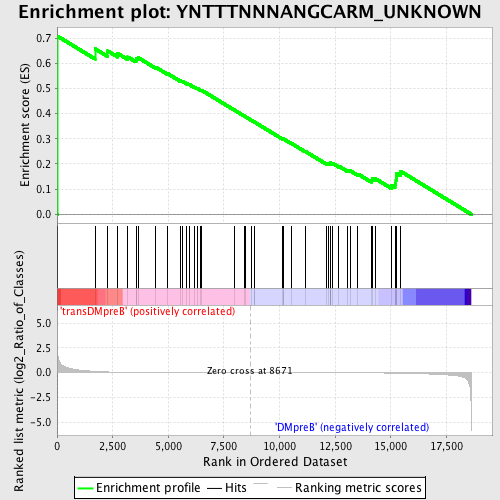
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMpreB_versus_DMpreB.cls#transDMpreB_versus_DMpreB_repos |
| Upregulated in class | transDMpreB |
| GeneSet | YNTTTNNNANGCARM_UNKNOWN |
| Enrichment Score (ES) | 0.7080401 |
| Normalized Enrichment Score (NES) | 1.4838094 |
| Nominal p-value | 0.027559055 |
| FDR q-value | 0.32243055 |
| FWER p-Value | 0.986 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | IL10 | 14145 1510 1553 22902 | 14 | 2.242 | 0.7080 | Yes | ||
| 2 | LEMD2 | 5893 | 1706 | 0.129 | 0.6576 | No | ||
| 3 | KCTD12 | 6240 | 2273 | 0.070 | 0.6491 | No | ||
| 4 | NRF1 | 17499 | 2724 | 0.046 | 0.6394 | No | ||
| 5 | C1S | 6950 6949 | 3178 | 0.032 | 0.6251 | No | ||
| 6 | PDE4D | 10722 6235 | 3548 | 0.024 | 0.6129 | No | ||
| 7 | OMG | 5209 | 3572 | 0.024 | 0.6192 | No | ||
| 8 | PVRL1 | 3006 19482 12211 7192 | 3636 | 0.023 | 0.6230 | No | ||
| 9 | FGD4 | 10306 5870 1717 1669 1700 | 4433 | 0.014 | 0.5845 | No | ||
| 10 | MSX2 | 21461 | 4979 | 0.010 | 0.5585 | No | ||
| 11 | FGF14 | 21716 3005 3463 | 5530 | 0.008 | 0.5314 | No | ||
| 12 | LDB2 | 16538 4989 | 5634 | 0.008 | 0.5282 | No | ||
| 13 | GABRG1 | 16521 | 5830 | 0.007 | 0.5199 | No | ||
| 14 | GABRA1 | 20492 | 5938 | 0.006 | 0.5162 | No | ||
| 15 | KIF13A | 21475 | 6166 | 0.006 | 0.5058 | No | ||
| 16 | KLHL5 | 3631 7753 | 6293 | 0.005 | 0.5007 | No | ||
| 17 | SYNPR | 22094 | 6465 | 0.005 | 0.4931 | No | ||
| 18 | EYA1 | 4695 4061 | 6502 | 0.005 | 0.4927 | No | ||
| 19 | TLL2 | 23680 | 7952 | 0.001 | 0.4151 | No | ||
| 20 | B3GNT7 | 14202 | 8410 | 0.001 | 0.3906 | No | ||
| 21 | NRK | 6559 | 8480 | 0.000 | 0.3870 | No | ||
| 22 | FGF12 | 1723 22621 | 8735 | -0.000 | 0.3734 | No | ||
| 23 | VAX1 | 23636 | 8882 | -0.000 | 0.3657 | No | ||
| 24 | DCX | 24040 8841 4604 | 8883 | -0.000 | 0.3658 | No | ||
| 25 | GCDH | 3913 18812 | 10110 | -0.003 | 0.3008 | No | ||
| 26 | EVX1 | 17442 | 10157 | -0.003 | 0.2994 | No | ||
| 27 | LRRTM1 | 17408 | 10170 | -0.003 | 0.2998 | No | ||
| 28 | MAFB | 14367 | 10526 | -0.004 | 0.2820 | No | ||
| 29 | SMPX | 24220 | 11161 | -0.006 | 0.2497 | No | ||
| 30 | VDR | 5852 10285 | 12119 | -0.009 | 0.2010 | No | ||
| 31 | DMD | 24295 2647 | 12181 | -0.009 | 0.2007 | No | ||
| 32 | DNAH5 | 8470 | 12271 | -0.010 | 0.1990 | No | ||
| 33 | HOXC5 | 22340 | 12284 | -0.010 | 0.2015 | No | ||
| 34 | HCRTR2 | 6907 | 12289 | -0.010 | 0.2044 | No | ||
| 35 | ELAVL2 | 15839 | 12384 | -0.010 | 0.2026 | No | ||
| 36 | GPM6A | 3834 10618 | 12666 | -0.012 | 0.1911 | No | ||
| 37 | ADAMTS5 | 6207 | 13035 | -0.014 | 0.1757 | No | ||
| 38 | SFRP2 | 15564 | 13173 | -0.015 | 0.1731 | No | ||
| 39 | KCNA2 | 15458 | 13523 | -0.018 | 0.1599 | No | ||
| 40 | ETV5 | 22630 | 14115 | -0.024 | 0.1356 | No | ||
| 41 | GRK5 | 9036 23814 | 14155 | -0.024 | 0.1412 | No | ||
| 42 | HOXB6 | 20688 | 14311 | -0.027 | 0.1413 | No | ||
| 43 | ETV6 | 17264 | 15027 | -0.042 | 0.1159 | No | ||
| 44 | HOXC4 | 4864 | 15189 | -0.046 | 0.1218 | No | ||
| 45 | GNAI1 | 9024 | 15230 | -0.048 | 0.1346 | No | ||
| 46 | CNN3 | 12983 | 15239 | -0.048 | 0.1493 | No | ||
| 47 | CDX2 | 16289 | 15271 | -0.049 | 0.1631 | No | ||
| 48 | GPR158 | 149 | 15441 | -0.054 | 0.1710 | No |

