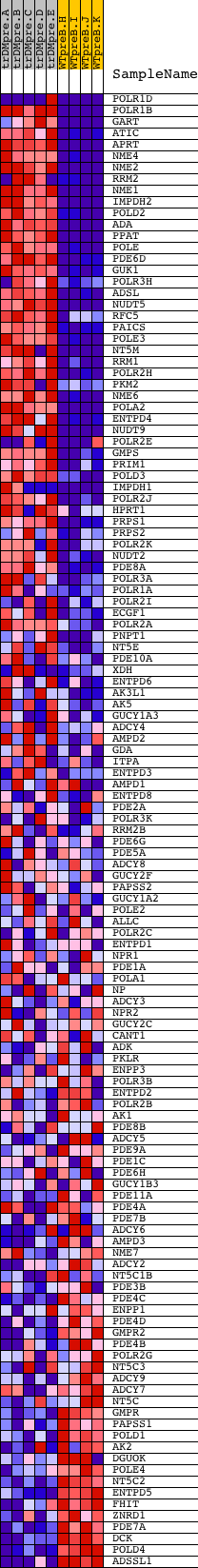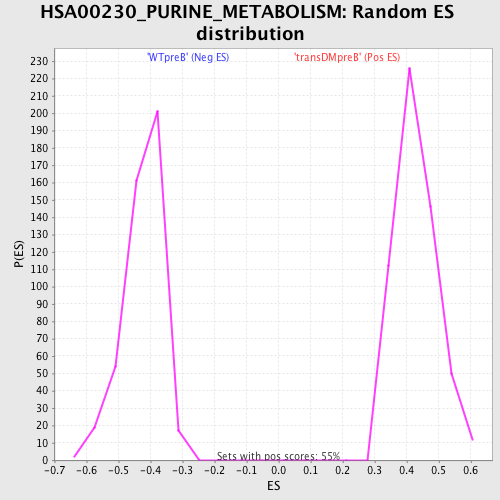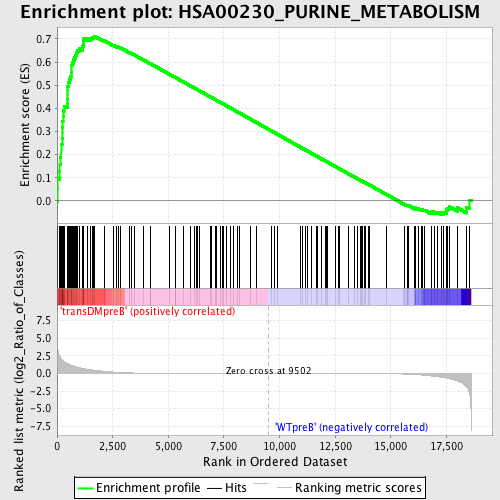
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_WTpreB.phenotype_transDMpreB_versus_WTpreB.cls #transDMpreB_versus_WTpreB |
| Phenotype | phenotype_transDMpreB_versus_WTpreB.cls#transDMpreB_versus_WTpreB |
| Upregulated in class | transDMpreB |
| GeneSet | HSA00230_PURINE_METABOLISM |
| Enrichment Score (ES) | 0.71174264 |
| Normalized Enrichment Score (NES) | 1.6629328 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.016217101 |
| FWER p-Value | 0.04 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | POLR1D | 3593 3658 16623 | 18 | 4.063 | 0.0541 | Yes | ||
| 2 | POLR1B | 14857 | 32 | 3.400 | 0.0994 | Yes | ||
| 3 | GART | 22543 1754 | 121 | 2.400 | 0.1272 | Yes | ||
| 4 | ATIC | 14231 3968 | 128 | 2.362 | 0.1589 | Yes | ||
| 5 | APRT | 8620 | 144 | 2.254 | 0.1886 | Yes | ||
| 6 | NME4 | 23065 1577 | 174 | 2.127 | 0.2158 | Yes | ||
| 7 | NME2 | 9468 | 175 | 2.112 | 0.2445 | Yes | ||
| 8 | RRM2 | 5401 5400 | 222 | 1.924 | 0.2680 | Yes | ||
| 9 | NME1 | 9467 | 229 | 1.906 | 0.2935 | Yes | ||
| 10 | IMPDH2 | 10730 | 230 | 1.901 | 0.3193 | Yes | ||
| 11 | POLD2 | 20537 | 245 | 1.841 | 0.3435 | Yes | ||
| 12 | ADA | 2703 14361 | 266 | 1.775 | 0.3664 | Yes | ||
| 13 | PPAT | 6081 | 271 | 1.764 | 0.3901 | Yes | ||
| 14 | POLE | 16755 | 314 | 1.622 | 0.4098 | Yes | ||
| 15 | PDE6D | 13897 3945 | 467 | 1.413 | 0.4207 | Yes | ||
| 16 | GUK1 | 20432 | 474 | 1.395 | 0.4393 | Yes | ||
| 17 | POLR3H | 13460 8135 | 481 | 1.385 | 0.4578 | Yes | ||
| 18 | ADSL | 4358 | 482 | 1.385 | 0.4765 | Yes | ||
| 19 | NUDT5 | 15121 | 484 | 1.379 | 0.4952 | Yes | ||
| 20 | RFC5 | 13005 7791 | 515 | 1.327 | 0.5115 | Yes | ||
| 21 | PAICS | 16820 | 552 | 1.262 | 0.5267 | Yes | ||
| 22 | POLE3 | 7200 | 616 | 1.176 | 0.5392 | Yes | ||
| 23 | NT5M | 8345 4175 | 637 | 1.153 | 0.5537 | Yes | ||
| 24 | RRM1 | 18163 | 645 | 1.145 | 0.5689 | Yes | ||
| 25 | POLR2H | 10888 | 646 | 1.144 | 0.5844 | Yes | ||
| 26 | PKM2 | 3642 9573 | 675 | 1.102 | 0.5978 | Yes | ||
| 27 | NME6 | 3139 19297 | 724 | 1.055 | 0.6095 | Yes | ||
| 28 | POLA2 | 23988 | 781 | 1.001 | 0.6200 | Yes | ||
| 29 | ENTPD4 | 7449 | 840 | 0.939 | 0.6296 | Yes | ||
| 30 | NUDT9 | 7897 | 873 | 0.916 | 0.6403 | Yes | ||
| 31 | POLR2E | 3325 19699 | 901 | 0.891 | 0.6509 | Yes | ||
| 32 | GMPS | 15578 | 996 | 0.810 | 0.6568 | Yes | ||
| 33 | PRIM1 | 19847 | 1125 | 0.723 | 0.6597 | Yes | ||
| 34 | POLD3 | 17742 | 1142 | 0.708 | 0.6684 | Yes | ||
| 35 | IMPDH1 | 17197 1131 | 1169 | 0.692 | 0.6763 | Yes | ||
| 36 | POLR2J | 16672 | 1186 | 0.681 | 0.6847 | Yes | ||
| 37 | HPRT1 | 2655 24339 408 | 1187 | 0.681 | 0.6939 | Yes | ||
| 38 | PRPS1 | 24233 | 1205 | 0.671 | 0.7021 | Yes | ||
| 39 | PRPS2 | 24003 | 1351 | 0.594 | 0.7023 | Yes | ||
| 40 | POLR2K | 9413 | 1517 | 0.524 | 0.7005 | Yes | ||
| 41 | NUDT2 | 16239 | 1570 | 0.503 | 0.7045 | Yes | ||
| 42 | PDE8A | 18202 | 1648 | 0.471 | 0.7067 | Yes | ||
| 43 | POLR3A | 21900 | 1672 | 0.462 | 0.7117 | Yes | ||
| 44 | POLR1A | 9749 5393 | 2115 | 0.308 | 0.6920 | No | ||
| 45 | POLR2I | 12839 | 2516 | 0.193 | 0.6730 | No | ||
| 46 | ECGF1 | 22160 | 2676 | 0.161 | 0.6666 | No | ||
| 47 | POLR2A | 5394 | 2747 | 0.149 | 0.6648 | No | ||
| 48 | PNPT1 | 20936 | 2830 | 0.133 | 0.6622 | No | ||
| 49 | NT5E | 19360 18702 | 3274 | 0.076 | 0.6392 | No | ||
| 50 | PDE10A | 23387 | 3355 | 0.070 | 0.6358 | No | ||
| 51 | XDH | 22897 | 3482 | 0.060 | 0.6298 | No | ||
| 52 | ENTPD6 | 4496 2678 | 3865 | 0.043 | 0.6098 | No | ||
| 53 | AK3L1 | 8564 | 4194 | 0.032 | 0.5925 | No | ||
| 54 | AK5 | 6042 | 4195 | 0.032 | 0.5929 | No | ||
| 55 | GUCY1A3 | 7216 12245 15310 | 5039 | 0.018 | 0.5475 | No | ||
| 56 | ADCY4 | 2980 21818 | 5328 | 0.015 | 0.5322 | No | ||
| 57 | AMPD2 | 1931 15197 | 5330 | 0.015 | 0.5323 | No | ||
| 58 | GDA | 4764 | 5670 | 0.013 | 0.5142 | No | ||
| 59 | ITPA | 9193 | 5993 | 0.011 | 0.4969 | No | ||
| 60 | ENTPD3 | 19266 | 6168 | 0.010 | 0.4876 | No | ||
| 61 | AMPD1 | 5051 | 6248 | 0.010 | 0.4835 | No | ||
| 62 | ENTPD8 | 12996 | 6288 | 0.009 | 0.4815 | No | ||
| 63 | PDE2A | 18170 | 6392 | 0.009 | 0.4760 | No | ||
| 64 | POLR3K | 12447 7372 | 6896 | 0.007 | 0.4489 | No | ||
| 65 | RRM2B | 6902 6903 11801 | 6930 | 0.007 | 0.4472 | No | ||
| 66 | PDE6G | 20119 | 7120 | 0.006 | 0.4371 | No | ||
| 67 | PDE5A | 15431 | 7151 | 0.006 | 0.4356 | No | ||
| 68 | ADCY8 | 22281 | 7339 | 0.006 | 0.4255 | No | ||
| 69 | GUCY2F | 24045 | 7454 | 0.005 | 0.4194 | No | ||
| 70 | PAPSS2 | 6258 | 7461 | 0.005 | 0.4192 | No | ||
| 71 | GUCY1A2 | 19580 | 7613 | 0.005 | 0.4111 | No | ||
| 72 | POLE2 | 21053 | 7815 | 0.004 | 0.4003 | No | ||
| 73 | ALLC | 21093 2130 | 7948 | 0.004 | 0.3932 | No | ||
| 74 | POLR2C | 9750 | 8110 | 0.003 | 0.3845 | No | ||
| 75 | ENTPD1 | 4495 | 8187 | 0.003 | 0.3804 | No | ||
| 76 | NPR1 | 9480 | 8679 | 0.002 | 0.3539 | No | ||
| 77 | PDE1A | 9541 5234 | 8950 | 0.001 | 0.3393 | No | ||
| 78 | POLA1 | 24112 | 9653 | -0.000 | 0.3014 | No | ||
| 79 | NP | 22027 9597 5273 5274 | 9773 | -0.001 | 0.2949 | No | ||
| 80 | ADCY3 | 21329 894 | 9899 | -0.001 | 0.2882 | No | ||
| 81 | NPR2 | 16230 | 10933 | -0.004 | 0.2324 | No | ||
| 82 | GUCY2C | 1140 16954 | 11013 | -0.004 | 0.2281 | No | ||
| 83 | CANT1 | 13304 | 11171 | -0.004 | 0.2197 | No | ||
| 84 | ADK | 3302 3454 8555 | 11260 | -0.005 | 0.2150 | No | ||
| 85 | PKLR | 1850 15545 | 11427 | -0.005 | 0.2061 | No | ||
| 86 | ENPP3 | 19803 | 11641 | -0.006 | 0.1947 | No | ||
| 87 | POLR3B | 12875 | 11695 | -0.006 | 0.1919 | No | ||
| 88 | ENTPD2 | 8713 | 11872 | -0.007 | 0.1824 | No | ||
| 89 | POLR2B | 16817 | 12059 | -0.007 | 0.1725 | No | ||
| 90 | AK1 | 4363 | 12126 | -0.007 | 0.1690 | No | ||
| 91 | PDE8B | 5756 | 12164 | -0.008 | 0.1671 | No | ||
| 92 | ADCY5 | 10312 | 12491 | -0.009 | 0.1496 | No | ||
| 93 | PDE9A | 23301 | 12652 | -0.010 | 0.1411 | No | ||
| 94 | PDE1C | 5235 | 12671 | -0.010 | 0.1403 | No | ||
| 95 | PDE6H | 17257 | 13103 | -0.013 | 0.1171 | No | ||
| 96 | GUCY1B3 | 15311 | 13349 | -0.014 | 0.1041 | No | ||
| 97 | PDE11A | 10805 | 13519 | -0.016 | 0.0951 | No | ||
| 98 | PDE4A | 3037 19542 3083 3023 | 13615 | -0.017 | 0.0902 | No | ||
| 99 | PDE7B | 19807 | 13687 | -0.018 | 0.0866 | No | ||
| 100 | ADCY6 | 22139 2283 8551 | 13729 | -0.018 | 0.0846 | No | ||
| 101 | AMPD3 | 4383 2110 | 13800 | -0.019 | 0.0811 | No | ||
| 102 | NME7 | 926 4101 14070 3986 3946 931 | 13857 | -0.020 | 0.0784 | No | ||
| 103 | ADCY2 | 21412 | 14014 | -0.022 | 0.0702 | No | ||
| 104 | NT5C1B | 21315 | 14053 | -0.022 | 0.0685 | No | ||
| 105 | PDE3B | 5236 | 14827 | -0.040 | 0.0272 | No | ||
| 106 | PDE4C | 8481 | 15624 | -0.088 | -0.0147 | No | ||
| 107 | ENPP1 | 19804 | 15742 | -0.101 | -0.0196 | No | ||
| 108 | PDE4D | 10722 6235 | 15749 | -0.102 | -0.0186 | No | ||
| 109 | GMPR2 | 2711 | 15809 | -0.107 | -0.0203 | No | ||
| 110 | PDE4B | 9543 | 16079 | -0.158 | -0.0327 | No | ||
| 111 | POLR2G | 23753 | 16110 | -0.165 | -0.0321 | No | ||
| 112 | NT5C3 | 17136 | 16113 | -0.167 | -0.0300 | No | ||
| 113 | ADCY9 | 22680 | 16230 | -0.191 | -0.0337 | No | ||
| 114 | ADCY7 | 8552 448 | 16369 | -0.222 | -0.0381 | No | ||
| 115 | NT5C | 20151 | 16435 | -0.236 | -0.0384 | No | ||
| 116 | GMPR | 21651 | 16520 | -0.264 | -0.0394 | No | ||
| 117 | PAPSS1 | 15419 | 16824 | -0.362 | -0.0509 | No | ||
| 118 | POLD1 | 17847 | 16839 | -0.368 | -0.0466 | No | ||
| 119 | AK2 | 8563 2479 16073 | 16960 | -0.419 | -0.0475 | No | ||
| 120 | DGUOK | 17099 1041 | 17097 | -0.468 | -0.0485 | No | ||
| 121 | POLE4 | 12441 | 17279 | -0.536 | -0.0510 | No | ||
| 122 | NT5C2 | 3768 8052 | 17383 | -0.587 | -0.0486 | No | ||
| 123 | ENTPD5 | 4497 | 17497 | -0.654 | -0.0459 | No | ||
| 124 | FHIT | 21920 | 17514 | -0.663 | -0.0378 | No | ||
| 125 | ZNRD1 | 1491 22987 | 17560 | -0.693 | -0.0308 | No | ||
| 126 | PDE7A | 9544 1788 | 17646 | -0.750 | -0.0252 | No | ||
| 127 | DCK | 16808 | 17984 | -1.054 | -0.0292 | No | ||
| 128 | POLD4 | 12822 | 18404 | -1.867 | -0.0266 | No | ||
| 129 | ADSSL1 | 21136 2087 | 18542 | -2.803 | 0.0040 | No |