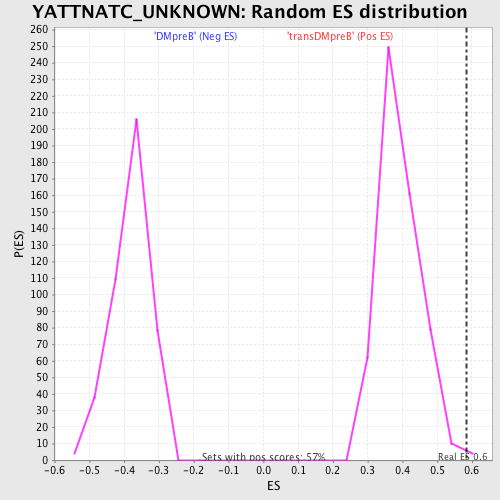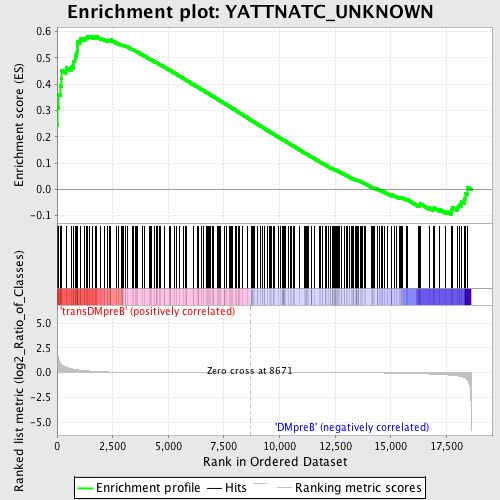
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMpreB_versus_DMpreB.cls#transDMpreB_versus_DMpreB_repos |
| Upregulated in class | transDMpreB |
| GeneSet | YATTNATC_UNKNOWN |
| Enrichment Score (ES) | 0.58301336 |
| Normalized Enrichment Score (NES) | 1.4902772 |
| Nominal p-value | 0.003539823 |
| FDR q-value | 0.33407745 |
| FWER p-Value | 0.981 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ATP2A3 | 20795 1314 | 0 | 6.225 | 0.2464 | Yes | ||
| 2 | ANKMY2 | 5721 21288 | 35 | 1.628 | 0.3090 | Yes | ||
| 3 | ARHGAP8 | 4268 2196 | 73 | 1.369 | 0.3612 | Yes | ||
| 4 | TAF15 | 20731 | 152 | 0.923 | 0.3935 | Yes | ||
| 5 | MAST2 | 9423 | 199 | 0.797 | 0.4226 | Yes | ||
| 6 | PPP1R10 | 6972 11960 6971 | 214 | 0.774 | 0.4524 | Yes | ||
| 7 | MAP4K2 | 6457 | 400 | 0.537 | 0.4636 | Yes | ||
| 8 | NFIX | 9458 | 630 | 0.396 | 0.4668 | Yes | ||
| 9 | DYRK1A | 4649 | 738 | 0.345 | 0.4747 | Yes | ||
| 10 | STAT3 | 5525 9906 | 746 | 0.342 | 0.4878 | Yes | ||
| 11 | PBXIP1 | 6028 | 805 | 0.321 | 0.4974 | Yes | ||
| 12 | ADD3 | 23812 3694 | 823 | 0.317 | 0.5090 | Yes | ||
| 13 | ATP2A2 | 4421 3481 | 863 | 0.303 | 0.5189 | Yes | ||
| 14 | UBE4B | 2433 15672 | 894 | 0.294 | 0.5289 | Yes | ||
| 15 | GOLGA1 | 14595 | 904 | 0.292 | 0.5399 | Yes | ||
| 16 | BCL9L | 19475 | 907 | 0.290 | 0.5513 | Yes | ||
| 17 | ARPC2 | 14225 | 929 | 0.283 | 0.5614 | Yes | ||
| 18 | ZMYND17 | 21909 | 1030 | 0.253 | 0.5660 | Yes | ||
| 19 | E2F1 | 14384 | 1040 | 0.251 | 0.5754 | Yes | ||
| 20 | PRICKLE1 | 8379 | 1226 | 0.206 | 0.5735 | Yes | ||
| 21 | BZW2 | 21081 | 1300 | 0.190 | 0.5771 | Yes | ||
| 22 | AP1GBP1 | 20725 | 1356 | 0.179 | 0.5812 | Yes | ||
| 23 | AXUD1 | 18967 | 1445 | 0.167 | 0.5830 | Yes | ||
| 24 | SNRPF | 7645 | 1583 | 0.147 | 0.5814 | No | ||
| 25 | H3F3B | 4839 | 1708 | 0.128 | 0.5797 | No | ||
| 26 | GPX1 | 19310 | 1767 | 0.122 | 0.5814 | No | ||
| 27 | LUC7L | 47 1520 | 1969 | 0.096 | 0.5742 | No | ||
| 28 | PITPNC1 | 7759 | 2112 | 0.082 | 0.5697 | No | ||
| 29 | CPNE1 | 11186 | 2252 | 0.071 | 0.5650 | No | ||
| 30 | KCTD12 | 6240 | 2273 | 0.070 | 0.5666 | No | ||
| 31 | FBXL19 | 6134 | 2360 | 0.064 | 0.5645 | No | ||
| 32 | AGTR2 | 24365 | 2380 | 0.063 | 0.5660 | No | ||
| 33 | SP6 | 8178 13678 13679 | 2411 | 0.061 | 0.5668 | No | ||
| 34 | APP | 4402 | 2419 | 0.061 | 0.5688 | No | ||
| 35 | AXIN2 | 20618 | 2667 | 0.048 | 0.5572 | No | ||
| 36 | NSD1 | 2134 5197 | 2777 | 0.044 | 0.5530 | No | ||
| 37 | BNC2 | 15850 | 2906 | 0.040 | 0.5477 | No | ||
| 38 | HOXB7 | 666 9110 | 2944 | 0.039 | 0.5472 | No | ||
| 39 | SLC16A10 | 7813 | 2948 | 0.039 | 0.5485 | No | ||
| 40 | PTPRJ | 9664 | 2977 | 0.037 | 0.5485 | No | ||
| 41 | CFL2 | 2155 21069 | 3082 | 0.034 | 0.5442 | No | ||
| 42 | FOXP2 | 4312 17523 8523 | 3091 | 0.034 | 0.5451 | No | ||
| 43 | S100A16 | 15530 | 3162 | 0.032 | 0.5426 | No | ||
| 44 | HOXB9 | 20689 | 3366 | 0.028 | 0.5326 | No | ||
| 45 | MBNL2 | 21932 | 3427 | 0.026 | 0.5304 | No | ||
| 46 | DNAJB4 | 12451 1916 1832 | 3542 | 0.024 | 0.5251 | No | ||
| 47 | OMG | 5209 | 3572 | 0.024 | 0.5245 | No | ||
| 48 | MBL2 | 23886 | 3590 | 0.024 | 0.5245 | No | ||
| 49 | ARF4 | 22072 | 3859 | 0.020 | 0.5107 | No | ||
| 50 | SCGB3A2 | 23570 | 3936 | 0.019 | 0.5073 | No | ||
| 51 | ZNRF1 | 9306 | 4174 | 0.016 | 0.4950 | No | ||
| 52 | UCN2 | 10308 | 4177 | 0.016 | 0.4956 | No | ||
| 53 | IL3 | 20453 | 4232 | 0.016 | 0.4932 | No | ||
| 54 | HOXA7 | 4862 | 4398 | 0.014 | 0.4848 | No | ||
| 55 | SEMA3A | 16919 | 4448 | 0.014 | 0.4827 | No | ||
| 56 | CSF3 | 1394 20671 | 4462 | 0.014 | 0.4825 | No | ||
| 57 | SNX26 | 10582 | 4492 | 0.013 | 0.4815 | No | ||
| 58 | SLC5A2 | 18057 10905 | 4607 | 0.013 | 0.4758 | No | ||
| 59 | FGF5 | 16785 | 4635 | 0.012 | 0.4748 | No | ||
| 60 | DACH1 | 4595 3371 | 4821 | 0.011 | 0.4652 | No | ||
| 61 | MYOD1 | 18237 | 4834 | 0.011 | 0.4650 | No | ||
| 62 | F9 | 24328 | 5042 | 0.010 | 0.4541 | No | ||
| 63 | ROBO3 | 9705 | 5076 | 0.010 | 0.4527 | No | ||
| 64 | MYOG | 14129 | 5107 | 0.010 | 0.4514 | No | ||
| 65 | POLR3D | 21760 12456 | 5283 | 0.009 | 0.4423 | No | ||
| 66 | COL3A1 | 14253 8765 | 5377 | 0.009 | 0.4375 | No | ||
| 67 | COL4A6 | 8248 2552 | 5496 | 0.008 | 0.4314 | No | ||
| 68 | FLRT3 | 7730 12930 2673 | 5660 | 0.007 | 0.4229 | No | ||
| 69 | DOCK7 | 12506 | 5779 | 0.007 | 0.4167 | No | ||
| 70 | KCNQ4 | 12247 2533 | 5818 | 0.007 | 0.4149 | No | ||
| 71 | EGR3 | 4656 | 5831 | 0.007 | 0.4145 | No | ||
| 72 | KCNIP2 | 23655 3725 | 6132 | 0.006 | 0.3984 | No | ||
| 73 | EHF | 4657 | 6135 | 0.006 | 0.3986 | No | ||
| 74 | GABRB3 | 4747 | 6324 | 0.005 | 0.3885 | No | ||
| 75 | TJP2 | 5761 3699 | 6336 | 0.005 | 0.3882 | No | ||
| 76 | HOXD3 | 9115 | 6372 | 0.005 | 0.3865 | No | ||
| 77 | EYA1 | 4695 4061 | 6502 | 0.005 | 0.3796 | No | ||
| 78 | TTYH1 | 2496 7176 | 6557 | 0.005 | 0.3769 | No | ||
| 79 | IL21 | 12241 | 6694 | 0.004 | 0.3696 | No | ||
| 80 | PNOC | 21783 | 6752 | 0.004 | 0.3667 | No | ||
| 81 | IL23A | 19598 | 6799 | 0.004 | 0.3644 | No | ||
| 82 | CRH | 8784 | 6828 | 0.004 | 0.3630 | No | ||
| 83 | WNT4 | 16025 | 6898 | 0.004 | 0.3594 | No | ||
| 84 | THBS1 | 5748 14910 | 7006 | 0.004 | 0.3537 | No | ||
| 85 | MOXD1 | 20069 | 7007 | 0.004 | 0.3538 | No | ||
| 86 | CA4 | 20723 | 7027 | 0.003 | 0.3529 | No | ||
| 87 | GLRA1 | 9021 | 7206 | 0.003 | 0.3434 | No | ||
| 88 | RASGEF1B | 11515 16469 | 7244 | 0.003 | 0.3415 | No | ||
| 89 | CALD1 | 4273 8463 | 7286 | 0.003 | 0.3394 | No | ||
| 90 | HOXB3 | 20685 | 7353 | 0.003 | 0.3359 | No | ||
| 91 | PI15 | 13586 | 7508 | 0.002 | 0.3276 | No | ||
| 92 | NAP1L5 | 17134 | 7523 | 0.002 | 0.3269 | No | ||
| 93 | PRL | 21690 | 7530 | 0.002 | 0.3267 | No | ||
| 94 | PPP1R14D | 14471 | 7635 | 0.002 | 0.3211 | No | ||
| 95 | GUCY2F | 24045 | 7763 | 0.002 | 0.3143 | No | ||
| 96 | RAB3C | 21351 | 7810 | 0.002 | 0.3118 | No | ||
| 97 | KCNA3 | 15460 | 7821 | 0.002 | 0.3113 | No | ||
| 98 | NKD1 | 8228 | 7868 | 0.002 | 0.3089 | No | ||
| 99 | TGM5 | 2687 14460 | 7996 | 0.001 | 0.3020 | No | ||
| 100 | FOLR2 | 17732 | 8083 | 0.001 | 0.2974 | No | ||
| 101 | ANGPTL3 | 16171 | 8143 | 0.001 | 0.2942 | No | ||
| 102 | COL19A1 | 4541 | 8174 | 0.001 | 0.2926 | No | ||
| 103 | APBA2BP | 14385 | 8199 | 0.001 | 0.2914 | No | ||
| 104 | CSF2 | 20454 | 8345 | 0.001 | 0.2835 | No | ||
| 105 | TBXAS1 | 17480 1105 | 8565 | 0.000 | 0.2716 | No | ||
| 106 | GRIN2B | 16957 | 8734 | -0.000 | 0.2624 | No | ||
| 107 | RARB | 3011 21915 | 8748 | -0.000 | 0.2617 | No | ||
| 108 | TCF1 | 16416 | 8752 | -0.000 | 0.2616 | No | ||
| 109 | SOAT2 | 22348 10302 | 8787 | -0.000 | 0.2597 | No | ||
| 110 | AQP5 | 22365 | 8799 | -0.000 | 0.2592 | No | ||
| 111 | KCNH2 | 16592 | 8828 | -0.000 | 0.2576 | No | ||
| 112 | DCX | 24040 8841 4604 | 8883 | -0.000 | 0.2547 | No | ||
| 113 | BHLHB5 | 15629 | 8991 | -0.001 | 0.2489 | No | ||
| 114 | RBPMS | 5369 | 9021 | -0.001 | 0.2474 | No | ||
| 115 | MITF | 17349 | 9127 | -0.001 | 0.2417 | No | ||
| 116 | ZIC1 | 19040 | 9234 | -0.001 | 0.2360 | No | ||
| 117 | TINAG | 3141 19060 | 9253 | -0.001 | 0.2350 | No | ||
| 118 | ANK2 | 4275 | 9300 | -0.001 | 0.2326 | No | ||
| 119 | DKK4 | 18653 | 9339 | -0.001 | 0.2306 | No | ||
| 120 | NXPH4 | 19602 | 9437 | -0.002 | 0.2254 | No | ||
| 121 | HNMT | 14672 | 9543 | -0.002 | 0.2197 | No | ||
| 122 | SLITRK5 | 21939 | 9587 | -0.002 | 0.2175 | No | ||
| 123 | PKD2L2 | 898 | 9646 | -0.002 | 0.2144 | No | ||
| 124 | SYT1 | 5565 | 9705 | -0.002 | 0.2113 | No | ||
| 125 | PCSK5 | 23714 | 9724 | -0.002 | 0.2104 | No | ||
| 126 | NPTX2 | 16630 | 9769 | -0.002 | 0.2081 | No | ||
| 127 | BMPR1B | 8661 15152 4451 | 9933 | -0.003 | 0.1994 | No | ||
| 128 | PKP4 | 2822 15008 | 10061 | -0.003 | 0.1926 | No | ||
| 129 | CHM | 4521 | 10113 | -0.003 | 0.1899 | No | ||
| 130 | SPAG6 | 22650 | 10114 | -0.003 | 0.1900 | No | ||
| 131 | ZIC2 | 21927 | 10130 | -0.003 | 0.1894 | No | ||
| 132 | EVX1 | 17442 | 10157 | -0.003 | 0.1881 | No | ||
| 133 | LRRTM1 | 17408 | 10170 | -0.003 | 0.1876 | No | ||
| 134 | EN1 | 387 14155 | 10175 | -0.003 | 0.1875 | No | ||
| 135 | HOXD10 | 14983 | 10203 | -0.003 | 0.1861 | No | ||
| 136 | TCF7L2 | 10048 5646 | 10213 | -0.003 | 0.1858 | No | ||
| 137 | GPR151 | 23446 | 10257 | -0.004 | 0.1836 | No | ||
| 138 | ERG | 1686 8915 | 10399 | -0.004 | 0.1761 | No | ||
| 139 | STMN2 | 15638 | 10500 | -0.004 | 0.1708 | No | ||
| 140 | CRY1 | 19662 | 10514 | -0.004 | 0.1702 | No | ||
| 141 | ARMET | 19005 | 10634 | -0.004 | 0.1639 | No | ||
| 142 | SLC12A1 | 14874 2772 2943 | 10636 | -0.005 | 0.1641 | No | ||
| 143 | NR3C2 | 18562 13798 | 10678 | -0.005 | 0.1620 | No | ||
| 144 | TRPC6 | 19565 | 10900 | -0.005 | 0.1502 | No | ||
| 145 | KCTD8 | 16523 | 11132 | -0.006 | 0.1378 | No | ||
| 146 | SMPX | 24220 | 11161 | -0.006 | 0.1365 | No | ||
| 147 | MEOX2 | 21285 | 11191 | -0.006 | 0.1352 | No | ||
| 148 | SOX2 | 9849 15612 | 11224 | -0.006 | 0.1337 | No | ||
| 149 | SSTR1 | 21264 | 11250 | -0.006 | 0.1326 | No | ||
| 150 | NPAS3 | 2077 21273 | 11296 | -0.006 | 0.1304 | No | ||
| 151 | PAX2 | 9530 24416 | 11416 | -0.007 | 0.1241 | No | ||
| 152 | POU6F2 | 21539 | 11421 | -0.007 | 0.1242 | No | ||
| 153 | CHES1 | 21007 11200 | 11432 | -0.007 | 0.1239 | No | ||
| 154 | SLC43A3 | 2824 14973 | 11455 | -0.007 | 0.1230 | No | ||
| 155 | ABLIM2 | 16866 3605 | 11579 | -0.007 | 0.1166 | No | ||
| 156 | GNAO1 | 4785 3829 | 11779 | -0.008 | 0.1060 | No | ||
| 157 | STAT5B | 20222 | 11851 | -0.008 | 0.1025 | No | ||
| 158 | SOX1 | 18685 | 11935 | -0.008 | 0.0983 | No | ||
| 159 | HAPLN1 | 21592 | 11940 | -0.008 | 0.0984 | No | ||
| 160 | IGF1 | 3352 9156 3409 | 12045 | -0.009 | 0.0931 | No | ||
| 161 | EMX2 | 23806 | 12077 | -0.009 | 0.0918 | No | ||
| 162 | C1QTNF3 | 2224 22505 | 12111 | -0.009 | 0.0903 | No | ||
| 163 | DMD | 24295 2647 | 12181 | -0.009 | 0.0869 | No | ||
| 164 | HCRTR2 | 6907 | 12289 | -0.010 | 0.0815 | No | ||
| 165 | NANOS1 | 6858 | 12290 | -0.010 | 0.0819 | No | ||
| 166 | HOXA2 | 17151 | 12362 | -0.010 | 0.0784 | No | ||
| 167 | LMO4 | 15151 | 12368 | -0.010 | 0.0786 | No | ||
| 168 | ELAVL2 | 15839 | 12384 | -0.010 | 0.0781 | No | ||
| 169 | GC | 16486 | 12402 | -0.010 | 0.0776 | No | ||
| 170 | IRX3 | 18793 | 12464 | -0.011 | 0.0747 | No | ||
| 171 | EIF4E | 15403 1827 8890 | 12487 | -0.011 | 0.0740 | No | ||
| 172 | DSG3 | 8866 | 12496 | -0.011 | 0.0740 | No | ||
| 173 | LRRC16 | 21515 21514 | 12526 | -0.011 | 0.0728 | No | ||
| 174 | LRP8 | 16149 2425 | 12534 | -0.011 | 0.0729 | No | ||
| 175 | GPC3 | 24151 | 12569 | -0.011 | 0.0715 | No | ||
| 176 | NEUROD6 | 17138 | 12592 | -0.011 | 0.0707 | No | ||
| 177 | GRIK3 | 16083 | 12613 | -0.011 | 0.0701 | No | ||
| 178 | CTBP2 | 17591 1137 | 12630 | -0.012 | 0.0697 | No | ||
| 179 | DLC1 | 11931 3855 | 12696 | -0.012 | 0.0666 | No | ||
| 180 | HOXA1 | 1005 17152 | 12766 | -0.012 | 0.0633 | No | ||
| 181 | SSPN | 9246 | 12794 | -0.012 | 0.0623 | No | ||
| 182 | QKI | 23128 5340 | 12898 | -0.013 | 0.0573 | No | ||
| 183 | SLC38A3 | 13319 | 12915 | -0.013 | 0.0569 | No | ||
| 184 | CALB1 | 4469 | 13020 | -0.014 | 0.0518 | No | ||
| 185 | TRERF1 | 1546 23213 | 13044 | -0.014 | 0.0511 | No | ||
| 186 | SLC13A1 | 17205 | 13120 | -0.015 | 0.0476 | No | ||
| 187 | GDF7 | 21113 | 13244 | -0.015 | 0.0415 | No | ||
| 188 | PALMD | 15181 | 13274 | -0.016 | 0.0405 | No | ||
| 189 | BRUNOL4 | 2010 4229 | 13325 | -0.016 | 0.0385 | No | ||
| 190 | NEB | 5158 | 13328 | -0.016 | 0.0390 | No | ||
| 191 | A2BP1 | 11212 1652 1689 | 13329 | -0.016 | 0.0396 | No | ||
| 192 | CEBPA | 4514 | 13397 | -0.017 | 0.0366 | No | ||
| 193 | OTOP3 | 20599 | 13448 | -0.017 | 0.0346 | No | ||
| 194 | GFI1 | 16451 | 13461 | -0.017 | 0.0346 | No | ||
| 195 | KIF1B | 15671 2476 4950 15673 | 13462 | -0.017 | 0.0353 | No | ||
| 196 | WNT6 | 5881 | 13475 | -0.017 | 0.0353 | No | ||
| 197 | HOXA10 | 17145 989 | 13494 | -0.017 | 0.0350 | No | ||
| 198 | RFX4 | 7721 3410 | 13532 | -0.018 | 0.0337 | No | ||
| 199 | GRIA3 | 24349 11996 2649 | 13541 | -0.018 | 0.0340 | No | ||
| 200 | PLEC1 | 2273 2198 2205 2178 2230 22247 2213 2231 2295 2172 5263 2266 | 13549 | -0.018 | 0.0343 | No | ||
| 201 | DAAM1 | 5536 | 13616 | -0.019 | 0.0314 | No | ||
| 202 | TMIE | 18984 | 13700 | -0.019 | 0.0277 | No | ||
| 203 | SEC14L2 | 20546 | 13725 | -0.020 | 0.0272 | No | ||
| 204 | ELMO1 | 3275 8939 3202 3271 3254 3235 3164 3175 3274 3188 3293 3214 3218 3239 | 13808 | -0.021 | 0.0235 | No | ||
| 205 | HOXC6 | 22339 2302 | 13875 | -0.021 | 0.0208 | No | ||
| 206 | LGP2 | 8155 | 14135 | -0.024 | 0.0076 | No | ||
| 207 | GRK5 | 9036 23814 | 14155 | -0.024 | 0.0075 | No | ||
| 208 | PTF1A | 15110 | 14194 | -0.025 | 0.0065 | No | ||
| 209 | CREB5 | 10551 | 14233 | -0.026 | 0.0054 | No | ||
| 210 | TCL1A | 20987 | 14273 | -0.026 | 0.0043 | No | ||
| 211 | LASS6 | 6307 10804 | 14387 | -0.028 | -0.0007 | No | ||
| 212 | KIF5A | 9222 | 14389 | -0.028 | 0.0003 | No | ||
| 213 | F12 | 3260 21453 | 14390 | -0.028 | 0.0014 | No | ||
| 214 | PRKG2 | 74 | 14478 | -0.030 | -0.0021 | No | ||
| 215 | ST7 | 1054 17519 | 14596 | -0.032 | -0.0072 | No | ||
| 216 | TAC1 | 17526 19853 | 14623 | -0.033 | -0.0073 | No | ||
| 217 | APOBEC2 | 22946 | 14694 | -0.034 | -0.0098 | No | ||
| 218 | SP4 | 20972 | 14839 | -0.037 | -0.0162 | No | ||
| 219 | G6PC | 20656 | 15012 | -0.041 | -0.0239 | No | ||
| 220 | CPB1 | 175 | 15013 | -0.041 | -0.0223 | No | ||
| 221 | DOCK4 | 10705 | 15019 | -0.041 | -0.0209 | No | ||
| 222 | ETV6 | 17264 | 15027 | -0.042 | -0.0196 | No | ||
| 223 | PRDM13 | 15933 | 15162 | -0.045 | -0.0251 | No | ||
| 224 | OVGP1 | 4519 8740 | 15233 | -0.048 | -0.0271 | No | ||
| 225 | SORT1 | 15452 5475 9847 | 15411 | -0.053 | -0.0346 | No | ||
| 226 | HOXB4 | 20686 | 15426 | -0.053 | -0.0333 | No | ||
| 227 | SPRED1 | 4331 | 15460 | -0.055 | -0.0329 | No | ||
| 228 | HOXC10 | 22342 | 15474 | -0.055 | -0.0314 | No | ||
| 229 | POU3F4 | 347 | 15540 | -0.058 | -0.0326 | No | ||
| 230 | LAMB2 | 9265 | 15724 | -0.066 | -0.0400 | No | ||
| 231 | ZBTB33 | 24352 | 15736 | -0.067 | -0.0379 | No | ||
| 232 | VAT1 | 11253 | 16241 | -0.095 | -0.0616 | No | ||
| 233 | HMGN2 | 9095 | 16299 | -0.098 | -0.0608 | No | ||
| 234 | CAMK2D | 4232 | 16308 | -0.099 | -0.0574 | No | ||
| 235 | HOXB2 | 8346 | 16315 | -0.099 | -0.0538 | No | ||
| 236 | RAB33A | 358 | 16718 | -0.133 | -0.0704 | No | ||
| 237 | CRSP8 | 15062 | 16896 | -0.153 | -0.0739 | No | ||
| 238 | MAP2K5 | 19088 | 16940 | -0.158 | -0.0700 | No | ||
| 239 | PJA2 | 5904 | 17203 | -0.185 | -0.0770 | No | ||
| 240 | NUDT2 | 16239 | 17472 | -0.221 | -0.0828 | No | ||
| 241 | STARD13 | 10833 | 17709 | -0.255 | -0.0856 | No | ||
| 242 | PRKAG1 | 22135 | 17739 | -0.261 | -0.0768 | No | ||
| 243 | MPP4 | 10399 | 17757 | -0.265 | -0.0672 | No | ||
| 244 | MXI1 | 23813 5139 | 17987 | -0.319 | -0.0671 | No | ||
| 245 | MRPS18B | 7365 | 18064 | -0.343 | -0.0576 | No | ||
| 246 | MARCKS | 9331 | 18173 | -0.404 | -0.0476 | No | ||
| 247 | SUPT4H1 | 9938 | 18315 | -0.497 | -0.0356 | No | ||
| 248 | ENO3 | 8905 | 18359 | -0.547 | -0.0163 | No | ||
| 249 | TLE4 | 23720 3697 | 18457 | -0.763 | 0.0087 | No |