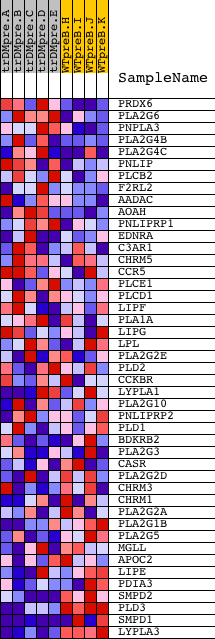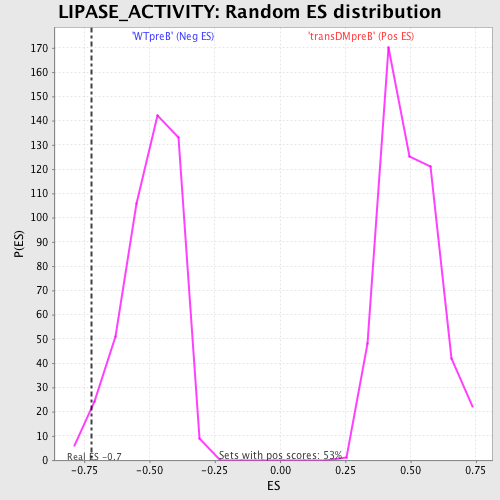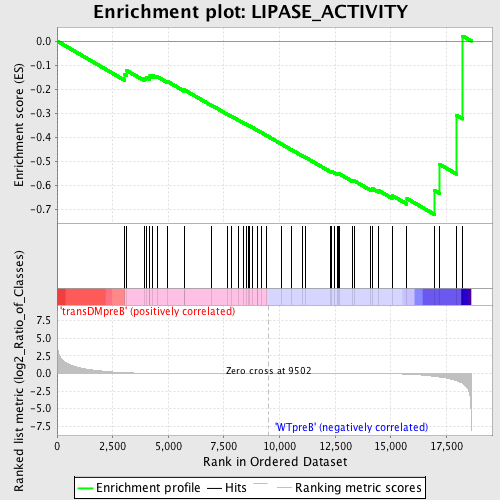
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_WTpreB.phenotype_transDMpreB_versus_WTpreB.cls #transDMpreB_versus_WTpreB |
| Phenotype | phenotype_transDMpreB_versus_WTpreB.cls#transDMpreB_versus_WTpreB |
| Upregulated in class | WTpreB |
| GeneSet | LIPASE_ACTIVITY |
| Enrichment Score (ES) | -0.7216056 |
| Normalized Enrichment Score (NES) | -1.466413 |
| Nominal p-value | 0.02972399 |
| FDR q-value | 0.5062587 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PRDX6 | 4389 8596 | 3041 | 0.101 | -0.1396 | No | ||
| 2 | PLA2G6 | 22209 | 3130 | 0.090 | -0.1227 | No | ||
| 3 | PNPLA3 | 22406 | 3910 | 0.041 | -0.1548 | No | ||
| 4 | PLA2G4B | 14896 | 3997 | 0.038 | -0.1503 | No | ||
| 5 | PLA2G4C | 147 | 4166 | 0.033 | -0.1514 | No | ||
| 6 | PNLIP | 23830 | 4173 | 0.033 | -0.1439 | No | ||
| 7 | PLCB2 | 5262 | 4304 | 0.030 | -0.1438 | No | ||
| 8 | F2RL2 | 21580 | 4490 | 0.026 | -0.1475 | No | ||
| 9 | AADAC | 15584 | 4963 | 0.019 | -0.1683 | No | ||
| 10 | AOAH | 11296 | 5712 | 0.012 | -0.2056 | No | ||
| 11 | PNLIPRP1 | 23835 | 5719 | 0.012 | -0.2029 | No | ||
| 12 | EDNRA | 18834 | 6961 | 0.007 | -0.2681 | No | ||
| 13 | C3AR1 | 8668 | 7644 | 0.005 | -0.3037 | No | ||
| 14 | CHRM5 | 10038 | 7860 | 0.004 | -0.3143 | No | ||
| 15 | CCR5 | 8754 4534 | 8141 | 0.003 | -0.3285 | No | ||
| 16 | PLCE1 | 13156 | 8377 | 0.003 | -0.3405 | No | ||
| 17 | PLCD1 | 18972 | 8511 | 0.002 | -0.3471 | No | ||
| 18 | LIPF | 23883 | 8597 | 0.002 | -0.3511 | No | ||
| 19 | PLA1A | 1737 22594 | 8656 | 0.002 | -0.3538 | No | ||
| 20 | LIPG | 23407 | 8800 | 0.002 | -0.3611 | No | ||
| 21 | LPL | 9283 5003 | 8994 | 0.001 | -0.3712 | No | ||
| 22 | PLA2G2E | 16016 | 9015 | 0.001 | -0.3719 | No | ||
| 23 | PLD2 | 20803 | 9178 | 0.001 | -0.3805 | No | ||
| 24 | CCKBR | 8706 | 9431 | 0.000 | -0.3940 | No | ||
| 25 | LYPLA1 | 4107 9580 | 10085 | -0.001 | -0.4288 | No | ||
| 26 | PLA2G10 | 22658 | 10543 | -0.003 | -0.4528 | No | ||
| 27 | PNLIPRP2 | 23836 | 11039 | -0.004 | -0.4785 | No | ||
| 28 | PLD1 | 15624 | 11167 | -0.004 | -0.4843 | No | ||
| 29 | BDKRB2 | 21163 | 12295 | -0.008 | -0.5430 | No | ||
| 30 | PLA2G3 | 20968 | 12336 | -0.008 | -0.5431 | No | ||
| 31 | CASR | 22602 | 12457 | -0.009 | -0.5474 | No | ||
| 32 | PLA2G2D | 16018 | 12613 | -0.010 | -0.5535 | No | ||
| 33 | CHRM3 | 21547 | 12665 | -0.010 | -0.5538 | No | ||
| 34 | CHRM1 | 23943 | 12673 | -0.010 | -0.5517 | No | ||
| 35 | PLA2G2A | 16017 | 13294 | -0.014 | -0.5818 | No | ||
| 36 | PLA2G1B | 9581 | 13366 | -0.015 | -0.5821 | No | ||
| 37 | PLA2G5 | 9583 | 14095 | -0.023 | -0.6158 | No | ||
| 38 | MGLL | 17368 1145 | 14183 | -0.025 | -0.6146 | No | ||
| 39 | APOC2 | 8616 | 14438 | -0.029 | -0.6212 | No | ||
| 40 | LIPE | 17927 | 15060 | -0.050 | -0.6428 | No | ||
| 41 | PDIA3 | 14888 | 15724 | -0.097 | -0.6552 | Yes | ||
| 42 | SMPD2 | 5462 | 16959 | -0.418 | -0.6213 | Yes | ||
| 43 | PLD3 | 17917 | 17197 | -0.505 | -0.5129 | Yes | ||
| 44 | SMPD1 | 18145 2658 | 17955 | -1.019 | -0.3093 | Yes | ||
| 45 | LYPLA3 | 18482 | 18236 | -1.438 | 0.0205 | Yes |