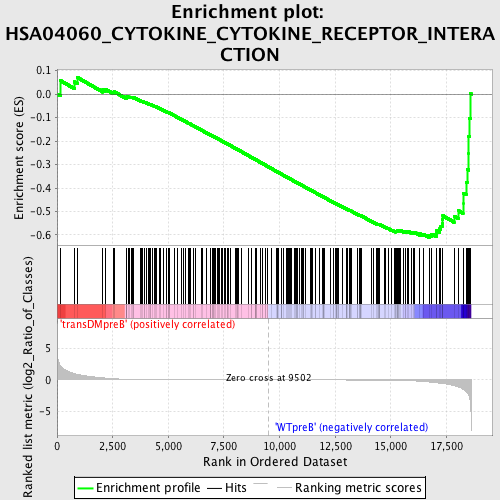
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_WTpreB.phenotype_transDMpreB_versus_WTpreB.cls #transDMpreB_versus_WTpreB |
| Phenotype | phenotype_transDMpreB_versus_WTpreB.cls#transDMpreB_versus_WTpreB |
| Upregulated in class | WTpreB |
| GeneSet | HSA04060_CYTOKINE_CYTOKINE_RECEPTOR_INTERACTION |
| Enrichment Score (ES) | -0.6106816 |
| Normalized Enrichment Score (NES) | -1.5313168 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.19116113 |
| FWER p-Value | 0.75 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | IL10 | 14145 1510 1553 22902 | 148 | 2.248 | 0.0575 | No | ||
| 2 | VEGFB | 23800 | 785 | 0.996 | 0.0520 | No | ||
| 3 | ACVR2B | 19276 | 921 | 0.880 | 0.0704 | No | ||
| 4 | VEGFA | 22969 | 2056 | 0.325 | 0.0182 | No | ||
| 5 | CSF1 | 15202 | 2160 | 0.294 | 0.0212 | No | ||
| 6 | IL1B | 14431 | 2517 | 0.193 | 0.0075 | No | ||
| 7 | IL12RB1 | 3916 18582 | 2569 | 0.181 | 0.0100 | No | ||
| 8 | EGF | 15169 | 3113 | 0.092 | -0.0168 | No | ||
| 9 | CCL5 | 20320 | 3122 | 0.091 | -0.0146 | No | ||
| 10 | IL1R1 | 14264 3987 | 3133 | 0.090 | -0.0125 | No | ||
| 11 | IL2RB | 22219 | 3136 | 0.090 | -0.0100 | No | ||
| 12 | IL6ST | 4920 | 3222 | 0.080 | -0.0123 | No | ||
| 13 | IL6R | 1862 4919 | 3264 | 0.077 | -0.0123 | No | ||
| 14 | TNFSF13 | 12806 | 3334 | 0.072 | -0.0139 | No | ||
| 15 | TGFB2 | 5743 | 3382 | 0.068 | -0.0145 | No | ||
| 16 | CCL27 | 5419 | 3426 | 0.064 | -0.0150 | No | ||
| 17 | IL9R | 20505 | 3440 | 0.063 | -0.0138 | No | ||
| 18 | IFNA14 | 11917 | 3727 | 0.048 | -0.0280 | No | ||
| 19 | IL18R1 | 14263 | 3794 | 0.045 | -0.0303 | No | ||
| 20 | IL15RA | 2899 2717 15118 2792 | 3833 | 0.044 | -0.0310 | No | ||
| 21 | IL4 | 9174 | 3933 | 0.040 | -0.0352 | No | ||
| 22 | IL24 | 976 13841 | 3943 | 0.040 | -0.0346 | No | ||
| 23 | TNFRSF10B | 21971 10205 5781 10204 | 4025 | 0.037 | -0.0379 | No | ||
| 24 | PPBP | 16801 | 4108 | 0.034 | -0.0413 | No | ||
| 25 | IL15 | 18826 3801 | 4158 | 0.033 | -0.0430 | No | ||
| 26 | CX3CR1 | 18965 | 4184 | 0.032 | -0.0435 | No | ||
| 27 | CCL17 | 18517 | 4191 | 0.032 | -0.0428 | No | ||
| 28 | CXCL9 | 5099 | 4301 | 0.030 | -0.0479 | No | ||
| 29 | AMHR2 | 8487 22345 | 4377 | 0.028 | -0.0512 | No | ||
| 30 | KIT | 16823 | 4403 | 0.027 | -0.0517 | No | ||
| 31 | IL17A | 14289 | 4452 | 0.027 | -0.0535 | No | ||
| 32 | PDGFRB | 23539 | 4594 | 0.024 | -0.0605 | No | ||
| 33 | TNFSF13B | 6291 10792 | 4648 | 0.023 | -0.0627 | No | ||
| 34 | XCL1 | 13781 | 4775 | 0.022 | -0.0689 | No | ||
| 35 | IFNA1 | 9147 | 4905 | 0.020 | -0.0753 | No | ||
| 36 | CCL1 | 9787 | 4907 | 0.020 | -0.0748 | No | ||
| 37 | TNFRSF17 | 22856 | 5000 | 0.019 | -0.0793 | No | ||
| 38 | IL5 | 20884 10220 | 5018 | 0.019 | -0.0797 | No | ||
| 39 | PDGFC | 7048 | 5029 | 0.018 | -0.0797 | No | ||
| 40 | IFNA7 | 9152 | 5049 | 0.018 | -0.0802 | No | ||
| 41 | CCL4 | 1362 20730 | 5266 | 0.016 | -0.0914 | No | ||
| 42 | CCL8 | 20740 | 5420 | 0.015 | -0.0993 | No | ||
| 43 | CSF2RA | 23631 | 5576 | 0.013 | -0.1073 | No | ||
| 44 | CX3CL1 | 9791 | 5699 | 0.013 | -0.1136 | No | ||
| 45 | CCL3 | 20317 | 5756 | 0.012 | -0.1163 | No | ||
| 46 | IL1A | 4915 | 5911 | 0.011 | -0.1243 | No | ||
| 47 | CNTFR | 2515 15906 | 5952 | 0.011 | -0.1262 | No | ||
| 48 | TNFSF15 | 15863 | 5973 | 0.011 | -0.1270 | No | ||
| 49 | IL6 | 16895 | 6131 | 0.010 | -0.1352 | No | ||
| 50 | CCL22 | 18518 | 6142 | 0.010 | -0.1354 | No | ||
| 51 | IL25 | 22014 | 6221 | 0.010 | -0.1394 | No | ||
| 52 | CCL2 | 9788 | 6475 | 0.009 | -0.1529 | No | ||
| 53 | CSF3 | 1394 20671 | 6543 | 0.008 | -0.1563 | No | ||
| 54 | IFNB1 | 15846 | 6706 | 0.008 | -0.1649 | No | ||
| 55 | TNFRSF8 | 2373 5784 | 6877 | 0.007 | -0.1739 | No | ||
| 56 | CXCL12 | 9792 1182 1031 | 6882 | 0.007 | -0.1739 | No | ||
| 57 | IL19 | 13839 | 6915 | 0.007 | -0.1754 | No | ||
| 58 | IL3 | 20453 | 6996 | 0.007 | -0.1796 | No | ||
| 59 | EPO | 8911 | 7000 | 0.007 | -0.1796 | No | ||
| 60 | TPO | 21092 | 7013 | 0.007 | -0.1800 | No | ||
| 61 | TNFSF18 | 10789 6290 | 7044 | 0.007 | -0.1815 | No | ||
| 62 | EDA2R | 6364 | 7057 | 0.007 | -0.1819 | No | ||
| 63 | IFNA4 | 9150 | 7127 | 0.006 | -0.1855 | No | ||
| 64 | IL8RA | 13925 | 7196 | 0.006 | -0.1890 | No | ||
| 65 | CCR2 | 19250 | 7243 | 0.006 | -0.1913 | No | ||
| 66 | IFNK | 11837 | 7304 | 0.006 | -0.1944 | No | ||
| 67 | IL13 | 20461 | 7370 | 0.005 | -0.1978 | No | ||
| 68 | IL23R | 17127 | 7448 | 0.005 | -0.2018 | No | ||
| 69 | FLT3 | 16288 | 7531 | 0.005 | -0.2061 | No | ||
| 70 | LTBR | 16993 | 7578 | 0.005 | -0.2085 | No | ||
| 71 | CCL11 | 20742 | 7646 | 0.005 | -0.2120 | No | ||
| 72 | ACVR2A | 8542 | 7663 | 0.005 | -0.2127 | No | ||
| 73 | IL22RA1 | 16037 | 7716 | 0.005 | -0.2154 | No | ||
| 74 | CNTF | 3741 8761 3680 3736 | 7810 | 0.004 | -0.2204 | No | ||
| 75 | IL1R2 | 14265 | 8033 | 0.004 | -0.2323 | No | ||
| 76 | CXCR3 | 24093 | 8072 | 0.004 | -0.2343 | No | ||
| 77 | CXCL14 | 21445 7165 | 8089 | 0.004 | -0.2350 | No | ||
| 78 | CD40LG | 24330 | 8128 | 0.003 | -0.2370 | No | ||
| 79 | CCR5 | 8754 4534 | 8141 | 0.003 | -0.2376 | No | ||
| 80 | PDGFB | 22199 2311 | 8288 | 0.003 | -0.2454 | No | ||
| 81 | CXCL11 | 16478 | 8593 | 0.002 | -0.2619 | No | ||
| 82 | IL9 | 21444 | 8717 | 0.002 | -0.2685 | No | ||
| 83 | CSF1R | 8800 23540 | 8745 | 0.002 | -0.2699 | No | ||
| 84 | EDAR | 19764 | 8899 | 0.001 | -0.2782 | No | ||
| 85 | IL12B | 20918 | 8911 | 0.001 | -0.2787 | No | ||
| 86 | CCL24 | 16342 | 8922 | 0.001 | -0.2792 | No | ||
| 87 | GHR | 22336 | 8939 | 0.001 | -0.2801 | No | ||
| 88 | TNFRSF4 | 15954 | 8960 | 0.001 | -0.2811 | No | ||
| 89 | CXCL5 | 16802 | 9130 | 0.001 | -0.2903 | No | ||
| 90 | CCR3 | 19251 | 9222 | 0.001 | -0.2952 | No | ||
| 91 | IL18RAP | 14262 | 9229 | 0.001 | -0.2955 | No | ||
| 92 | CCR1 | 18951 | 9345 | 0.000 | -0.3017 | No | ||
| 93 | IFNE1 | 15842 | 9356 | 0.000 | -0.3023 | No | ||
| 94 | IL5RA | 17051 | 9451 | 0.000 | -0.3074 | No | ||
| 95 | TNFSF8 | 15862 | 9455 | 0.000 | -0.3075 | No | ||
| 96 | GDF5 | 4768 | 9648 | -0.000 | -0.3180 | No | ||
| 97 | LTA | 23003 | 9882 | -0.001 | -0.3306 | No | ||
| 98 | IL11 | 17981 | 9889 | -0.001 | -0.3309 | No | ||
| 99 | CCR8 | 19272 | 9903 | -0.001 | -0.3316 | No | ||
| 100 | PDGFRA | 16824 | 9909 | -0.001 | -0.3318 | No | ||
| 101 | BMP2 | 14833 | 9956 | -0.001 | -0.3343 | No | ||
| 102 | IL22RA2 | 20082 | 10071 | -0.001 | -0.3404 | No | ||
| 103 | LEPR | 2531 2349 | 10089 | -0.002 | -0.3413 | No | ||
| 104 | IL20 | 13840 | 10096 | -0.002 | -0.3416 | No | ||
| 105 | MPL | 15780 | 10175 | -0.002 | -0.3458 | No | ||
| 106 | IL17B | 12074 | 10294 | -0.002 | -0.3521 | No | ||
| 107 | CXCR6 | 19252 | 10365 | -0.002 | -0.3559 | No | ||
| 108 | TNFSF9 | 23175 | 10392 | -0.002 | -0.3572 | No | ||
| 109 | KITLG | 19889 3342 | 10459 | -0.002 | -0.3607 | No | ||
| 110 | IL22 | 11948 | 10470 | -0.002 | -0.3612 | No | ||
| 111 | KDR | 16509 | 10530 | -0.003 | -0.3643 | No | ||
| 112 | IFNAR1 | 22703 | 10662 | -0.003 | -0.3714 | No | ||
| 113 | CXCL2 | 9790 | 10702 | -0.003 | -0.3734 | No | ||
| 114 | HGF | 16916 | 10749 | -0.003 | -0.3758 | No | ||
| 115 | IL13RA1 | 24361 | 10779 | -0.003 | -0.3773 | No | ||
| 116 | IFNA2 | 9149 | 10783 | -0.003 | -0.3774 | No | ||
| 117 | CXCL1 | 9044 | 10817 | -0.003 | -0.3791 | No | ||
| 118 | LEP | 17505 | 10909 | -0.004 | -0.3839 | No | ||
| 119 | INHBC | 19603 | 10966 | -0.004 | -0.3868 | No | ||
| 120 | TNFSF4 | 14082 | 11021 | -0.004 | -0.3896 | No | ||
| 121 | CCR9 | 8753 | 11073 | -0.004 | -0.3923 | No | ||
| 122 | TNFRSF21 | 8244 | 11088 | -0.004 | -0.3929 | No | ||
| 123 | PRL | 21690 | 11180 | -0.004 | -0.3978 | No | ||
| 124 | IFNA13 | 15844 | 11380 | -0.005 | -0.4084 | No | ||
| 125 | CCL19 | 10773 | 11416 | -0.005 | -0.4102 | No | ||
| 126 | TSLP | 23604 1955 | 11418 | -0.005 | -0.4101 | No | ||
| 127 | TNFRSF9 | 10208 5785 | 11497 | -0.005 | -0.4142 | No | ||
| 128 | TNFSF10 | 15626 | 11600 | -0.006 | -0.4196 | No | ||
| 129 | IL21 | 12241 | 11628 | -0.006 | -0.4209 | No | ||
| 130 | CTF1 | 18064 | 11810 | -0.006 | -0.4305 | No | ||
| 131 | TNFSF14 | 1584 22914 | 11949 | -0.007 | -0.4378 | No | ||
| 132 | IFNA6 | 17749 | 11958 | -0.007 | -0.4381 | No | ||
| 133 | XCR1 | 18953 | 11973 | -0.007 | -0.4386 | No | ||
| 134 | IL7 | 4921 | 11981 | -0.007 | -0.4388 | No | ||
| 135 | IL12RB2 | 17128 | 11997 | -0.007 | -0.4394 | No | ||
| 136 | CSF2 | 20454 | 12278 | -0.008 | -0.4544 | No | ||
| 137 | TNFRSF25 | 2334 15983 | 12403 | -0.009 | -0.4609 | No | ||
| 138 | CXCL10 | 16477 | 12408 | -0.009 | -0.4608 | No | ||
| 139 | TNFRSF11B | 22296 | 12524 | -0.009 | -0.4668 | No | ||
| 140 | CXCL16 | 7251 | 12542 | -0.009 | -0.4675 | No | ||
| 141 | TNFSF12 | 10209 | 12545 | -0.009 | -0.4673 | No | ||
| 142 | BMP7 | 14324 | 12554 | -0.010 | -0.4674 | No | ||
| 143 | IL2 | 15354 | 12604 | -0.010 | -0.4698 | No | ||
| 144 | TGFBR2 | 2994 3001 10167 | 12641 | -0.010 | -0.4715 | No | ||
| 145 | TNF | 23004 | 12661 | -0.010 | -0.4722 | No | ||
| 146 | LIF | 956 667 20960 | 12848 | -0.011 | -0.4820 | No | ||
| 147 | IL23A | 19598 | 12991 | -0.012 | -0.4894 | No | ||
| 148 | TNFSF11 | 21745 | 12992 | -0.012 | -0.4891 | No | ||
| 149 | FLT1 | 3483 16287 | 13003 | -0.012 | -0.4892 | No | ||
| 150 | LIFR | 22515 9277 | 13037 | -0.012 | -0.4907 | No | ||
| 151 | OSMR | 22333 | 13144 | -0.013 | -0.4961 | No | ||
| 152 | NGFR | 5174 | 13147 | -0.013 | -0.4958 | No | ||
| 153 | BMPR2 | 14241 4081 | 13160 | -0.013 | -0.4961 | No | ||
| 154 | TNFRSF1B | 5782 2346 | 13166 | -0.013 | -0.4960 | No | ||
| 155 | TNFRSF11A | 14167 | 13169 | -0.013 | -0.4957 | No | ||
| 156 | INHBB | 13858 | 13234 | -0.013 | -0.4988 | No | ||
| 157 | BMPR1A | 8660 4450 | 13483 | -0.015 | -0.5118 | No | ||
| 158 | INHBE | 19604 | 13486 | -0.016 | -0.5115 | No | ||
| 159 | CXCL13 | 16789 | 13588 | -0.017 | -0.5165 | No | ||
| 160 | MET | 17520 | 13603 | -0.017 | -0.5167 | No | ||
| 161 | BMPR1B | 8661 15152 4451 | 13620 | -0.017 | -0.5171 | No | ||
| 162 | EPOR | 19204 | 13635 | -0.017 | -0.5174 | No | ||
| 163 | CD70 | 22916 | 13658 | -0.017 | -0.5181 | No | ||
| 164 | FLT4 | 20904 1461 | 13670 | -0.017 | -0.5182 | No | ||
| 165 | EDA | 24284 | 14108 | -0.023 | -0.5412 | No | ||
| 166 | IL17RB | 21898 | 14199 | -0.025 | -0.5454 | No | ||
| 167 | CD27 | 16994 | 14372 | -0.028 | -0.5539 | No | ||
| 168 | IL28RA | 10823 16038 | 14378 | -0.028 | -0.5534 | No | ||
| 169 | CCR4 | 18978 | 14443 | -0.029 | -0.5560 | No | ||
| 170 | FASLG | 13789 | 14448 | -0.030 | -0.5554 | No | ||
| 171 | CCL28 | 7156 | 14472 | -0.030 | -0.5557 | No | ||
| 172 | IL21R | 1398 18084 | 14478 | -0.030 | -0.5551 | No | ||
| 173 | AMH | 19939 | 14484 | -0.030 | -0.5545 | No | ||
| 174 | TGFBR1 | 16213 10165 5745 | 14721 | -0.037 | -0.5663 | No | ||
| 175 | IFNA5 | 9151 | 14782 | -0.039 | -0.5684 | No | ||
| 176 | CCL7 | 20743 | 14892 | -0.043 | -0.5731 | No | ||
| 177 | IL8RB | 8752 4533 | 15028 | -0.048 | -0.5790 | No | ||
| 178 | EGFR | 1329 20944 | 15177 | -0.055 | -0.5855 | No | ||
| 179 | IFNG | 19869 | 15219 | -0.057 | -0.5860 | No | ||
| 180 | TNFRSF1A | 1181 10206 | 15227 | -0.057 | -0.5847 | No | ||
| 181 | CLCF1 | 12160 3742 | 15257 | -0.059 | -0.5846 | No | ||
| 182 | CCR6 | 1505 23390 | 15278 | -0.060 | -0.5839 | No | ||
| 183 | IFNAR2 | 22705 1699 | 15300 | -0.062 | -0.5832 | No | ||
| 184 | CCL25 | 9789 3848 | 15303 | -0.062 | -0.5815 | No | ||
| 185 | ACVR1B | 4335 22353 | 15353 | -0.065 | -0.5823 | No | ||
| 186 | IL2RG | 24096 | 15358 | -0.066 | -0.5806 | No | ||
| 187 | IL1RAP | 1653 1676 4917 1748 1712 9173 | 15369 | -0.067 | -0.5792 | No | ||
| 188 | CSF3R | 8804 8803 2326 4565 | 15454 | -0.074 | -0.5816 | No | ||
| 189 | PF4 | 16800 | 15585 | -0.085 | -0.5862 | No | ||
| 190 | ACVR1 | 4334 | 15671 | -0.092 | -0.5881 | No | ||
| 191 | CD40 | 10207 2961 2968 5783 | 15678 | -0.093 | -0.5857 | No | ||
| 192 | FAS | 23881 3750 | 15769 | -0.103 | -0.5876 | No | ||
| 193 | PRLR | 9620 5291 | 15808 | -0.107 | -0.5865 | No | ||
| 194 | TGFB3 | 10161 | 15910 | -0.122 | -0.5885 | No | ||
| 195 | IFNGR1 | 20083 | 16034 | -0.145 | -0.5909 | No | ||
| 196 | CRLF2 | 16442 | 16067 | -0.154 | -0.5882 | No | ||
| 197 | IL3RA | 22091 | 16307 | -0.207 | -0.5951 | No | ||
| 198 | TNFRSF12A | 23107 | 16459 | -0.243 | -0.5962 | No | ||
| 199 | VEGFC | 10286 | 16726 | -0.325 | -0.6012 | Yes | ||
| 200 | IL17RA | 4914 | 16832 | -0.365 | -0.5962 | Yes | ||
| 201 | IL4R | 18085 | 17054 | -0.452 | -0.5951 | Yes | ||
| 202 | IFNGR2 | 9155 22702 | 17065 | -0.457 | -0.5823 | Yes | ||
| 203 | IL10RB | 9169 | 17191 | -0.504 | -0.5744 | Yes | ||
| 204 | TNFRSF14 | 15650 | 17247 | -0.522 | -0.5621 | Yes | ||
| 205 | TNFRSF18 | 15955 2394 | 17308 | -0.549 | -0.5493 | Yes | ||
| 206 | IL10RA | 19137 | 17318 | -0.555 | -0.5336 | Yes | ||
| 207 | CCR7 | 20254 | 17339 | -0.564 | -0.5183 | Yes | ||
| 208 | IL2RA | 4918 | 17844 | -0.900 | -0.5194 | Yes | ||
| 209 | IL18 | 9172 | 18052 | -1.147 | -0.4972 | Yes | ||
| 210 | TNFRSF13B | 7181 | 18247 | -1.472 | -0.4648 | Yes | ||
| 211 | IL12A | 4913 | 18250 | -1.479 | -0.4217 | Yes | ||
| 212 | TGFB1 | 18332 | 18387 | -1.827 | -0.3758 | Yes | ||
| 213 | LTB | 23259 | 18432 | -1.982 | -0.3204 | Yes | ||
| 214 | TNFRSF19 | 21799 3033 | 18498 | -2.427 | -0.2531 | Yes | ||
| 215 | TNFRSF13C | 22191 | 18504 | -2.490 | -0.1807 | Yes | ||
| 216 | CXCR4 | 13844 | 18537 | -2.753 | -0.1021 | Yes | ||
| 217 | IL7R | 9175 4922 | 18575 | -3.646 | 0.0022 | Yes |

