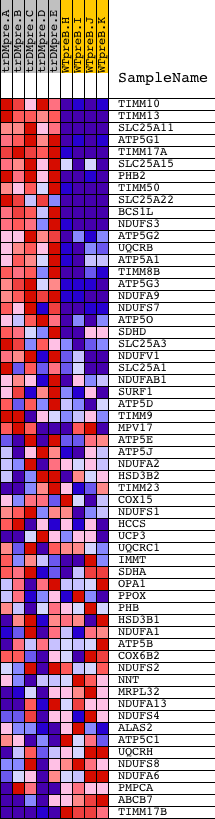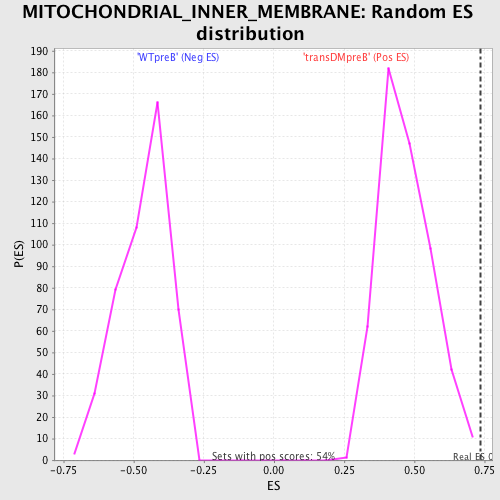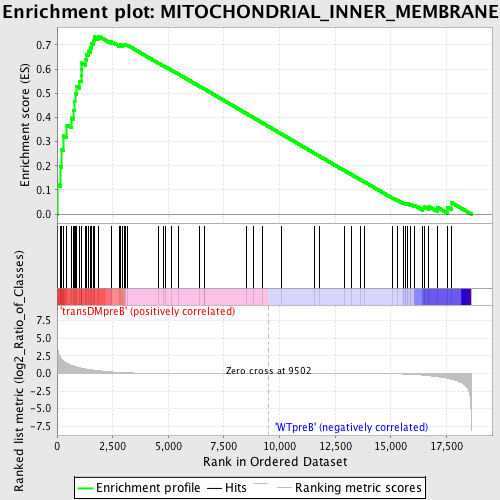
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_WTpreB.phenotype_transDMpreB_versus_WTpreB.cls #transDMpreB_versus_WTpreB |
| Phenotype | phenotype_transDMpreB_versus_WTpreB.cls#transDMpreB_versus_WTpreB |
| Upregulated in class | transDMpreB |
| GeneSet | MITOCHONDRIAL_INNER_MEMBRANE |
| Enrichment Score (ES) | 0.73659974 |
| Normalized Enrichment Score (NES) | 1.5724602 |
| Nominal p-value | 0.0036832413 |
| FDR q-value | 0.07775739 |
| FWER p-Value | 0.61 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | TIMM10 | 11390 2158 | 30 | 3.408 | 0.1220 | Yes | ||
| 2 | TIMM13 | 14974 | 149 | 2.246 | 0.1970 | Yes | ||
| 3 | SLC25A11 | 20366 1368 | 203 | 2.002 | 0.2668 | Yes | ||
| 4 | ATP5G1 | 8636 | 288 | 1.688 | 0.3235 | Yes | ||
| 5 | TIMM17A | 13822 | 434 | 1.463 | 0.3687 | Yes | ||
| 6 | SLC25A15 | 356 3819 | 656 | 1.131 | 0.3978 | Yes | ||
| 7 | PHB2 | 17286 | 747 | 1.032 | 0.4304 | Yes | ||
| 8 | TIMM50 | 17911 | 768 | 1.015 | 0.4661 | Yes | ||
| 9 | SLC25A22 | 12671 | 805 | 0.976 | 0.4995 | Yes | ||
| 10 | BCS1L | 14223 | 887 | 0.904 | 0.5280 | Yes | ||
| 11 | NDUFS3 | 7553 12683 | 1010 | 0.800 | 0.5504 | Yes | ||
| 12 | ATP5G2 | 12610 | 1079 | 0.751 | 0.5740 | Yes | ||
| 13 | UQCRB | 12545 | 1095 | 0.737 | 0.5999 | Yes | ||
| 14 | ATP5A1 | 23505 | 1111 | 0.728 | 0.6255 | Yes | ||
| 15 | TIMM8B | 11391 | 1293 | 0.625 | 0.6384 | Yes | ||
| 16 | ATP5G3 | 14558 | 1321 | 0.611 | 0.6591 | Yes | ||
| 17 | NDUFA9 | 12288 | 1407 | 0.566 | 0.6750 | Yes | ||
| 18 | NDUFS7 | 19946 | 1495 | 0.532 | 0.6896 | Yes | ||
| 19 | ATP5O | 22539 | 1544 | 0.516 | 0.7057 | Yes | ||
| 20 | SDHD | 19125 | 1650 | 0.470 | 0.7171 | Yes | ||
| 21 | SLC25A3 | 19644 | 1665 | 0.464 | 0.7332 | Yes | ||
| 22 | NDUFV1 | 23953 | 1865 | 0.389 | 0.7366 | Yes | ||
| 23 | SLC25A1 | 22645 | 2447 | 0.212 | 0.7130 | No | ||
| 24 | NDUFAB1 | 7667 | 2801 | 0.139 | 0.6990 | No | ||
| 25 | SURF1 | 14645 | 2860 | 0.127 | 0.7005 | No | ||
| 26 | ATP5D | 19949 | 2957 | 0.113 | 0.6994 | No | ||
| 27 | TIMM9 | 19686 | 3007 | 0.107 | 0.7006 | No | ||
| 28 | MPV17 | 9407 3505 3543 | 3066 | 0.097 | 0.7010 | No | ||
| 29 | ATP5E | 14321 | 3159 | 0.087 | 0.6992 | No | ||
| 30 | ATP5J | 880 22555 | 4557 | 0.025 | 0.6248 | No | ||
| 31 | NDUFA2 | 9450 | 4802 | 0.021 | 0.6125 | No | ||
| 32 | HSD3B2 | 4873 4874 15229 4875 9126 | 4862 | 0.021 | 0.6100 | No | ||
| 33 | TIMM23 | 7004 5768 | 5158 | 0.017 | 0.5947 | No | ||
| 34 | COX15 | 5931 | 5470 | 0.014 | 0.5785 | No | ||
| 35 | NDUFS1 | 13940 | 6418 | 0.009 | 0.5278 | No | ||
| 36 | HCCS | 9079 4842 | 6631 | 0.008 | 0.5167 | No | ||
| 37 | UCP3 | 3972 18175 | 6636 | 0.008 | 0.5167 | No | ||
| 38 | UQCRC1 | 10259 | 8510 | 0.002 | 0.4159 | No | ||
| 39 | IMMT | 367 8029 | 8809 | 0.002 | 0.3999 | No | ||
| 40 | SDHA | 12436 | 9232 | 0.001 | 0.3772 | No | ||
| 41 | OPA1 | 13169 22800 7894 | 10063 | -0.001 | 0.3325 | No | ||
| 42 | PPOX | 13761 4104 | 11558 | -0.005 | 0.2522 | No | ||
| 43 | PHB | 9558 | 11778 | -0.006 | 0.2406 | No | ||
| 44 | HSD3B1 | 15227 | 12902 | -0.011 | 0.1805 | No | ||
| 45 | NDUFA1 | 24170 | 13236 | -0.013 | 0.1631 | No | ||
| 46 | ATP5B | 19846 | 13623 | -0.017 | 0.1429 | No | ||
| 47 | COX6B2 | 17983 4110 | 13808 | -0.019 | 0.1336 | No | ||
| 48 | NDUFS2 | 4089 13760 | 15088 | -0.051 | 0.0666 | No | ||
| 49 | NNT | 3273 5181 9471 3238 | 15316 | -0.063 | 0.0566 | No | ||
| 50 | MRPL32 | 7961 | 15580 | -0.085 | 0.0455 | No | ||
| 51 | NDUFA13 | 7400 | 15666 | -0.092 | 0.0442 | No | ||
| 52 | NDUFS4 | 9452 5157 3173 | 15760 | -0.102 | 0.0429 | No | ||
| 53 | ALAS2 | 24229 | 15878 | -0.117 | 0.0409 | No | ||
| 54 | ATP5C1 | 8635 | 16066 | -0.152 | 0.0363 | No | ||
| 55 | UQCRH | 12378 | 16440 | -0.237 | 0.0248 | No | ||
| 56 | NDUFS8 | 23950 | 16507 | -0.260 | 0.0307 | No | ||
| 57 | NDUFA6 | 12473 | 16709 | -0.319 | 0.0314 | No | ||
| 58 | PMPCA | 12421 7350 | 17106 | -0.473 | 0.0273 | No | ||
| 59 | ABCB7 | 24085 | 17557 | -0.692 | 0.0281 | No | ||
| 60 | TIMM17B | 24390 | 17712 | -0.797 | 0.0487 | No |