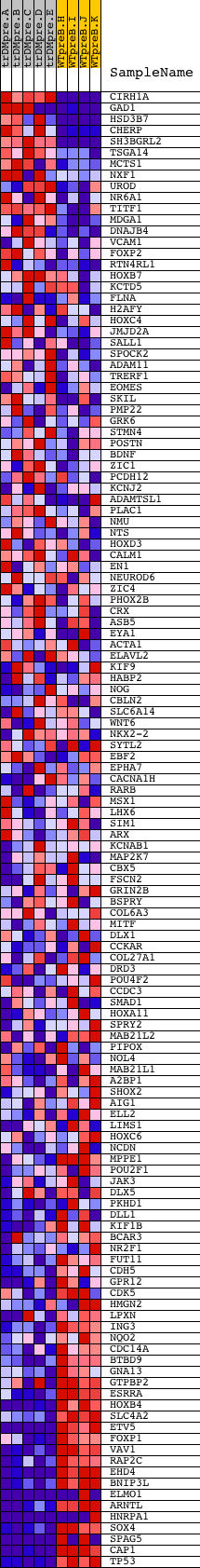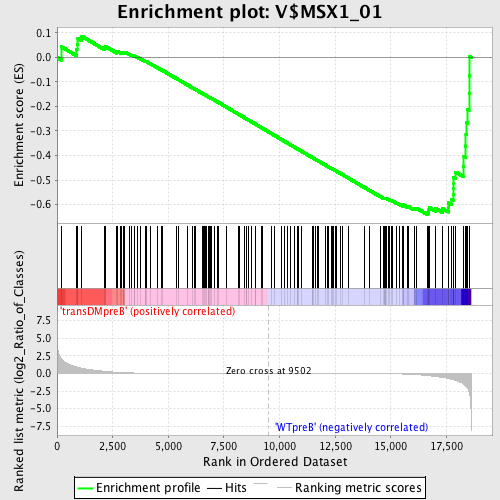
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_WTpreB.phenotype_transDMpreB_versus_WTpreB.cls #transDMpreB_versus_WTpreB |
| Phenotype | phenotype_transDMpreB_versus_WTpreB.cls#transDMpreB_versus_WTpreB |
| Upregulated in class | WTpreB |
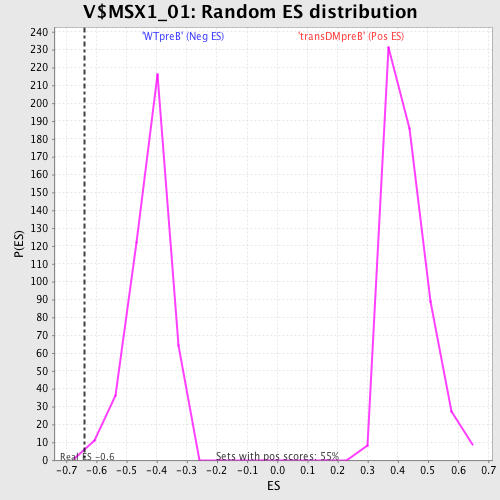
| GeneSet | V$MSX1_01 |
| Enrichment Score (ES) | -0.64069813 |
| Normalized Enrichment Score (NES) | -1.5222709 |
| Nominal p-value | 0.0022222223 |
| FDR q-value | 0.15375887 |
| FWER p-Value | 0.684 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CIRH1A | 18478 | 198 | 2.016 | 0.0437 | No | ||
| 2 | GAD1 | 14993 2690 | 876 | 0.913 | 0.0316 | No | ||
| 3 | HSD3B7 | 4158 | 895 | 0.897 | 0.0549 | No | ||
| 4 | CHERP | 11367 | 934 | 0.868 | 0.0762 | No | ||
| 5 | SH3BGRL2 | 5601 | 1101 | 0.736 | 0.0871 | No | ||
| 6 | TSGA14 | 17194 | 2125 | 0.305 | 0.0400 | No | ||
| 7 | MCTS1 | 24351 | 2174 | 0.291 | 0.0452 | No | ||
| 8 | NXF1 | 11986 | 2679 | 0.160 | 0.0223 | No | ||
| 9 | UROD | 15791 | 2717 | 0.154 | 0.0244 | No | ||
| 10 | NR6A1 | 14596 | 2857 | 0.128 | 0.0204 | No | ||
| 11 | TITF1 | 10180 21062 | 2912 | 0.120 | 0.0207 | No | ||
| 12 | MDGA1 | 13231 | 2974 | 0.110 | 0.0203 | No | ||
| 13 | DNAJB4 | 12451 1916 1832 | 2999 | 0.108 | 0.0219 | No | ||
| 14 | VCAM1 | 5851 | 3050 | 0.100 | 0.0219 | No | ||
| 15 | FOXP2 | 4312 17523 8523 | 3267 | 0.076 | 0.0123 | No | ||
| 16 | RTN4RL1 | 20782 | 3348 | 0.070 | 0.0099 | No | ||
| 17 | HOXB7 | 666 9110 | 3467 | 0.061 | 0.0051 | No | ||
| 18 | KCTD5 | 23096 | 3480 | 0.060 | 0.0061 | No | ||
| 19 | FLNA | 24129 | 3592 | 0.054 | 0.0016 | No | ||
| 20 | H2AFY | 21447 | 3733 | 0.048 | -0.0047 | No | ||
| 21 | HOXC4 | 4864 | 3955 | 0.039 | -0.0156 | No | ||
| 22 | JMJD2A | 10505 6058 | 3974 | 0.039 | -0.0155 | No | ||
| 23 | SALL1 | 18796 12207 | 4028 | 0.037 | -0.0174 | No | ||
| 24 | SPOCK2 | 13585 | 4198 | 0.032 | -0.0257 | No | ||
| 25 | ADAM11 | 4340 1432 | 4514 | 0.026 | -0.0420 | No | ||
| 26 | TRERF1 | 1546 23213 | 4713 | 0.022 | -0.0521 | No | ||
| 27 | EOMES | 4669 | 4740 | 0.022 | -0.0530 | No | ||
| 28 | SKIL | 15617 | 5343 | 0.015 | -0.0851 | No | ||
| 29 | PMP22 | 9594 1312 | 5451 | 0.014 | -0.0905 | No | ||
| 30 | GRK6 | 6449 | 5869 | 0.012 | -0.1128 | No | ||
| 31 | STMN4 | 21977 | 6079 | 0.010 | -0.1238 | No | ||
| 32 | POSTN | 1801 11927 | 6178 | 0.010 | -0.1288 | No | ||
| 33 | BDNF | 14926 2797 | 6229 | 0.010 | -0.1313 | No | ||
| 34 | ZIC1 | 19040 | 6535 | 0.008 | -0.1475 | No | ||
| 35 | PCDH12 | 7005 | 6584 | 0.008 | -0.1499 | No | ||
| 36 | KCNJ2 | 20612 | 6625 | 0.008 | -0.1519 | No | ||
| 37 | ADAMTSL1 | 16181 | 6670 | 0.008 | -0.1540 | No | ||
| 38 | PLAC1 | 12079 | 6700 | 0.008 | -0.1554 | No | ||
| 39 | NMU | 16506 | 6804 | 0.007 | -0.1607 | No | ||
| 40 | NTS | 19635 3297 | 6806 | 0.007 | -0.1606 | No | ||
| 41 | HOXD3 | 9115 | 6843 | 0.007 | -0.1624 | No | ||
| 42 | CALM1 | 21184 | 6895 | 0.007 | -0.1649 | No | ||
| 43 | EN1 | 387 14155 | 6944 | 0.007 | -0.1673 | No | ||
| 44 | NEUROD6 | 17138 | 7095 | 0.006 | -0.1753 | No | ||
| 45 | ZIC4 | 5992 | 7199 | 0.006 | -0.1807 | No | ||
| 46 | PHOX2B | 16524 | 7272 | 0.006 | -0.1844 | No | ||
| 47 | CRX | 17961 4694 | 7608 | 0.005 | -0.2024 | No | ||
| 48 | ASB5 | 13322 | 8155 | 0.003 | -0.2319 | No | ||
| 49 | EYA1 | 4695 4061 | 8207 | 0.003 | -0.2345 | No | ||
| 50 | ACTA1 | 18715 | 8440 | 0.003 | -0.2470 | No | ||
| 51 | ELAVL2 | 15839 | 8502 | 0.002 | -0.2502 | No | ||
| 52 | KIF9 | 19292 | 8521 | 0.002 | -0.2512 | No | ||
| 53 | HABP2 | 10364 23825 | 8591 | 0.002 | -0.2548 | No | ||
| 54 | NOG | 13663 | 8741 | 0.002 | -0.2628 | No | ||
| 55 | CBLN2 | 23498 | 8895 | 0.001 | -0.2711 | No | ||
| 56 | SLC6A14 | 12168 | 8927 | 0.001 | -0.2727 | No | ||
| 57 | WNT6 | 5881 | 9202 | 0.001 | -0.2875 | No | ||
| 58 | NKX2-2 | 5176 | 9246 | 0.001 | -0.2898 | No | ||
| 59 | SYTL2 | 18190 1539 3730 | 9646 | -0.000 | -0.3114 | No | ||
| 60 | EBF2 | 21975 | 9754 | -0.001 | -0.3172 | No | ||
| 61 | EPHA7 | 8909 4674 | 10087 | -0.002 | -0.3351 | No | ||
| 62 | CACNA1H | 23077 | 10200 | -0.002 | -0.3411 | No | ||
| 63 | RARB | 3011 21915 | 10377 | -0.002 | -0.3506 | No | ||
| 64 | MSX1 | 16547 | 10509 | -0.003 | -0.3576 | No | ||
| 65 | LHX6 | 9275 2914 | 10689 | -0.003 | -0.3672 | No | ||
| 66 | SIM1 | 20031 | 10815 | -0.003 | -0.3739 | No | ||
| 67 | ARX | 24290 | 10825 | -0.003 | -0.3743 | No | ||
| 68 | KCNAB1 | 1791 15577 | 10868 | -0.003 | -0.3764 | No | ||
| 69 | MAP2K7 | 6453 | 10987 | -0.004 | -0.3827 | No | ||
| 70 | CBX5 | 22108 4485 | 11480 | -0.005 | -0.4092 | No | ||
| 71 | FSCN2 | 20569 | 11496 | -0.005 | -0.4099 | No | ||
| 72 | GRIN2B | 16957 | 11537 | -0.005 | -0.4119 | No | ||
| 73 | BSPRY | 5307 | 11623 | -0.006 | -0.4163 | No | ||
| 74 | COL6A3 | 4546 13885 13884 | 11709 | -0.006 | -0.4208 | No | ||
| 75 | MITF | 17349 | 11764 | -0.006 | -0.4235 | No | ||
| 76 | DLX1 | 14988 | 12045 | -0.007 | -0.4385 | No | ||
| 77 | CCKAR | 16531 | 12133 | -0.007 | -0.4430 | No | ||
| 78 | COL27A1 | 6885 | 12216 | -0.008 | -0.4472 | No | ||
| 79 | DRD3 | 22750 | 12341 | -0.008 | -0.4537 | No | ||
| 80 | POU4F2 | 9603 5276 | 12393 | -0.009 | -0.4562 | No | ||
| 81 | CCDC3 | 15122 | 12405 | -0.009 | -0.4566 | No | ||
| 82 | SMAD1 | 18831 922 | 12519 | -0.009 | -0.4624 | No | ||
| 83 | HOXA11 | 17146 | 12539 | -0.009 | -0.4632 | No | ||
| 84 | SPRY2 | 21725 | 12759 | -0.011 | -0.4748 | No | ||
| 85 | MAB21L2 | 15305 | 12840 | -0.011 | -0.4788 | No | ||
| 86 | PIPOX | 20340 | 13081 | -0.012 | -0.4914 | No | ||
| 87 | NOL4 | 2028 11432 | 13819 | -0.019 | -0.5308 | No | ||
| 88 | MAB21L1 | 15589 5046 | 14032 | -0.022 | -0.5417 | No | ||
| 89 | A2BP1 | 11212 1652 1689 | 14530 | -0.031 | -0.5677 | No | ||
| 90 | SHOX2 | 1802 5432 | 14688 | -0.036 | -0.5752 | No | ||
| 91 | AIG1 | 7272 | 14698 | -0.036 | -0.5747 | No | ||
| 92 | ELL2 | 5329 | 14734 | -0.037 | -0.5756 | No | ||
| 93 | LIMS1 | 8493 4290 | 14750 | -0.038 | -0.5754 | No | ||
| 94 | HOXC6 | 22339 2302 | 14775 | -0.039 | -0.5757 | No | ||
| 95 | NCDN | 15756 2487 6471 | 14793 | -0.039 | -0.5756 | No | ||
| 96 | MPPE1 | 23417 2013 | 14903 | -0.043 | -0.5803 | No | ||
| 97 | POU2F1 | 5275 3989 4065 4010 | 14921 | -0.044 | -0.5800 | No | ||
| 98 | JAK3 | 9198 4936 | 15025 | -0.048 | -0.5843 | No | ||
| 99 | DLX5 | 17219 1058 | 15063 | -0.050 | -0.5850 | No | ||
| 100 | PKHD1 | 6292 4106 | 15271 | -0.060 | -0.5945 | No | ||
| 101 | DLL1 | 23119 | 15370 | -0.067 | -0.5980 | No | ||
| 102 | KIF1B | 15671 2476 4950 15673 | 15536 | -0.081 | -0.6048 | No | ||
| 103 | BCAR3 | 15432 | 15560 | -0.083 | -0.6038 | No | ||
| 104 | NR2F1 | 7015 21401 | 15561 | -0.083 | -0.6016 | No | ||
| 105 | FUT11 | 13084 | 15733 | -0.099 | -0.6081 | No | ||
| 106 | CDH5 | 18512 | 15813 | -0.108 | -0.6095 | No | ||
| 107 | GPR12 | 16291 | 16045 | -0.147 | -0.6181 | No | ||
| 108 | CDK5 | 16591 | 16077 | -0.157 | -0.6155 | No | ||
| 109 | HMGN2 | 9095 | 16174 | -0.179 | -0.6159 | No | ||
| 110 | LPXN | 8399 | 16634 | -0.295 | -0.6327 | Yes | ||
| 111 | ING3 | 17514 1019 | 16687 | -0.312 | -0.6272 | Yes | ||
| 112 | NQO2 | 9469 3167 | 16712 | -0.320 | -0.6198 | Yes | ||
| 113 | CDC14A | 15184 | 16732 | -0.329 | -0.6120 | Yes | ||
| 114 | BTBD9 | 23040 | 17010 | -0.437 | -0.6152 | Yes | ||
| 115 | GNA13 | 20617 | 17337 | -0.563 | -0.6176 | Yes | ||
| 116 | GTPBP2 | 23221 | 17578 | -0.701 | -0.6117 | Yes | ||
| 117 | ESRRA | 23802 | 17590 | -0.710 | -0.5932 | Yes | ||
| 118 | HOXB4 | 20686 | 17727 | -0.811 | -0.5787 | Yes | ||
| 119 | SLC4A2 | 9828 | 17799 | -0.866 | -0.5592 | Yes | ||
| 120 | ETV5 | 22630 | 17811 | -0.879 | -0.5361 | Yes | ||
| 121 | FOXP1 | 4242 | 17816 | -0.882 | -0.5125 | Yes | ||
| 122 | VAV1 | 23173 | 17823 | -0.886 | -0.4889 | Yes | ||
| 123 | RAP2C | 24156 | 17921 | -0.982 | -0.4677 | Yes | ||
| 124 | EHD4 | 8275 | 18265 | -1.508 | -0.4456 | Yes | ||
| 125 | BNIP3L | 21775 | 18286 | -1.555 | -0.4048 | Yes | ||
| 126 | ELMO1 | 3275 8939 3202 3271 3254 3235 3164 3175 3274 3188 3293 3214 3218 3239 | 18347 | -1.737 | -0.3612 | Yes | ||
| 127 | ARNTL | 18119 | 18377 | -1.808 | -0.3140 | Yes | ||
| 128 | HNRPA1 | 9103 4860 | 18392 | -1.837 | -0.2652 | Yes | ||
| 129 | SOX4 | 5479 | 18455 | -2.118 | -0.2114 | Yes | ||
| 130 | SPAG5 | 20760 1341 | 18515 | -2.537 | -0.1462 | Yes | ||
| 131 | CAP1 | 8684 | 18528 | -2.681 | -0.0746 | Yes | ||
| 132 | TP53 | 20822 | 18549 | -2.941 | 0.0036 | Yes |