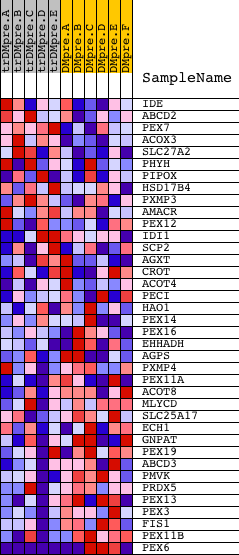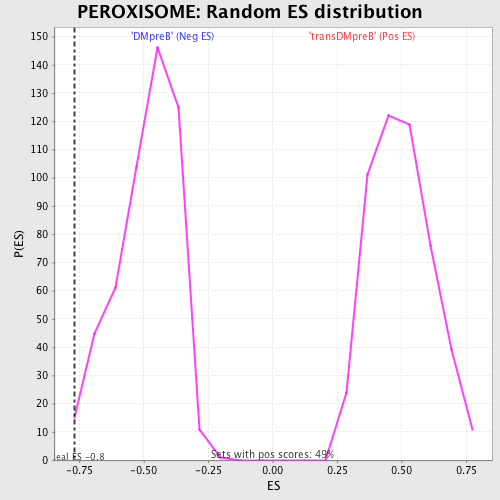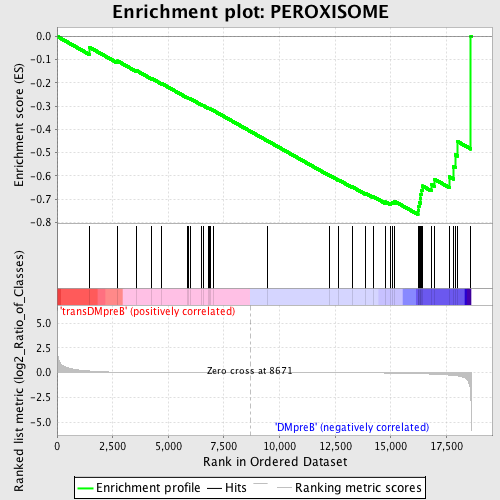
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMpreB_versus_DMpreB.cls#transDMpreB_versus_DMpreB_repos |
| Upregulated in class | DMpreB |
| GeneSet | PEROXISOME |
| Enrichment Score (ES) | -0.7662733 |
| Normalized Enrichment Score (NES) | -1.5687301 |
| Nominal p-value | 0.011811024 |
| FDR q-value | 0.44533896 |
| FWER p-Value | 0.991 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | IDE | 4893 | 1464 | 0.165 | -0.0475 | No | ||
| 2 | ABCD2 | 11204 6494 | 2712 | 0.046 | -0.1059 | No | ||
| 3 | PEX7 | 19808 3305 | 3574 | 0.024 | -0.1477 | No | ||
| 4 | ACOX3 | 3612 16867 | 4252 | 0.015 | -0.1812 | No | ||
| 5 | SLC27A2 | 6468 | 4706 | 0.012 | -0.2033 | No | ||
| 6 | PHYH | 15124 | 5849 | 0.007 | -0.2635 | No | ||
| 7 | PIPOX | 20340 | 5918 | 0.007 | -0.2659 | No | ||
| 8 | HSD17B4 | 23560 | 5988 | 0.006 | -0.2684 | No | ||
| 9 | PXMP3 | 15361 1786 15381 | 6467 | 0.005 | -0.2932 | No | ||
| 10 | AMACR | 22506 | 6566 | 0.005 | -0.2976 | No | ||
| 11 | PEX12 | 20321 | 6789 | 0.004 | -0.3088 | No | ||
| 12 | IDI1 | 11452 | 6859 | 0.004 | -0.3117 | No | ||
| 13 | SCP2 | 5415 2422 9785 5416 | 6887 | 0.004 | -0.3125 | No | ||
| 14 | AGXT | 14178 | 7040 | 0.003 | -0.3200 | No | ||
| 15 | CROT | 13165 | 9446 | -0.002 | -0.4491 | No | ||
| 16 | ACOT4 | 21214 | 12258 | -0.010 | -0.5986 | No | ||
| 17 | PECI | 10754 | 12660 | -0.012 | -0.6179 | No | ||
| 18 | HAO1 | 14418 | 13259 | -0.016 | -0.6472 | No | ||
| 19 | PEX14 | 15668 | 13879 | -0.021 | -0.6765 | No | ||
| 20 | PEX16 | 2681 | 14226 | -0.025 | -0.6903 | No | ||
| 21 | EHHADH | 22635 | 14760 | -0.035 | -0.7123 | No | ||
| 22 | AGPS | 14979 | 15000 | -0.041 | -0.7173 | No | ||
| 23 | PXMP4 | 12226 | 15069 | -0.043 | -0.7129 | No | ||
| 24 | PEX11A | 1056 5241 | 15165 | -0.045 | -0.7094 | No | ||
| 25 | ACOT8 | 14344 2749 | 16223 | -0.093 | -0.7485 | Yes | ||
| 26 | MLYCD | 7143 | 16256 | -0.096 | -0.7320 | Yes | ||
| 27 | SLC25A17 | 22197 | 16273 | -0.097 | -0.7145 | Yes | ||
| 28 | ECH1 | 18316 | 16332 | -0.101 | -0.6985 | Yes | ||
| 29 | GNPAT | 18420 | 16348 | -0.102 | -0.6800 | Yes | ||
| 30 | PEX19 | 9673 | 16386 | -0.105 | -0.6621 | Yes | ||
| 31 | ABCD3 | 5337 | 16422 | -0.108 | -0.6436 | Yes | ||
| 32 | PMVK | 15539 | 16825 | -0.145 | -0.6377 | Yes | ||
| 33 | PRDX5 | 7053 3732 | 16960 | -0.160 | -0.6145 | Yes | ||
| 34 | PEX13 | 20513 | 17630 | -0.243 | -0.6045 | Yes | ||
| 35 | PEX3 | 7134 | 17808 | -0.278 | -0.5612 | Yes | ||
| 36 | FIS1 | 16669 3579 | 17903 | -0.299 | -0.5095 | Yes | ||
| 37 | PEX11B | 9550 | 18008 | -0.327 | -0.4530 | Yes | ||
| 38 | PEX6 | 5900 10331 | 18599 | -2.558 | 0.0009 | Yes |