 | 11764 | -0.011 | 0.1505 | No | ||
| 118 | CSMD3 | 10735 7938 | 11884 | -0.012 | 0.1442 | No | ||
| 119 | NKD1 | 8228 | 11920 | -0.012 | 0.1425 | No | ||
| 120 | GRIK2 | 3318 | 11987 | -0.013 | 0.1391 | No | ||
| 121 | HOXA6 | 17150 | 12117 | -0.013 | 0.1323 | No | ||
| 122 | RBP3 | 22047 | 12125 | -0.013 | 0.1321 | No | ||
| 123 | FGF17 | 21757 | 12265 | -0.014 | 0.1248 | No | ||
| 124 | CD36 | 8712 | 12331 | -0.015 | 0.1215 | No | ||
| 125 | GATA6 | 96 | 12472 | -0.016 | 0.1142 | No | ||
| 126 | SOX3 | 24145 | 12728 | -0.018 | 0.1006 | No | ||
| 127 | STX16 | 10464 | 12791 | -0.019 | 0.0975 | No | ||
| 128 | P2RY10 | 2613 24269 | 12813 | -0.019 | 0.0967 | No | ||
| 129 | GPR142 | 20604 | 12858 | -0.019 | 0.0946 | No | ||
| 130 | C1QTNF6 | 13063 | 13017 | -0.021 | 0.0863 | No | ||
| 131 | SLIT3 | 20925 | 13141 | -0.022 | 0.0800 | No | ||
| 132 | HOXA10 | 17145 989 | 13144 | -0.023 | 0.0802 | No | ||
| 133 | KLF12 | 4960 2637 21732 9228 | 13305 | -0.025 | 0.0719 | No | ||
| 134 | A2BP1 | 11212 1652 1689 | 13350 | -0.025 | 0.0699 | No | ||
| 135 | SNTB2 | 3887 18476 | 13531 | -0.028 | 0.0606 | No | ||
| 136 | CAST | 3182 4478 3177 | 13653 | -0.030 | 0.0544 | No | ||
| 137 | GRIK3 | 16083 | 13849 | -0.034 | 0.0444 | No | ||
| 138 | TBX3 | 16723 | 14201 | -0.041 | 0.0260 | No | ||
| 139 | SLC39A8 | 12547 1776 | 14341 | -0.045 | 0.0191 | No | ||
| 140 | NR2F2 | 8619 409 | 14533 | -0.052 | 0.0095 | No | ||
| 141 | STAG2 | 5521 | 14577 | -0.054 | 0.0080 | No | ||
| 142 | MCC | 552 | 14604 | -0.055 | 0.0074 | No | ||
| 143 | DTNA | 8870 4647 23611 1991 | 14606 | -0.055 | 0.0081 | No | ||
| 144 | ATOH8 | 17120 | 14742 | -0.061 | 0.0017 | No | ||
| 145 | SIN3A | 5442 | 14777 | -0.063 | 0.0008 | No | ||
| 146 | BCL11B | 7190 | 14847 | -0.066 | -0.0019 | No | ||
| 147 | RAD21 | 22298 | 14922 | -0.070 | -0.0049 | No | ||
| 148 | SCG3 | 19061 | 14997 | -0.075 | -0.0078 | No | ||
| 149 | UBE3A | 1209 1084 1830 | 15158 | -0.086 | -0.0152 | No | ||
| 150 | CRIM1 | 403 | 15384 | -0.108 | -0.0258 | No | ||
| 151 | PALMD | 15181 | 15393 | -0.109 | -0.0246 | No | ||
| 152 | CDC2L2 | 15964 2419 | 15414 | -0.111 | -0.0240 | No | ||
| 153 | BUB3 | 18045 | 15989 | -0.198 | -0.0522 | No | ||
| 154 | GPX1 | 19310 | 16019 | -0.202 | -0.0508 | No | ||
| 155 | HSF2 | 9128 | 16075 | -0.212 | -0.0506 | No | ||
| 156 | STARD13 | 10833 | 16291 | -0.259 | -0.0584 | No | ||
| 157 | HESX1 | 22070 | 16435 | -0.293 | -0.0618 | No | ||
| 158 | ING3 | 17514 1019 | 16657 | -0.358 | -0.0685 | No | ||
| 159 | CLN5 | 21943 | 16732 | -0.377 | -0.0669 | No | ||
| 160 | PGM2 | 16168 | 16759 | -0.387 | -0.0626 | No | ||
| 161 | ATP2A2 | 4421 3481 | 16775 | -0.391 | -0.0576 | No | ||
| 162 | GPR85 | 7227 1048 | 16840 | -0.410 | -0.0549 | No | ||
| 163 | ZHX2 | 11842 | 16891 | -0.433 | -0.0512 | No | ||
| 164 | INSM1 | 14815 | 17045 | -0.486 | -0.0523 | No | ||
| 165 | NEO1 | 5161 | 17088 | -0.506 | -0.0470 | No | ||
| 166 | NCOA5 | 14341 | 17359 | -0.623 | -0.0524 | No | ||
| 167 | PDCD4 | 5232 23816 | 17403 | -0.647 | -0.0452 | No | ||
| 168 | EIF4G1 | 22818 | 17505 | -0.698 | -0.0403 | No | ||
| 169 | HSPA2 | 21237 | 17844 | -0.934 | -0.0447 | No | ||
| 170 | IRF4 | 21679 | 18156 | -1.281 | -0.0425 | No | ||
| 171 | DOCK3 | 9920 5537 10751 | 18459 | -2.097 | -0.0278 | No | ||
| 172 | FSTL1 | 22765 | 18512 | -2.438 | 0.0056 | No |