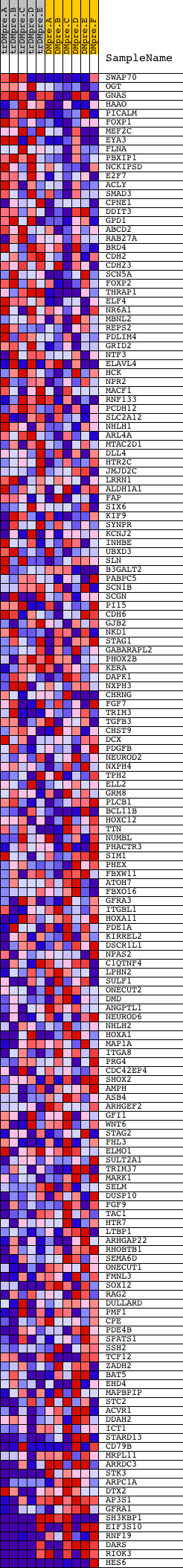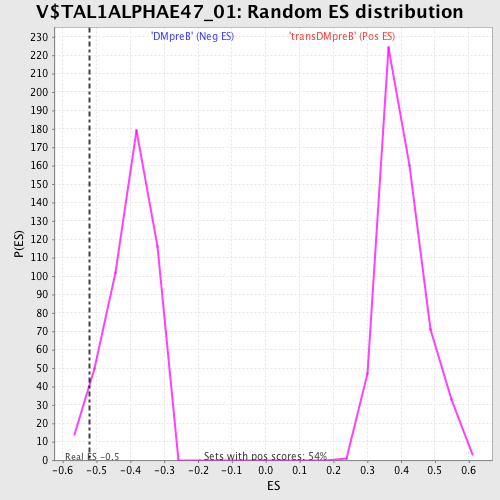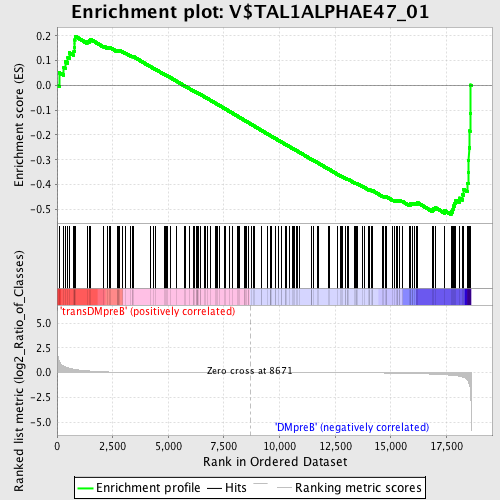
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMpreB_versus_DMpreB.cls#transDMpreB_versus_DMpreB_repos |
| Upregulated in class | DMpreB |
| GeneSet | V$TAL1ALPHAE47_01 |
| Enrichment Score (ES) | -0.5206106 |
| Normalized Enrichment Score (NES) | -1.3035283 |
| Nominal p-value | 0.04338395 |
| FDR q-value | 0.9863204 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SWAP70 | 18126 5553 9948 | 102 | 1.145 | 0.0510 | No | ||
| 2 | OGT | 4241 24274 | 300 | 0.635 | 0.0716 | No | ||
| 3 | GNAS | 9025 2963 2752 | 376 | 0.559 | 0.0952 | No | ||
| 4 | HAAO | 22885 1578 | 483 | 0.481 | 0.1131 | No | ||
| 5 | PICALM | 18191 | 549 | 0.442 | 0.1314 | No | ||
| 6 | FOXP1 | 4242 | 755 | 0.338 | 0.1370 | No | ||
| 7 | MEF2C | 3204 9378 | 787 | 0.327 | 0.1515 | No | ||
| 8 | EYA3 | 8936 2352 | 797 | 0.325 | 0.1670 | No | ||
| 9 | FLNA | 24129 | 798 | 0.324 | 0.1830 | No | ||
| 10 | PBXIP1 | 6028 | 805 | 0.321 | 0.1985 | No | ||
| 11 | NCKIPSD | 19306 | 1347 | 0.180 | 0.1781 | No | ||
| 12 | E2F7 | 19884 | 1442 | 0.168 | 0.1813 | No | ||
| 13 | ACLY | 8348 | 1491 | 0.160 | 0.1866 | No | ||
| 14 | SMAD3 | 19084 | 2085 | 0.085 | 0.1586 | No | ||
| 15 | CPNE1 | 11186 | 2252 | 0.071 | 0.1531 | No | ||
| 16 | DDIT3 | 19857 | 2355 | 0.065 | 0.1508 | No | ||
| 17 | GPD1 | 22361 | 2391 | 0.063 | 0.1520 | No | ||
| 18 | ABCD2 | 11204 6494 | 2712 | 0.046 | 0.1369 | No | ||
| 19 | RAB27A | 19381 | 2716 | 0.046 | 0.1390 | No | ||
| 20 | BRD4 | 7164 1609 1628 1598 1552 1601 12176 1630 1501 | 2775 | 0.044 | 0.1380 | No | ||
| 21 | CDH2 | 1963 8727 8726 4508 | 2780 | 0.044 | 0.1400 | No | ||
| 22 | CDH23 | 3369 19759 3442 | 2820 | 0.042 | 0.1400 | No | ||
| 23 | SCN5A | 5412 | 2936 | 0.039 | 0.1356 | No | ||
| 24 | FOXP2 | 4312 17523 8523 | 3091 | 0.034 | 0.1290 | No | ||
| 25 | THRAP1 | 20309 | 3304 | 0.029 | 0.1189 | No | ||
| 26 | ELF4 | 24162 | 3391 | 0.027 | 0.1156 | No | ||
| 27 | NR6A1 | 14596 | 3407 | 0.026 | 0.1161 | No | ||
| 28 | MBNL2 | 21932 | 3427 | 0.026 | 0.1163 | No | ||
| 29 | REPS2 | 24018 | 4178 | 0.016 | 0.0765 | No | ||
| 30 | PDLIM4 | 20455 | 4341 | 0.015 | 0.0684 | No | ||
| 31 | GRID2 | 4803 | 4417 | 0.014 | 0.0651 | No | ||
| 32 | NTF3 | 16991 | 4830 | 0.011 | 0.0433 | No | ||
| 33 | ELAVL4 | 15805 4889 9137 | 4871 | 0.011 | 0.0417 | No | ||
| 34 | HCK | 14787 | 4930 | 0.011 | 0.0390 | No | ||
| 35 | NPR2 | 16230 | 4966 | 0.011 | 0.0377 | No | ||
| 36 | MACF1 | 4316 15763 2382 15764 | 5115 | 0.010 | 0.0301 | No | ||
| 37 | RNF133 | 11813 | 5356 | 0.009 | 0.0175 | No | ||
| 38 | PCDH12 | 7005 | 5726 | 0.007 | -0.0021 | No | ||
| 39 | SLC2A12 | 20073 | 5781 | 0.007 | -0.0047 | No | ||
| 40 | NHLH1 | 13754 | 5950 | 0.006 | -0.0135 | No | ||
| 41 | ARL4A | 8627 | 6114 | 0.006 | -0.0220 | No | ||
| 42 | MTAC2D1 | 21003 | 6136 | 0.006 | -0.0229 | No | ||
| 43 | DLL4 | 7040 | 6155 | 0.006 | -0.0235 | No | ||
| 44 | HTR2C | 24230 | 6285 | 0.005 | -0.0303 | No | ||
| 45 | JMJD2C | 16188 | 6286 | 0.005 | -0.0300 | No | ||
| 46 | LRRN1 | 17343 | 6290 | 0.005 | -0.0299 | No | ||
| 47 | ALDH1A1 | 8569 | 6313 | 0.005 | -0.0308 | No | ||
| 48 | FAP | 14577 | 6318 | 0.005 | -0.0308 | No | ||
| 49 | SIX6 | 21247 | 6339 | 0.005 | -0.0316 | No | ||
| 50 | KIF9 | 19292 | 6445 | 0.005 | -0.0370 | No | ||
| 51 | SYNPR | 22094 | 6465 | 0.005 | -0.0378 | No | ||
| 52 | KCNJ2 | 20612 | 6638 | 0.004 | -0.0469 | No | ||
| 53 | INHBE | 19604 | 6675 | 0.004 | -0.0487 | No | ||
| 54 | UBXD3 | 15701 | 6763 | 0.004 | -0.0532 | No | ||
| 55 | SLN | 19450 | 6880 | 0.004 | -0.0593 | No | ||
| 56 | B3GALT2 | 6498 | 7136 | 0.003 | -0.0729 | No | ||
| 57 | PABPC5 | 24077 | 7155 | 0.003 | -0.0737 | No | ||
| 58 | SCN1B | 1148 17872 | 7207 | 0.003 | -0.0764 | No | ||
| 59 | SCGN | 3197 21516 | 7293 | 0.003 | -0.0808 | No | ||
| 60 | PI15 | 13586 | 7508 | 0.002 | -0.0923 | No | ||
| 61 | CDH6 | 22320 | 7546 | 0.002 | -0.0942 | No | ||
| 62 | GJB2 | 9018 21806 | 7743 | 0.002 | -0.1047 | No | ||
| 63 | NKD1 | 8228 | 7868 | 0.002 | -0.1114 | No | ||
| 64 | STAG1 | 5520 9905 2987 | 8109 | 0.001 | -0.1243 | No | ||
| 65 | GABARAPL2 | 8211 13552 | 8166 | 0.001 | -0.1273 | No | ||
| 66 | PHOX2B | 16524 | 8188 | 0.001 | -0.1284 | No | ||
| 67 | KERA | 19895 | 8416 | 0.001 | -0.1407 | No | ||
| 68 | DAPK1 | 7632 | 8443 | 0.000 | -0.1421 | No | ||
| 69 | NXPH3 | 20281 | 8463 | 0.000 | -0.1431 | No | ||
| 70 | CHRNG | 4323 | 8502 | 0.000 | -0.1451 | No | ||
| 71 | FGF7 | 8965 | 8584 | 0.000 | -0.1495 | No | ||
| 72 | TRIM3 | 17703 | 8603 | 0.000 | -0.1505 | No | ||
| 73 | TGFB3 | 10161 | 8719 | -0.000 | -0.1567 | No | ||
| 74 | CHST9 | 7727 | 8831 | -0.000 | -0.1627 | No | ||
| 75 | DCX | 24040 8841 4604 | 8883 | -0.000 | -0.1654 | No | ||
| 76 | PDGFB | 22199 2311 | 9170 | -0.001 | -0.1809 | No | ||
| 77 | NEUROD2 | 20262 | 9202 | -0.001 | -0.1825 | No | ||
| 78 | NXPH4 | 19602 | 9437 | -0.002 | -0.1951 | No | ||
| 79 | TPH2 | 19626 | 9569 | -0.002 | -0.2021 | No | ||
| 80 | ELL2 | 5329 | 9593 | -0.002 | -0.2033 | No | ||
| 81 | GRM8 | 4805 | 9608 | -0.002 | -0.2039 | No | ||
| 82 | PLCB1 | 14832 2821 | 9655 | -0.002 | -0.2063 | No | ||
| 83 | BCL11B | 7190 | 9826 | -0.003 | -0.2154 | No | ||
| 84 | HOXC12 | 22343 | 9835 | -0.003 | -0.2157 | No | ||
| 85 | TTN | 14552 2743 14553 | 9972 | -0.003 | -0.2229 | No | ||
| 86 | NUMBL | 1663 9496 | 10087 | -0.003 | -0.2290 | No | ||
| 87 | PHACTR3 | 7900 2770 | 10090 | -0.003 | -0.2289 | No | ||
| 88 | SIM1 | 20031 | 10272 | -0.004 | -0.2386 | No | ||
| 89 | PHEX | 24024 | 10275 | -0.004 | -0.2385 | No | ||
| 90 | FBXW11 | 20926 | 10329 | -0.004 | -0.2412 | No | ||
| 91 | ATOH7 | 20000 | 10441 | -0.004 | -0.2470 | No | ||
| 92 | FBXO16 | 11929 | 10576 | -0.004 | -0.2540 | No | ||
| 93 | GFRA3 | 23469 | 10642 | -0.005 | -0.2573 | No | ||
| 94 | ITGBL1 | 5850 | 10689 | -0.005 | -0.2596 | No | ||
| 95 | HOXA11 | 17146 | 10774 | -0.005 | -0.2639 | No | ||
| 96 | PDE1A | 9541 5234 | 10809 | -0.005 | -0.2655 | No | ||
| 97 | KIRREL2 | 17889 3922 | 10912 | -0.005 | -0.2708 | No | ||
| 98 | DSCR1L1 | 23231 | 11415 | -0.007 | -0.2977 | No | ||
| 99 | NPAS2 | 5186 | 11428 | -0.007 | -0.2980 | No | ||
| 100 | C1QTNF4 | 14953 | 11434 | -0.007 | -0.2979 | No | ||
| 101 | LPHN2 | 13619 | 11529 | -0.007 | -0.3027 | No | ||
| 102 | SULF1 | 6285 | 11714 | -0.008 | -0.3123 | No | ||
| 103 | ONECUT2 | 10346 | 11762 | -0.008 | -0.3145 | No | ||
| 104 | DMD | 24295 2647 | 12181 | -0.009 | -0.3367 | No | ||
| 105 | ANGPTL1 | 10195 8589 13064 | 12225 | -0.010 | -0.3385 | No | ||
| 106 | NEUROD6 | 17138 | 12592 | -0.011 | -0.3578 | No | ||
| 107 | NHLH2 | 9462 | 12748 | -0.012 | -0.3656 | No | ||
| 108 | HOXA1 | 1005 17152 | 12766 | -0.012 | -0.3659 | No | ||
| 109 | MAP1A | 5124 | 12820 | -0.013 | -0.3682 | No | ||
| 110 | ITGA8 | 14685 | 12976 | -0.014 | -0.3759 | No | ||
| 111 | PRG4 | 13592 4036 | 12982 | -0.014 | -0.3755 | No | ||
| 112 | CDC42EP4 | 20163 | 13046 | -0.014 | -0.3782 | No | ||
| 113 | SHOX2 | 1802 5432 | 13074 | -0.014 | -0.3790 | No | ||
| 114 | AMPH | 3256 21700 | 13096 | -0.014 | -0.3794 | No | ||
| 115 | ASB4 | 990 17529 | 13355 | -0.016 | -0.3926 | No | ||
| 116 | ARHGEF2 | 4987 | 13419 | -0.017 | -0.3952 | No | ||
| 117 | GFI1 | 16451 | 13461 | -0.017 | -0.3965 | No | ||
| 118 | WNT6 | 5881 | 13475 | -0.017 | -0.3964 | No | ||
| 119 | STAG2 | 5521 | 13520 | -0.018 | -0.3979 | No | ||
| 120 | FHL3 | 16092 2457 | 13715 | -0.019 | -0.4075 | No | ||
| 121 | ELMO1 | 3275 8939 3202 3271 3254 3235 3164 3175 3274 3188 3293 3214 3218 3239 | 13808 | -0.021 | -0.4114 | No | ||
| 122 | SULT2A1 | 5527 | 14010 | -0.023 | -0.4212 | No | ||
| 123 | TRIM37 | 20715 | 14029 | -0.023 | -0.4211 | No | ||
| 124 | MARK1 | 5955 | 14035 | -0.023 | -0.4202 | No | ||
| 125 | SELM | 20967 | 14142 | -0.024 | -0.4248 | No | ||
| 126 | DUSP10 | 4003 14016 | 14161 | -0.024 | -0.4245 | No | ||
| 127 | FGF9 | 8966 | 14196 | -0.025 | -0.4251 | No | ||
| 128 | TAC1 | 17526 19853 | 14623 | -0.033 | -0.4466 | No | ||
| 129 | HTR7 | 23690 | 14690 | -0.034 | -0.4485 | No | ||
| 130 | LTBP1 | 1536 1604 6503 1559 1564 1518 | 14738 | -0.035 | -0.4494 | No | ||
| 131 | ARHGAP22 | 22050 | 14788 | -0.036 | -0.4502 | No | ||
| 132 | RHOBTB1 | 12789 | 14801 | -0.036 | -0.4491 | No | ||
| 133 | SEMA6D | 2800 14876 90 | 15085 | -0.043 | -0.4623 | No | ||
| 134 | ONECUT1 | 4858 | 15168 | -0.045 | -0.4645 | No | ||
| 135 | FMNL3 | 22134 5866 | 15243 | -0.048 | -0.4661 | No | ||
| 136 | SOX12 | 5478 | 15278 | -0.049 | -0.4656 | No | ||
| 137 | RAG2 | 14941 | 15316 | -0.050 | -0.4651 | No | ||
| 138 | DULLARD | 20816 | 15397 | -0.052 | -0.4669 | No | ||
| 139 | PMF1 | 12452 | 15398 | -0.052 | -0.4643 | No | ||
| 140 | CPE | 18868 | 15507 | -0.057 | -0.4673 | No | ||
| 141 | PDE4B | 9543 | 15823 | -0.071 | -0.4809 | No | ||
| 142 | SPATS1 | 22973 | 15873 | -0.073 | -0.4800 | No | ||
| 143 | SSH2 | 6202 | 15880 | -0.074 | -0.4766 | No | ||
| 144 | TCF12 | 10044 3125 | 15972 | -0.078 | -0.4777 | No | ||
| 145 | ZADH2 | 23501 | 15985 | -0.079 | -0.4745 | No | ||
| 146 | BAT5 | 23261 | 16073 | -0.084 | -0.4750 | No | ||
| 147 | EHD4 | 8275 | 16169 | -0.090 | -0.4757 | No | ||
| 148 | MAPBPIP | 15288 | 16204 | -0.092 | -0.4730 | No | ||
| 149 | STC2 | 5526 | 16878 | -0.152 | -0.5020 | No | ||
| 150 | ACVR1 | 4334 | 16905 | -0.154 | -0.4958 | No | ||
| 151 | DDAH2 | 23263 | 16998 | -0.164 | -0.4927 | No | ||
| 152 | ICT1 | 20598 | 17417 | -0.213 | -0.5048 | No | ||
| 153 | STARD13 | 10833 | 17709 | -0.255 | -0.5080 | Yes | ||
| 154 | CD79B | 20185 1309 | 17788 | -0.272 | -0.4988 | Yes | ||
| 155 | MRPL11 | 23774 | 17807 | -0.278 | -0.4861 | Yes | ||
| 156 | ARRDC3 | 4191 | 17852 | -0.287 | -0.4743 | Yes | ||
| 157 | STK3 | 7084 | 17919 | -0.304 | -0.4629 | Yes | ||
| 158 | ARPC1A | 3541 16628 3519 | 18073 | -0.346 | -0.4541 | Yes | ||
| 159 | DTX2 | 16674 3488 | 18229 | -0.441 | -0.4407 | Yes | ||
| 160 | AP3S1 | 8604 | 18273 | -0.467 | -0.4200 | Yes | ||
| 161 | GFRA1 | 9015 | 18437 | -0.707 | -0.3940 | Yes | ||
| 162 | SH3KBP1 | 2631 12206 7189 | 18489 | -0.920 | -0.3513 | Yes | ||
| 163 | EIF3S10 | 4659 8887 | 18500 | -0.972 | -0.3039 | Yes | ||
| 164 | RNF19 | 22314 | 18513 | -1.086 | -0.2509 | Yes | ||
| 165 | DARS | 10375 13846 | 18551 | -1.401 | -0.1838 | Yes | ||
| 166 | RIOK3 | 7354 12425 1973 | 18565 | -1.473 | -0.1118 | Yes | ||
| 167 | HES6 | 13882 4058 | 18589 | -2.320 | 0.0015 | Yes |