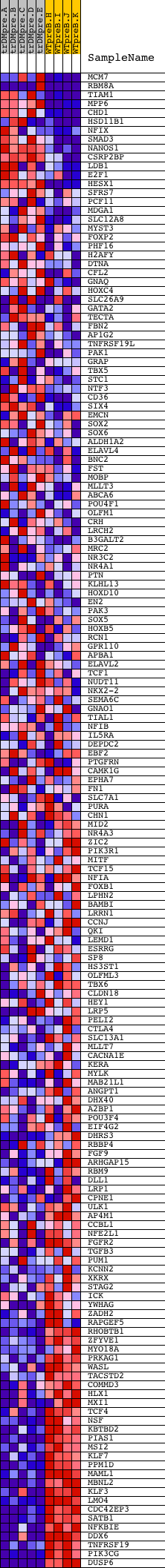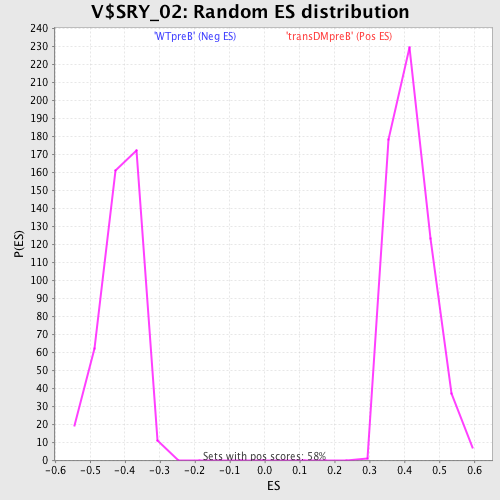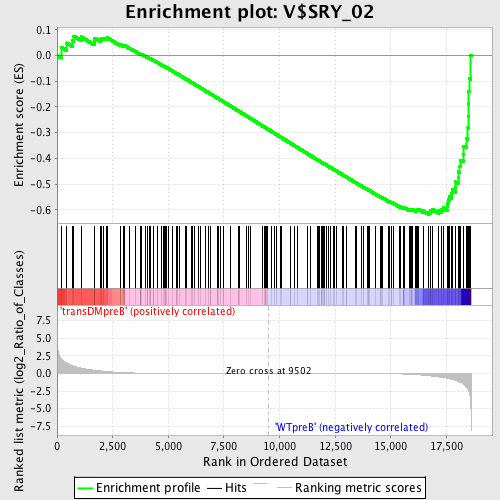
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_WTpreB.phenotype_transDMpreB_versus_WTpreB.cls #transDMpreB_versus_WTpreB |
| Phenotype | phenotype_transDMpreB_versus_WTpreB.cls#transDMpreB_versus_WTpreB |
| Upregulated in class | WTpreB |
| GeneSet | V$SRY_02 |
| Enrichment Score (ES) | -0.61817044 |
| Normalized Enrichment Score (NES) | -1.5031215 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.15553841 |
| FWER p-Value | 0.783 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | MCM7 | 9372 3568 | 178 | 2.099 | 0.0332 | No | ||
| 2 | RBM8A | 12239 7212 | 441 | 1.459 | 0.0488 | No | ||
| 3 | TIAM1 | 22547 | 687 | 1.091 | 0.0578 | No | ||
| 4 | MPP6 | 17448 | 749 | 1.030 | 0.0756 | No | ||
| 5 | CHD1 | 23377 | 1087 | 0.744 | 0.0725 | No | ||
| 6 | HSD11B1 | 9125 4019 | 1667 | 0.464 | 0.0506 | No | ||
| 7 | NFIX | 9458 | 1690 | 0.456 | 0.0587 | No | ||
| 8 | SMAD3 | 19084 | 1695 | 0.455 | 0.0678 | No | ||
| 9 | NANOS1 | 6858 | 1948 | 0.360 | 0.0615 | No | ||
| 10 | CSRP2BP | 2696 10458 | 2011 | 0.340 | 0.0651 | No | ||
| 11 | LDB1 | 23654 | 2084 | 0.318 | 0.0676 | No | ||
| 12 | E2F1 | 14384 | 2208 | 0.278 | 0.0667 | No | ||
| 13 | HESX1 | 22070 | 2270 | 0.261 | 0.0687 | No | ||
| 14 | SFRS7 | 22889 | 2839 | 0.130 | 0.0406 | No | ||
| 15 | PCF11 | 121 | 2855 | 0.128 | 0.0423 | No | ||
| 16 | MDGA1 | 13231 | 2974 | 0.110 | 0.0382 | No | ||
| 17 | SLC12A8 | 5059 | 3021 | 0.104 | 0.0378 | No | ||
| 18 | MYST3 | 18651 | 3040 | 0.101 | 0.0389 | No | ||
| 19 | FOXP2 | 4312 17523 8523 | 3267 | 0.076 | 0.0282 | No | ||
| 20 | PHF16 | 11795 6901 | 3534 | 0.057 | 0.0150 | No | ||
| 21 | H2AFY | 21447 | 3733 | 0.048 | 0.0052 | No | ||
| 22 | DTNA | 8870 4647 23611 1991 | 3759 | 0.047 | 0.0048 | No | ||
| 23 | CFL2 | 2155 21069 | 3769 | 0.046 | 0.0053 | No | ||
| 24 | GNAQ | 4786 23909 3685 | 3787 | 0.045 | 0.0053 | No | ||
| 25 | HOXC4 | 4864 | 3955 | 0.039 | -0.0030 | No | ||
| 26 | SLC26A9 | 11560 980 4067 | 4057 | 0.036 | -0.0077 | No | ||
| 27 | GATA2 | 17370 | 4069 | 0.036 | -0.0076 | No | ||
| 28 | TECTA | 19159 | 4159 | 0.033 | -0.0117 | No | ||
| 29 | FBN2 | 23433 | 4192 | 0.032 | -0.0128 | No | ||
| 30 | AP1G2 | 2677 21827 | 4196 | 0.032 | -0.0123 | No | ||
| 31 | TNFRSF19L | 11498 6727 6726 | 4323 | 0.029 | -0.0186 | No | ||
| 32 | PAK1 | 9527 | 4349 | 0.029 | -0.0193 | No | ||
| 33 | GRAP | 20851 | 4498 | 0.026 | -0.0268 | No | ||
| 34 | TBX5 | 3531 5634 | 4706 | 0.022 | -0.0376 | No | ||
| 35 | STC1 | 21974 | 4788 | 0.021 | -0.0415 | No | ||
| 36 | NTF3 | 16991 | 4821 | 0.021 | -0.0428 | No | ||
| 37 | CD36 | 8712 | 4851 | 0.021 | -0.0440 | No | ||
| 38 | SIX4 | 5445 | 4869 | 0.021 | -0.0445 | No | ||
| 39 | EMCN | 7211 | 4925 | 0.020 | -0.0471 | No | ||
| 40 | SOX2 | 9849 15612 | 4989 | 0.019 | -0.0501 | No | ||
| 41 | SOX6 | 3684 5481 9851 | 5190 | 0.017 | -0.0606 | No | ||
| 42 | ALDH1A2 | 19386 | 5372 | 0.015 | -0.0701 | No | ||
| 43 | ELAVL4 | 15805 4889 9137 | 5378 | 0.015 | -0.0701 | No | ||
| 44 | BNC2 | 15850 | 5416 | 0.015 | -0.0718 | No | ||
| 45 | FST | 8985 | 5505 | 0.014 | -0.0763 | No | ||
| 46 | MOBP | 5112 | 5775 | 0.012 | -0.0906 | No | ||
| 47 | MLLT3 | 2359 15847 2413 | 5810 | 0.012 | -0.0922 | No | ||
| 48 | ABCA6 | 1470 20165 | 6026 | 0.011 | -0.1036 | No | ||
| 49 | POU4F1 | 21727 | 6064 | 0.011 | -0.1054 | No | ||
| 50 | OLFM1 | 2918 2971 | 6165 | 0.010 | -0.1106 | No | ||
| 51 | CRH | 8784 | 6342 | 0.009 | -0.1200 | No | ||
| 52 | LRCH2 | 24035 | 6438 | 0.009 | -0.1250 | No | ||
| 53 | B3GALT2 | 6498 | 6661 | 0.008 | -0.1368 | No | ||
| 54 | MRC2 | 1386 5117 | 6814 | 0.007 | -0.1449 | No | ||
| 55 | NR3C2 | 18562 13798 | 6901 | 0.007 | -0.1494 | No | ||
| 56 | NR4A1 | 9099 | 6906 | 0.007 | -0.1495 | No | ||
| 57 | PTN | 5323 | 7188 | 0.006 | -0.1646 | No | ||
| 58 | KLHL13 | 12536 | 7210 | 0.006 | -0.1656 | No | ||
| 59 | HOXD10 | 14983 | 7260 | 0.006 | -0.1682 | No | ||
| 60 | EN2 | 16898 | 7358 | 0.006 | -0.1733 | No | ||
| 61 | PAK3 | 9528 | 7456 | 0.005 | -0.1785 | No | ||
| 62 | SOX5 | 9850 1044 5480 16937 | 7475 | 0.005 | -0.1793 | No | ||
| 63 | HOXB5 | 20687 | 7797 | 0.004 | -0.1966 | No | ||
| 64 | RCN1 | 9710 | 8161 | 0.003 | -0.2162 | No | ||
| 65 | GPR110 | 23234 | 8164 | 0.003 | -0.2163 | No | ||
| 66 | APBA1 | 11483 | 8189 | 0.003 | -0.2175 | No | ||
| 67 | ELAVL2 | 15839 | 8502 | 0.002 | -0.2344 | No | ||
| 68 | TCF1 | 16416 | 8601 | 0.002 | -0.2396 | No | ||
| 69 | NUDT11 | 24387 | 8703 | 0.002 | -0.2451 | No | ||
| 70 | NKX2-2 | 5176 | 9246 | 0.001 | -0.2744 | No | ||
| 71 | SEMA6C | 1797 5424 | 9312 | 0.000 | -0.2780 | No | ||
| 72 | GNAO1 | 4785 3829 | 9360 | 0.000 | -0.2805 | No | ||
| 73 | TIAL1 | 996 5753 | 9368 | 0.000 | -0.2809 | No | ||
| 74 | NFIB | 15855 | 9395 | 0.000 | -0.2823 | No | ||
| 75 | IL5RA | 17051 | 9451 | 0.000 | -0.2853 | No | ||
| 76 | DEPDC2 | 14295 | 9655 | -0.000 | -0.2962 | No | ||
| 77 | EBF2 | 21975 | 9754 | -0.001 | -0.3015 | No | ||
| 78 | PTGFRN | 15224 | 9842 | -0.001 | -0.3062 | No | ||
| 79 | CAMK1G | 13710 4129 | 10039 | -0.001 | -0.3168 | No | ||
| 80 | EPHA7 | 8909 4674 | 10087 | -0.002 | -0.3194 | No | ||
| 81 | FN1 | 4091 4094 971 13929 | 10493 | -0.002 | -0.3413 | No | ||
| 82 | SLC7A1 | 4430 16285 | 10496 | -0.002 | -0.3413 | No | ||
| 83 | PURA | 9670 | 10677 | -0.003 | -0.3510 | No | ||
| 84 | CHN1 | 4246 | 10792 | -0.003 | -0.3571 | No | ||
| 85 | MID2 | 24232 | 11233 | -0.004 | -0.3809 | No | ||
| 86 | NR4A3 | 9473 16212 5183 | 11248 | -0.004 | -0.3816 | No | ||
| 87 | ZIC2 | 21927 | 11405 | -0.005 | -0.3899 | No | ||
| 88 | PIK3R1 | 3170 | 11690 | -0.006 | -0.4052 | No | ||
| 89 | MITF | 17349 | 11764 | -0.006 | -0.4090 | No | ||
| 90 | TCF15 | 14798 | 11788 | -0.006 | -0.4101 | No | ||
| 91 | NFIA | 16172 5170 | 11867 | -0.006 | -0.4142 | No | ||
| 92 | FOXB1 | 12254 19069 | 11900 | -0.007 | -0.4158 | No | ||
| 93 | LPHN2 | 13619 | 11921 | -0.007 | -0.4168 | No | ||
| 94 | BAMBI | 23629 | 11962 | -0.007 | -0.4188 | No | ||
| 95 | LRRN1 | 17343 | 12007 | -0.007 | -0.4210 | No | ||
| 96 | CCNJ | 23859 | 12040 | -0.007 | -0.4226 | No | ||
| 97 | QKI | 23128 5340 | 12095 | -0.007 | -0.4254 | No | ||
| 98 | LEMD1 | 10023 537 | 12199 | -0.008 | -0.4308 | No | ||
| 99 | ESRRG | 6447 | 12271 | -0.008 | -0.4345 | No | ||
| 100 | SP8 | 21122 | 12434 | -0.009 | -0.4431 | No | ||
| 101 | HS3ST1 | 16542 | 12466 | -0.009 | -0.4446 | No | ||
| 102 | OLFML3 | 15216 | 12546 | -0.009 | -0.4487 | No | ||
| 103 | TBX6 | 1993 18075 | 12820 | -0.011 | -0.4633 | No | ||
| 104 | CLDN18 | 19028 | 12890 | -0.011 | -0.4668 | No | ||
| 105 | HEY1 | 15378 | 13020 | -0.012 | -0.4735 | No | ||
| 106 | LRP5 | 23948 9285 | 13023 | -0.012 | -0.4734 | No | ||
| 107 | PELI2 | 8217 | 13394 | -0.015 | -0.4931 | No | ||
| 108 | CTLA4 | 14238 | 13455 | -0.015 | -0.4961 | No | ||
| 109 | SLC13A1 | 17205 | 13667 | -0.017 | -0.5072 | No | ||
| 110 | MLLT7 | 24278 | 13782 | -0.019 | -0.5130 | No | ||
| 111 | CACNA1E | 4465 | 13948 | -0.021 | -0.5215 | No | ||
| 112 | KERA | 19895 | 13975 | -0.021 | -0.5225 | No | ||
| 113 | MYLK | 22778 4213 | 14009 | -0.022 | -0.5238 | No | ||
| 114 | MAB21L1 | 15589 5046 | 14032 | -0.022 | -0.5245 | No | ||
| 115 | ANGPT1 | 8588 2260 22303 | 14316 | -0.027 | -0.5393 | No | ||
| 116 | DHX40 | 20307 | 14523 | -0.031 | -0.5499 | No | ||
| 117 | A2BP1 | 11212 1652 1689 | 14530 | -0.031 | -0.5495 | No | ||
| 118 | POU3F4 | 347 | 14572 | -0.033 | -0.5511 | No | ||
| 119 | EIF4G2 | 1908 8892 | 14640 | -0.035 | -0.5540 | No | ||
| 120 | DHRS3 | 15997 | 14906 | -0.043 | -0.5675 | No | ||
| 121 | RBBP4 | 5364 | 14955 | -0.045 | -0.5692 | No | ||
| 122 | FGF9 | 8966 | 15013 | -0.048 | -0.5713 | No | ||
| 123 | ARHGAP15 | 8004 377 13310 | 15045 | -0.049 | -0.5720 | No | ||
| 124 | RBM9 | 2276 13528 | 15102 | -0.051 | -0.5740 | No | ||
| 125 | DLL1 | 23119 | 15370 | -0.067 | -0.5871 | No | ||
| 126 | LRP1 | 9284 | 15413 | -0.071 | -0.5879 | No | ||
| 127 | CPNE1 | 11186 | 15562 | -0.083 | -0.5943 | No | ||
| 128 | ULK1 | 16436 | 15588 | -0.085 | -0.5939 | No | ||
| 129 | AP4M1 | 3499 16656 | 15591 | -0.085 | -0.5922 | No | ||
| 130 | CCBL1 | 14630 | 15598 | -0.086 | -0.5908 | No | ||
| 131 | NFE2L1 | 9457 | 15839 | -0.111 | -0.6016 | No | ||
| 132 | FGFR2 | 1917 4722 1119 | 15861 | -0.114 | -0.6004 | No | ||
| 133 | TGFB3 | 10161 | 15910 | -0.122 | -0.6005 | No | ||
| 134 | PUM1 | 8160 | 15923 | -0.126 | -0.5985 | No | ||
| 135 | KCNN2 | 8933 1949 | 15971 | -0.135 | -0.5983 | No | ||
| 136 | XKRX | 301 | 16127 | -0.169 | -0.6033 | No | ||
| 137 | STAG2 | 5521 | 16171 | -0.178 | -0.6020 | No | ||
| 138 | ICK | 7135 12146 | 16203 | -0.186 | -0.5999 | No | ||
| 139 | YWHAG | 16339 | 16238 | -0.194 | -0.5978 | No | ||
| 140 | ZADH2 | 23501 | 16450 | -0.240 | -0.6043 | No | ||
| 141 | RAPGEF5 | 5739 10155 21123 | 16707 | -0.319 | -0.6117 | Yes | ||
| 142 | RHOBTB1 | 12789 | 16786 | -0.344 | -0.6089 | Yes | ||
| 143 | ZFYVE1 | 10138 | 16800 | -0.350 | -0.6024 | Yes | ||
| 144 | MYO18A | 20766 | 16865 | -0.380 | -0.5981 | Yes | ||
| 145 | PRKAG1 | 22135 | 17156 | -0.489 | -0.6038 | Yes | ||
| 146 | WASL | 17203 | 17292 | -0.543 | -0.6001 | Yes | ||
| 147 | TACSTD2 | 17126 | 17349 | -0.572 | -0.5914 | Yes | ||
| 148 | COMMD3 | 73 | 17530 | -0.674 | -0.5874 | Yes | ||
| 149 | HLX1 | 9093 | 17549 | -0.689 | -0.5743 | Yes | ||
| 150 | MXI1 | 23813 5139 | 17574 | -0.700 | -0.5613 | Yes | ||
| 151 | TCF4 | 5642 | 17625 | -0.736 | -0.5490 | Yes | ||
| 152 | NSF | 20192 | 17728 | -0.812 | -0.5380 | Yes | ||
| 153 | KBTBD2 | 17137 | 17749 | -0.823 | -0.5222 | Yes | ||
| 154 | PIAS1 | 7126 | 17914 | -0.971 | -0.5113 | Yes | ||
| 155 | MSI2 | 8030 1268 | 17922 | -0.983 | -0.4916 | Yes | ||
| 156 | KLF7 | 8205 | 18024 | -1.105 | -0.4745 | Yes | ||
| 157 | PPM1D | 20721 | 18030 | -1.116 | -0.4520 | Yes | ||
| 158 | MAML1 | 20477 | 18098 | -1.213 | -0.4308 | Yes | ||
| 159 | MBNL2 | 21932 | 18117 | -1.242 | -0.4064 | Yes | ||
| 160 | KLF3 | 4961 | 18251 | -1.482 | -0.3834 | Yes | ||
| 161 | LMO4 | 15151 | 18276 | -1.537 | -0.3533 | Yes | ||
| 162 | CDC42EP3 | 6436 | 18383 | -1.819 | -0.3219 | Yes | ||
| 163 | SATB1 | 5408 | 18466 | -2.201 | -0.2814 | Yes | ||
| 164 | NFKBIE | 23225 1556 | 18480 | -2.288 | -0.2353 | Yes | ||
| 165 | DDX6 | 3086 8844 | 18487 | -2.335 | -0.1880 | Yes | ||
| 166 | TNFRSF19 | 21799 3033 | 18498 | -2.427 | -0.1389 | Yes | ||
| 167 | PIK3CG | 6635 | 18517 | -2.540 | -0.0880 | Yes | ||
| 168 | DUSP6 | 19891 3399 | 18597 | -4.571 | 0.0010 | Yes |