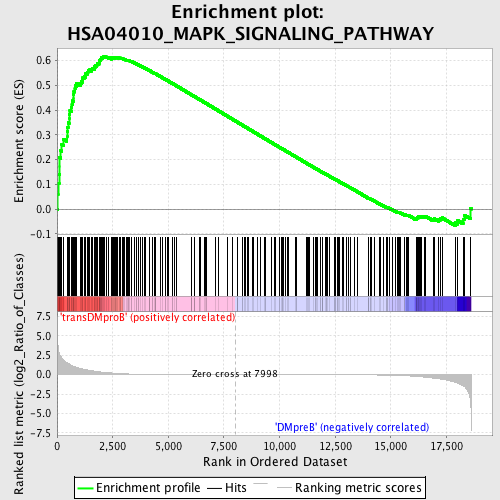
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_DMpreB.cls#transDMproB_versus_DMpreB_repos |
| Upregulated in class | transDMproB |
| GeneSet | HSA04010_MAPK_SIGNALING_PATHWAY |
| Enrichment Score (ES) | 0.6150473 |
| Normalized Enrichment Score (NES) | 1.5331838 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.15083128 |
| FWER p-Value | 0.735 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | DUSP6 | 19891 3399 | 19 | 4.580 | 0.0588 | Yes | ||
| 2 | PRKCB1 | 1693 9574 | 42 | 3.635 | 0.1051 | Yes | ||
| 3 | MAPK9 | 1233 20903 1383 | 89 | 2.817 | 0.1394 | Yes | ||
| 4 | MAPK1 | 1642 11167 | 108 | 2.667 | 0.1733 | Yes | ||
| 5 | RRAS2 | 12431 | 109 | 2.666 | 0.2081 | Yes | ||
| 6 | MAPKAPK2 | 13838 | 165 | 2.313 | 0.2354 | Yes | ||
| 7 | MAP4K2 | 6457 | 210 | 2.155 | 0.2611 | Yes | ||
| 8 | RASGRP2 | 9693 | 284 | 1.912 | 0.2821 | Yes | ||
| 9 | MAP3K7IP1 | 2193 2171 22419 | 444 | 1.544 | 0.2937 | Yes | ||
| 10 | FGF13 | 4720 8963 2627 | 450 | 1.532 | 0.3134 | Yes | ||
| 11 | TGFB1 | 18332 | 479 | 1.483 | 0.3313 | Yes | ||
| 12 | MAP3K3 | 20626 | 496 | 1.448 | 0.3493 | Yes | ||
| 13 | GADD45A | 17129 | 540 | 1.376 | 0.3649 | Yes | ||
| 14 | CACNA2D1 | 8676 | 546 | 1.366 | 0.3825 | Yes | ||
| 15 | PPM1B | 5280 1567 5281 | 575 | 1.325 | 0.3983 | Yes | ||
| 16 | MAP3K14 | 11998 | 645 | 1.197 | 0.4102 | Yes | ||
| 17 | PRKACB | 15140 | 668 | 1.157 | 0.4241 | Yes | ||
| 18 | RASA1 | 10174 | 698 | 1.106 | 0.4370 | Yes | ||
| 19 | DDIT3 | 19857 | 742 | 1.063 | 0.4486 | Yes | ||
| 20 | NFKB2 | 23810 | 744 | 1.056 | 0.4623 | Yes | ||
| 21 | DUSP1 | 23061 | 755 | 1.046 | 0.4754 | Yes | ||
| 22 | RAC2 | 22217 | 803 | 0.996 | 0.4859 | Yes | ||
| 23 | MAP3K7IP2 | 19827 | 816 | 0.979 | 0.4980 | Yes | ||
| 24 | MAPK14 | 23313 | 872 | 0.931 | 0.5072 | Yes | ||
| 25 | DUSP2 | 14865 | 1051 | 0.797 | 0.5079 | Yes | ||
| 26 | MAP3K5 | 20079 11166 | 1088 | 0.768 | 0.5160 | Yes | ||
| 27 | PTPN7 | 11500 | 1144 | 0.719 | 0.5224 | Yes | ||
| 28 | MAP4K3 | 22887 | 1158 | 0.707 | 0.5310 | Yes | ||
| 29 | RRAS | 18256 | 1216 | 0.672 | 0.5366 | Yes | ||
| 30 | MEF2C | 3204 9378 | 1283 | 0.629 | 0.5413 | Yes | ||
| 31 | MAP3K1 | 21348 | 1290 | 0.626 | 0.5491 | Yes | ||
| 32 | TP53 | 20822 | 1359 | 0.593 | 0.5532 | Yes | ||
| 33 | MKNK2 | 3299 9392 | 1390 | 0.578 | 0.5591 | Yes | ||
| 34 | RPS6KA4 | 23804 | 1464 | 0.544 | 0.5622 | Yes | ||
| 35 | STK4 | 14743 | 1551 | 0.500 | 0.5641 | Yes | ||
| 36 | TRAF6 | 5797 14940 | 1581 | 0.489 | 0.5689 | Yes | ||
| 37 | PPM1A | 5279 9607 | 1677 | 0.448 | 0.5696 | Yes | ||
| 38 | KRAS | 9247 | 1683 | 0.444 | 0.5751 | Yes | ||
| 39 | PPP3CC | 21763 | 1709 | 0.428 | 0.5793 | Yes | ||
| 40 | MAP4K4 | 11241 | 1759 | 0.408 | 0.5820 | Yes | ||
| 41 | PPP3CB | 5285 | 1794 | 0.394 | 0.5853 | Yes | ||
| 42 | CRKL | 4560 | 1800 | 0.391 | 0.5901 | Yes | ||
| 43 | SOS2 | 21049 | 1883 | 0.362 | 0.5904 | Yes | ||
| 44 | NFKB1 | 15160 | 1890 | 0.360 | 0.5948 | Yes | ||
| 45 | AKT3 | 13739 982 | 1896 | 0.357 | 0.5992 | Yes | ||
| 46 | GRB2 | 20149 | 1927 | 0.347 | 0.6021 | Yes | ||
| 47 | IL1B | 14431 | 1961 | 0.336 | 0.6047 | Yes | ||
| 48 | MAP2K3 | 20856 | 1975 | 0.333 | 0.6083 | Yes | ||
| 49 | ARRB2 | 20806 | 1981 | 0.330 | 0.6124 | Yes | ||
| 50 | JUN | 15832 | 2037 | 0.313 | 0.6135 | Yes | ||
| 51 | GNG12 | 4789 | 2082 | 0.298 | 0.6150 | Yes | ||
| 52 | RAC1 | 16302 | 2149 | 0.281 | 0.6150 | Yes | ||
| 53 | MAX | 21034 | 2221 | 0.262 | 0.6146 | No | ||
| 54 | EGF | 15169 | 2310 | 0.241 | 0.6130 | No | ||
| 55 | MAPK3 | 6458 11170 | 2440 | 0.214 | 0.6087 | No | ||
| 56 | MAP3K4 | 23126 | 2446 | 0.212 | 0.6112 | No | ||
| 57 | TRAF2 | 14657 | 2502 | 0.200 | 0.6109 | No | ||
| 58 | RASA2 | 4329 | 2532 | 0.194 | 0.6118 | No | ||
| 59 | CACNB2 | 4466 8678 | 2590 | 0.180 | 0.6111 | No | ||
| 60 | MAP3K8 | 23495 | 2611 | 0.176 | 0.6123 | No | ||
| 61 | RAP1B | 10083 | 2625 | 0.172 | 0.6138 | No | ||
| 62 | SRF | 22961 1597 | 2654 | 0.165 | 0.6145 | No | ||
| 63 | RPS6KA1 | 15725 | 2699 | 0.155 | 0.6141 | No | ||
| 64 | FOS | 21202 | 2816 | 0.135 | 0.6095 | No | ||
| 65 | PAK1 | 9527 | 2858 | 0.127 | 0.6090 | No | ||
| 66 | CD14 | 23452 | 2861 | 0.126 | 0.6105 | No | ||
| 67 | ACVR1B | 4335 22353 | 2930 | 0.114 | 0.6083 | No | ||
| 68 | MAP3K7 | 16255 | 3003 | 0.105 | 0.6058 | No | ||
| 69 | RAF1 | 17035 | 3038 | 0.101 | 0.6052 | No | ||
| 70 | MYC | 22465 9435 | 3120 | 0.093 | 0.6020 | No | ||
| 71 | MAP2K1 | 19082 | 3174 | 0.087 | 0.6003 | No | ||
| 72 | MAPK13 | 23314 | 3189 | 0.086 | 0.6007 | No | ||
| 73 | NF1 | 5165 | 3219 | 0.082 | 0.6001 | No | ||
| 74 | FGFR2 | 1917 4722 1119 | 3240 | 0.081 | 0.6001 | No | ||
| 75 | PDGFRB | 23539 | 3339 | 0.072 | 0.5957 | No | ||
| 76 | RASGRP4 | 6119 | 3340 | 0.072 | 0.5967 | No | ||
| 77 | MAPK8IP3 | 11402 23080 | 3458 | 0.062 | 0.5911 | No | ||
| 78 | CDC25B | 14841 | 3547 | 0.056 | 0.5870 | No | ||
| 79 | MAP2K5 | 19088 | 3584 | 0.054 | 0.5858 | No | ||
| 80 | GNA12 | 9023 | 3653 | 0.049 | 0.5827 | No | ||
| 81 | DUSP7 | 10661 | 3679 | 0.048 | 0.5820 | No | ||
| 82 | RASGRP1 | 5360 14476 9694 | 3767 | 0.043 | 0.5778 | No | ||
| 83 | PDGFA | 5237 | 3830 | 0.040 | 0.5750 | No | ||
| 84 | MAP3K10 | 3831 17916 | 3932 | 0.036 | 0.5700 | No | ||
| 85 | CASP3 | 8693 | 3967 | 0.034 | 0.5685 | No | ||
| 86 | FAS | 23881 3750 | 4136 | 0.028 | 0.5598 | No | ||
| 87 | ARRB1 | 3890 8465 18178 4276 | 4151 | 0.028 | 0.5594 | No | ||
| 88 | DUSP4 | 18632 3820 | 4155 | 0.028 | 0.5596 | No | ||
| 89 | MAPK7 | 1381 20414 | 4301 | 0.024 | 0.5520 | No | ||
| 90 | STK3 | 7084 | 4305 | 0.024 | 0.5522 | No | ||
| 91 | FGF18 | 4721 1351 | 4370 | 0.022 | 0.5490 | No | ||
| 92 | TNF | 23004 | 4411 | 0.021 | 0.5471 | No | ||
| 93 | DAXX | 23290 | 4418 | 0.021 | 0.5470 | No | ||
| 94 | CACNA1E | 4465 | 4441 | 0.020 | 0.5461 | No | ||
| 95 | MAP3K6 | 16054 | 4644 | 0.016 | 0.5353 | No | ||
| 96 | SOS1 | 5476 | 4722 | 0.015 | 0.5313 | No | ||
| 97 | ACVR1C | 14585 | 4862 | 0.013 | 0.5239 | No | ||
| 98 | CRK | 4559 1249 | 4892 | 0.013 | 0.5225 | No | ||
| 99 | CACNA1F | 12054 24393 | 4942 | 0.013 | 0.5200 | No | ||
| 100 | RAC3 | 20561 | 4987 | 0.012 | 0.5178 | No | ||
| 101 | PLA2G2D | 16018 | 5193 | 0.010 | 0.5068 | No | ||
| 102 | CDC42 | 4503 8722 4504 2465 | 5208 | 0.010 | 0.5061 | No | ||
| 103 | PPP3R1 | 9611 489 | 5297 | 0.010 | 0.5015 | No | ||
| 104 | TGFBR1 | 16213 10165 5745 | 5364 | 0.009 | 0.4980 | No | ||
| 105 | FGF4 | 8964 | 5376 | 0.009 | 0.4975 | No | ||
| 106 | PPP3R2 | 9612 | 6019 | 0.006 | 0.4627 | No | ||
| 107 | NR4A1 | 9099 | 6022 | 0.006 | 0.4626 | No | ||
| 108 | MAP3K2 | 11165 | 6028 | 0.006 | 0.4624 | No | ||
| 109 | DUSP9 | 24307 | 6164 | 0.005 | 0.4551 | No | ||
| 110 | FGF21 | 17830 | 6170 | 0.005 | 0.4549 | No | ||
| 111 | FGF1 | 1994 23447 | 6176 | 0.005 | 0.4547 | No | ||
| 112 | DUSP16 | 1004 7699 | 6391 | 0.004 | 0.4431 | No | ||
| 113 | CACNA1D | 4464 8675 | 6392 | 0.004 | 0.4432 | No | ||
| 114 | IL1R2 | 14265 | 6450 | 0.004 | 0.4402 | No | ||
| 115 | PDGFB | 22199 2311 | 6629 | 0.003 | 0.4305 | No | ||
| 116 | PLA2G12B | 20016 | 6687 | 0.003 | 0.4275 | No | ||
| 117 | PLA2G1B | 9581 | 6716 | 0.003 | 0.4260 | No | ||
| 118 | FGF16 | 24270 | 6721 | 0.003 | 0.4258 | No | ||
| 119 | CACNA1H | 23077 | 7106 | 0.002 | 0.4049 | No | ||
| 120 | PRKX | 9619 5290 | 7254 | 0.002 | 0.3970 | No | ||
| 121 | FASLG | 13789 | 7673 | 0.001 | 0.3742 | No | ||
| 122 | FGF7 | 8965 | 7875 | 0.000 | 0.3633 | No | ||
| 123 | FGF14 | 21716 3005 3463 | 8086 | -0.000 | 0.3519 | No | ||
| 124 | FGF5 | 16785 | 8104 | -0.000 | 0.3509 | No | ||
| 125 | FGFR1 | 3789 8968 | 8129 | -0.000 | 0.3496 | No | ||
| 126 | NTF5 | 18248 | 8312 | -0.001 | 0.3397 | No | ||
| 127 | PLA2G3 | 20968 | 8330 | -0.001 | 0.3388 | No | ||
| 128 | CACNG3 | 12029 | 8441 | -0.001 | 0.3329 | No | ||
| 129 | RPS6KA2 | 9759 9758 | 8473 | -0.001 | 0.3312 | No | ||
| 130 | RPS6KA5 | 2076 21005 | 8560 | -0.001 | 0.3265 | No | ||
| 131 | MRAS | 9409 | 8608 | -0.001 | 0.3240 | No | ||
| 132 | NTRK2 | 9491 | 8794 | -0.002 | 0.3139 | No | ||
| 133 | CACNG7 | 13497 | 8816 | -0.002 | 0.3128 | No | ||
| 134 | PLA2G5 | 9583 | 9022 | -0.002 | 0.3017 | No | ||
| 135 | CACNG6 | 7033 | 9161 | -0.003 | 0.2942 | No | ||
| 136 | MOS | 16254 | 9303 | -0.003 | 0.2866 | No | ||
| 137 | CACNG5 | 1339 20175 | 9354 | -0.003 | 0.2839 | No | ||
| 138 | NFATC2 | 5168 2866 | 9388 | -0.003 | 0.2822 | No | ||
| 139 | MAPKAPK5 | 16387 | 9650 | -0.004 | 0.2680 | No | ||
| 140 | RPS6KA6 | 12458 | 9781 | -0.004 | 0.2610 | No | ||
| 141 | FGF8 | 23656 | 9836 | -0.005 | 0.2581 | No | ||
| 142 | TAOK2 | 10610 3794 | 9984 | -0.005 | 0.2502 | No | ||
| 143 | FGF12 | 1723 22621 | 9991 | -0.005 | 0.2499 | No | ||
| 144 | AKT2 | 4365 4366 | 10087 | -0.005 | 0.2448 | No | ||
| 145 | PLA2G2E | 16016 | 10108 | -0.005 | 0.2438 | No | ||
| 146 | CACNG4 | 20176 | 10149 | -0.005 | 0.2417 | No | ||
| 147 | CACNG8 | 13498 | 10182 | -0.006 | 0.2400 | No | ||
| 148 | RAP1A | 8467 | 10259 | -0.006 | 0.2360 | No | ||
| 149 | IKBKG | 2570 2562 4908 | 10346 | -0.006 | 0.2314 | No | ||
| 150 | EVI1 | 4689 8926 | 10409 | -0.006 | 0.2281 | No | ||
| 151 | FGF11 | 20383 | 10692 | -0.007 | 0.2128 | No | ||
| 152 | FGF10 | 4719 8962 | 10747 | -0.007 | 0.2100 | No | ||
| 153 | CACNB3 | 22373 8679 | 10777 | -0.007 | 0.2085 | No | ||
| 154 | FLNC | 4302 8510 | 11215 | -0.009 | 0.1849 | No | ||
| 155 | CACNA1A | 4461 | 11243 | -0.009 | 0.1835 | No | ||
| 156 | PLA2G10 | 22658 | 11321 | -0.009 | 0.1794 | No | ||
| 157 | NFATC4 | 22002 | 11335 | -0.009 | 0.1788 | No | ||
| 158 | MAP2K7 | 6453 | 11501 | -0.010 | 0.1700 | No | ||
| 159 | PLA2G4A | 13809 | 11620 | -0.011 | 0.1637 | No | ||
| 160 | PLA2G2A | 16017 | 11629 | -0.011 | 0.1634 | No | ||
| 161 | PRKCA | 20174 | 11674 | -0.011 | 0.1612 | No | ||
| 162 | BDNF | 14926 2797 | 11684 | -0.011 | 0.1608 | No | ||
| 163 | FGF20 | 18892 | 11825 | -0.012 | 0.1534 | No | ||
| 164 | CACNA1B | 8673 4462 | 11859 | -0.012 | 0.1517 | No | ||
| 165 | MAP3K13 | 22814 | 11915 | -0.012 | 0.1489 | No | ||
| 166 | PDGFRA | 16824 | 11924 | -0.012 | 0.1486 | No | ||
| 167 | PRKACA | 18549 3844 | 11943 | -0.012 | 0.1478 | No | ||
| 168 | FGFR3 | 8969 3566 | 12054 | -0.013 | 0.1420 | No | ||
| 169 | PTPRR | 3436 19876 | 12101 | -0.013 | 0.1396 | No | ||
| 170 | IL1A | 4915 | 12109 | -0.013 | 0.1394 | No | ||
| 171 | NTF3 | 16991 | 12165 | -0.014 | 0.1366 | No | ||
| 172 | FGF17 | 21757 | 12265 | -0.014 | 0.1314 | No | ||
| 173 | DUSP10 | 4003 14016 | 12461 | -0.016 | 0.1210 | No | ||
| 174 | FGF22 | 12468 7385 | 12479 | -0.016 | 0.1203 | No | ||
| 175 | PLA2G6 | 22209 | 12521 | -0.016 | 0.1183 | No | ||
| 176 | FGF9 | 8966 | 12523 | -0.016 | 0.1185 | No | ||
| 177 | EGFR | 1329 20944 | 12610 | -0.017 | 0.1140 | No | ||
| 178 | CACNA1C | 1158 1166 24464 4463 8674 | 12619 | -0.017 | 0.1138 | No | ||
| 179 | CACNB4 | 4467 | 12626 | -0.017 | 0.1137 | No | ||
| 180 | FGFR4 | 3226 | 12709 | -0.018 | 0.1095 | No | ||
| 181 | ELK4 | 8895 4664 3937 | 12836 | -0.019 | 0.1029 | No | ||
| 182 | MAPK8IP2 | 77 | 12876 | -0.020 | 0.1010 | No | ||
| 183 | FGF2 | 15608 | 13003 | -0.021 | 0.0944 | No | ||
| 184 | TGFBR2 | 2994 3001 10167 | 13012 | -0.021 | 0.0943 | No | ||
| 185 | NGFB | 15476 | 13093 | -0.022 | 0.0902 | No | ||
| 186 | CACNA1G | 1237 940 20292 | 13112 | -0.022 | 0.0895 | No | ||
| 187 | TGFB3 | 10161 | 13172 | -0.023 | 0.0866 | No | ||
| 188 | AKT1 | 8568 | 13176 | -0.023 | 0.0867 | No | ||
| 189 | IL1R1 | 14264 3987 | 13385 | -0.026 | 0.0757 | No | ||
| 190 | MAPK10 | 11169 | 13498 | -0.027 | 0.0700 | No | ||
| 191 | DUSP14 | 12124 | 13996 | -0.036 | 0.0434 | No | ||
| 192 | MAP2K1IP1 | 12158 | 13997 | -0.036 | 0.0439 | No | ||
| 193 | NTRK1 | 15299 | 14008 | -0.037 | 0.0438 | No | ||
| 194 | MAPK11 | 2264 9618 22163 | 14100 | -0.039 | 0.0394 | No | ||
| 195 | MAPT | 9420 1347 | 14103 | -0.039 | 0.0398 | No | ||
| 196 | TNFRSF1A | 1181 10206 | 14144 | -0.040 | 0.0381 | No | ||
| 197 | RASGRF1 | 3050 19357 | 14265 | -0.043 | 0.0322 | No | ||
| 198 | PLA2G2F | 15700 | 14489 | -0.050 | 0.0207 | No | ||
| 199 | NLK | 5179 5178 | 14530 | -0.051 | 0.0192 | No | ||
| 200 | FGF3 | 17997 | 14667 | -0.058 | 0.0125 | No | ||
| 201 | CACNG1 | 20177 | 14800 | -0.064 | 0.0062 | No | ||
| 202 | FGF23 | 17275 | 14821 | -0.065 | 0.0060 | No | ||
| 203 | DUSP3 | 1358 13022 | 14853 | -0.067 | 0.0051 | No | ||
| 204 | RASGRF2 | 21397 | 14857 | -0.067 | 0.0059 | No | ||
| 205 | RPS6KA3 | 8490 | 14937 | -0.071 | 0.0025 | No | ||
| 206 | DUSP8 | 9493 | 15055 | -0.079 | -0.0028 | No | ||
| 207 | CACNA2D2 | 7154 | 15056 | -0.079 | -0.0018 | No | ||
| 208 | MAP3K12 | 6454 11164 | 15194 | -0.089 | -0.0081 | No | ||
| 209 | ATF2 | 4418 2759 | 15321 | -0.100 | -0.0137 | No | ||
| 210 | CACNB1 | 1443 1304 | 15347 | -0.103 | -0.0137 | No | ||
| 211 | MAP2K6 | 20614 1414 | 15369 | -0.106 | -0.0134 | No | ||
| 212 | CACNA2D3 | 8677 | 15449 | -0.114 | -0.0162 | No | ||
| 213 | MAP2K4 | 20405 | 15611 | -0.134 | -0.0233 | No | ||
| 214 | MKNK1 | 2504 | 15624 | -0.136 | -0.0221 | No | ||
| 215 | MAP2K2 | 19933 | 15726 | -0.151 | -0.0256 | No | ||
| 216 | PTPN5 | 17817 | 15755 | -0.154 | -0.0252 | No | ||
| 217 | GADD45G | 21637 | 15775 | -0.157 | -0.0241 | No | ||
| 218 | PAK2 | 10310 5875 | 16141 | -0.223 | -0.0411 | No | ||
| 219 | PPP5C | 17952 | 16156 | -0.226 | -0.0389 | No | ||
| 220 | STMN1 | 9261 | 16175 | -0.230 | -0.0369 | No | ||
| 221 | CACNA1S | 14111 | 16179 | -0.231 | -0.0340 | No | ||
| 222 | NRAS | 5191 | 16222 | -0.243 | -0.0331 | No | ||
| 223 | MAPK8 | 6459 | 16226 | -0.245 | -0.0301 | No | ||
| 224 | IKBKB | 4907 | 16287 | -0.259 | -0.0300 | No | ||
| 225 | FLNA | 24129 | 16355 | -0.272 | -0.0300 | No | ||
| 226 | RASGRP3 | 10767 | 16399 | -0.284 | -0.0287 | No | ||
| 227 | GADD45B | 19935 | 16500 | -0.310 | -0.0301 | No | ||
| 228 | HRAS | 4868 | 16569 | -0.329 | -0.0295 | No | ||
| 229 | FLNB | 13742 | 16904 | -0.436 | -0.0419 | No | ||
| 230 | MAPKAPK3 | 19003 | 16967 | -0.461 | -0.0393 | No | ||
| 231 | MAPK12 | 2182 2215 22164 | 17159 | -0.534 | -0.0427 | No | ||
| 232 | CHUK | 23665 | 17214 | -0.556 | -0.0384 | No | ||
| 233 | ATF4 | 22417 2239 | 17300 | -0.600 | -0.0352 | No | ||
| 234 | TGFB2 | 5743 | 17912 | -0.982 | -0.0556 | No | ||
| 235 | MAP4K1 | 18313 | 18009 | -1.088 | -0.0466 | No | ||
| 236 | PPP3CA | 1863 5284 | 18249 | -1.415 | -0.0411 | No | ||
| 237 | ECSIT | 11248 | 18302 | -1.518 | -0.0241 | No | ||
| 238 | PLA2G12A | 1782 7281 1839 | 18573 | -3.146 | 0.0023 | No |

