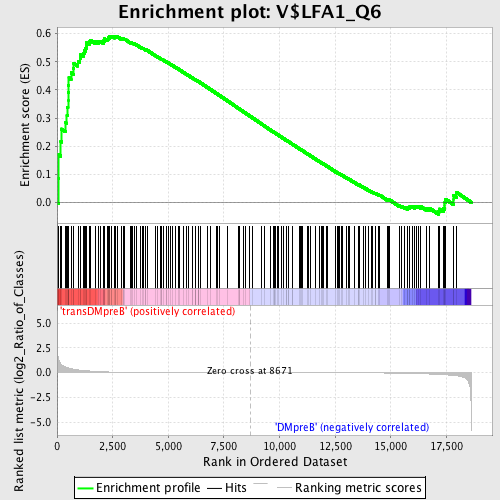
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMpreB_versus_DMpreB.cls#transDMpreB_versus_DMpreB_repos |
| Upregulated in class | transDMpreB |
| GeneSet | V$LFA1_Q6 |
| Enrichment Score (ES) | 0.5911681 |
| Normalized Enrichment Score (NES) | 1.4657848 |
| Nominal p-value | 0.009505703 |
| FDR q-value | 0.3589648 |
| FWER p-Value | 0.997 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | TPCN1 | 16398 6383 10915 | 57 | 1.528 | 0.0862 | Yes | ||
| 2 | RBM12 | 7980 2710 13280 | 68 | 1.432 | 0.1693 | Yes | ||
| 3 | GAD1 | 14993 2690 | 168 | 0.889 | 0.2159 | Yes | ||
| 4 | CCNL1 | 15325 | 198 | 0.798 | 0.2610 | Yes | ||
| 5 | CIC | 18338 | 388 | 0.550 | 0.2829 | Yes | ||
| 6 | ZFPM1 | 18439 | 443 | 0.505 | 0.3095 | Yes | ||
| 7 | BRPF1 | 17332 | 464 | 0.490 | 0.3370 | Yes | ||
| 8 | UPF2 | 11591 | 490 | 0.477 | 0.3636 | Yes | ||
| 9 | RBBP6 | 5365 | 495 | 0.475 | 0.3911 | Yes | ||
| 10 | BACE1 | 6214 | 520 | 0.464 | 0.4169 | Yes | ||
| 11 | HES1 | 22798 | 533 | 0.455 | 0.4429 | Yes | ||
| 12 | PHF12 | 11192 6483 | 629 | 0.396 | 0.4609 | Yes | ||
| 13 | NXF1 | 11986 | 728 | 0.350 | 0.4760 | Yes | ||
| 14 | CEBPB | 8733 | 751 | 0.341 | 0.4947 | Yes | ||
| 15 | RHOG | 17727 | 938 | 0.280 | 0.5010 | Yes | ||
| 16 | RASGRP2 | 9693 | 1028 | 0.254 | 0.5110 | Yes | ||
| 17 | CKB | 4523 | 1047 | 0.250 | 0.5246 | Yes | ||
| 18 | POLD4 | 12822 | 1173 | 0.217 | 0.5306 | Yes | ||
| 19 | DCTN1 | 1100 17392 | 1228 | 0.206 | 0.5397 | Yes | ||
| 20 | GBA2 | 15898 | 1285 | 0.193 | 0.5479 | Yes | ||
| 21 | TAL1 | 16129 | 1309 | 0.187 | 0.5576 | Yes | ||
| 22 | PPP1R16A | 22441 | 1329 | 0.183 | 0.5673 | Yes | ||
| 23 | NFKBIA | 21065 | 1447 | 0.167 | 0.5707 | Yes | ||
| 24 | ADM | 18125 | 1508 | 0.157 | 0.5766 | Yes | ||
| 25 | FLI1 | 4729 | 1716 | 0.128 | 0.5728 | Yes | ||
| 26 | TITF1 | 10180 21062 | 1852 | 0.110 | 0.5720 | Yes | ||
| 27 | ZFP36 | 84 | 1950 | 0.098 | 0.5724 | Yes | ||
| 28 | SMAD3 | 19084 | 2085 | 0.085 | 0.5701 | Yes | ||
| 29 | COX7A2L | 9819 | 2102 | 0.083 | 0.5741 | Yes | ||
| 30 | PYGO2 | 12748 | 2131 | 0.081 | 0.5773 | Yes | ||
| 31 | SMARCD2 | 13515 | 2140 | 0.080 | 0.5815 | Yes | ||
| 32 | CPNE1 | 11186 | 2252 | 0.071 | 0.5797 | Yes | ||
| 33 | STOML2 | 15902 | 2295 | 0.068 | 0.5814 | Yes | ||
| 34 | OBSCN | 20434 | 2300 | 0.068 | 0.5851 | Yes | ||
| 35 | UBL3 | 16284 | 2350 | 0.065 | 0.5862 | Yes | ||
| 36 | CRMP1 | 16860 | 2353 | 0.065 | 0.5899 | Yes | ||
| 37 | OLFML2A | 6304 | 2438 | 0.060 | 0.5888 | Yes | ||
| 38 | SLC26A9 | 11560 980 4067 | 2459 | 0.058 | 0.5912 | Yes | ||
| 39 | HNRPR | 13196 7909 2500 | 2587 | 0.051 | 0.5873 | No | ||
| 40 | RAB5C | 20226 | 2613 | 0.050 | 0.5889 | No | ||
| 41 | HIRA | 4852 9090 | 2626 | 0.049 | 0.5911 | No | ||
| 42 | PCBP2 | 9533 | 2698 | 0.047 | 0.5900 | No | ||
| 43 | BNC2 | 15850 | 2906 | 0.040 | 0.5811 | No | ||
| 44 | MAZ | 1327 17623 | 2964 | 0.038 | 0.5802 | No | ||
| 45 | ANXA4 | 17083 | 2972 | 0.037 | 0.5820 | No | ||
| 46 | IFRD2 | 19320 | 3032 | 0.036 | 0.5809 | No | ||
| 47 | THRAP1 | 20309 | 3304 | 0.029 | 0.5679 | No | ||
| 48 | TCF7 | 1467 20466 | 3342 | 0.028 | 0.5675 | No | ||
| 49 | ELF4 | 24162 | 3391 | 0.027 | 0.5665 | No | ||
| 50 | SALL1 | 18796 12207 | 3456 | 0.026 | 0.5645 | No | ||
| 51 | NFAT5 | 3921 7037 12036 | 3565 | 0.024 | 0.5601 | No | ||
| 52 | ARID1A | 8214 2501 13554 | 3753 | 0.021 | 0.5512 | No | ||
| 53 | NEURL | 5164 | 3853 | 0.020 | 0.5469 | No | ||
| 54 | PTRF | 9667 | 3864 | 0.020 | 0.5476 | No | ||
| 55 | MOV10 | 9403 9402 5114 | 3984 | 0.018 | 0.5422 | No | ||
| 56 | LCAT | 18762 | 3992 | 0.018 | 0.5429 | No | ||
| 57 | SEMA6A | 23439 9802 | 4066 | 0.017 | 0.5399 | No | ||
| 58 | TSP50 | 19286 | 4414 | 0.014 | 0.5219 | No | ||
| 59 | SLC2A4 | 20380 | 4515 | 0.013 | 0.5173 | No | ||
| 60 | UQCRC1 | 10259 | 4648 | 0.012 | 0.5108 | No | ||
| 61 | LAG3 | 17000 | 4709 | 0.012 | 0.5083 | No | ||
| 62 | CAV1 | 8698 | 4710 | 0.012 | 0.5090 | No | ||
| 63 | EPO | 8911 | 4801 | 0.011 | 0.5048 | No | ||
| 64 | CREB3L2 | 17182 | 4934 | 0.011 | 0.4982 | No | ||
| 65 | ADAMTS4 | 10791 | 4935 | 0.011 | 0.4989 | No | ||
| 66 | SOX5 | 9850 1044 5480 16937 | 5016 | 0.010 | 0.4951 | No | ||
| 67 | MYOG | 14129 | 5107 | 0.010 | 0.4908 | No | ||
| 68 | S100A5 | 15529 | 5180 | 0.009 | 0.4875 | No | ||
| 69 | USH1G | 20153 | 5326 | 0.009 | 0.4801 | No | ||
| 70 | ACVR1C | 14585 | 5434 | 0.008 | 0.4748 | No | ||
| 71 | EPHB2 | 4675 2440 8910 | 5507 | 0.008 | 0.4714 | No | ||
| 72 | POU4F1 | 21727 | 5678 | 0.007 | 0.4626 | No | ||
| 73 | KCNQ4 | 12247 2533 | 5818 | 0.007 | 0.4554 | No | ||
| 74 | NR4A1 | 9099 | 5883 | 0.007 | 0.4524 | No | ||
| 75 | ETV1 | 4688 8925 8924 | 6070 | 0.006 | 0.4426 | No | ||
| 76 | CHX10 | 21208 | 6074 | 0.006 | 0.4428 | No | ||
| 77 | PRRX1 | 9595 5271 | 6205 | 0.006 | 0.4361 | No | ||
| 78 | CNKSR2 | 2553 24021 | 6206 | 0.006 | 0.4364 | No | ||
| 79 | GSS | 14380 9047 | 6214 | 0.006 | 0.4364 | No | ||
| 80 | GPR4 | 18364 | 6349 | 0.005 | 0.4294 | No | ||
| 81 | OTOP2 | 20600 | 6351 | 0.005 | 0.4297 | No | ||
| 82 | ACCN1 | 20327 1194 | 6373 | 0.005 | 0.4289 | No | ||
| 83 | SH3BP1 | 632 22431 | 6456 | 0.005 | 0.4247 | No | ||
| 84 | PNOC | 21783 | 6752 | 0.004 | 0.4090 | No | ||
| 85 | WNT4 | 16025 | 6898 | 0.004 | 0.4013 | No | ||
| 86 | MGEA5 | 8001 | 7152 | 0.003 | 0.3878 | No | ||
| 87 | LMOD3 | 17059 | 7209 | 0.003 | 0.3849 | No | ||
| 88 | CLIC5 | 23229 | 7318 | 0.003 | 0.3792 | No | ||
| 89 | CNN2 | 19953 3391 | 7657 | 0.002 | 0.3610 | No | ||
| 90 | DPF2 | 9719 | 8147 | 0.001 | 0.3346 | No | ||
| 91 | SIX4 | 5445 | 8186 | 0.001 | 0.3326 | No | ||
| 92 | RHO | 17314 | 8216 | 0.001 | 0.3310 | No | ||
| 93 | EDG1 | 15187 | 8398 | 0.001 | 0.3213 | No | ||
| 94 | HARS2 | 23592 14820 | 8487 | 0.000 | 0.3165 | No | ||
| 95 | SESN3 | 19561 | 8626 | 0.000 | 0.3090 | No | ||
| 96 | EMILIN1 | 16891 | 8786 | -0.000 | 0.3004 | No | ||
| 97 | DUOX2 | 14453 | 9172 | -0.001 | 0.2796 | No | ||
| 98 | RABGGTA | 2669 21820 | 9337 | -0.001 | 0.2708 | No | ||
| 99 | IL1F10 | 15097 | 9577 | -0.002 | 0.2580 | No | ||
| 100 | MYBPH | 14130 | 9737 | -0.002 | 0.2495 | No | ||
| 101 | NPTXR | 7854 22200 | 9748 | -0.002 | 0.2491 | No | ||
| 102 | CASQ2 | 15477 | 9785 | -0.002 | 0.2473 | No | ||
| 103 | BCL11B | 7190 | 9826 | -0.003 | 0.2452 | No | ||
| 104 | JPH4 | 21826 | 9919 | -0.003 | 0.2404 | No | ||
| 105 | NR5A2 | 4105 13817 | 9940 | -0.003 | 0.2395 | No | ||
| 106 | PPY | 20208 | 10101 | -0.003 | 0.2310 | No | ||
| 107 | BOC | 22587 | 10176 | -0.003 | 0.2272 | No | ||
| 108 | IMMT | 367 8029 | 10292 | -0.004 | 0.2212 | No | ||
| 109 | MNT | 20784 | 10328 | -0.004 | 0.2195 | No | ||
| 110 | CTGF | 20068 | 10412 | -0.004 | 0.2152 | No | ||
| 111 | LIN28 | 15723 | 10416 | -0.004 | 0.2153 | No | ||
| 112 | GRIN2A | 22668 | 10568 | -0.004 | 0.2073 | No | ||
| 113 | XYLT1 | 18113 | 10572 | -0.004 | 0.2074 | No | ||
| 114 | SLC7A10 | 18294 | 10910 | -0.005 | 0.1895 | No | ||
| 115 | PALM | 19960 3350 | 10937 | -0.005 | 0.1884 | No | ||
| 116 | MSX1 | 16547 | 10998 | -0.005 | 0.1854 | No | ||
| 117 | SP7 | 22110 | 11041 | -0.006 | 0.1835 | No | ||
| 118 | CACNB1 | 1443 1304 | 11269 | -0.006 | 0.1715 | No | ||
| 119 | NPAS3 | 2077 21273 | 11296 | -0.006 | 0.1705 | No | ||
| 120 | NGFB | 15476 | 11384 | -0.006 | 0.1661 | No | ||
| 121 | NYX | 24378 | 11616 | -0.007 | 0.1540 | No | ||
| 122 | GNAO1 | 4785 3829 | 11779 | -0.008 | 0.1457 | No | ||
| 123 | FXYD3 | 9370 | 11878 | -0.008 | 0.1409 | No | ||
| 124 | DHH | 4629 | 11950 | -0.008 | 0.1375 | No | ||
| 125 | NKX2-3 | 5177 23845 | 11953 | -0.008 | 0.1379 | No | ||
| 126 | GABRG2 | 1259 4748 8996 1331 | 12126 | -0.009 | 0.1291 | No | ||
| 127 | KIF1C | 4951 | 12162 | -0.009 | 0.1277 | No | ||
| 128 | LRP8 | 16149 2425 | 12534 | -0.011 | 0.1083 | No | ||
| 129 | TNRC4 | 15514 | 12597 | -0.011 | 0.1056 | No | ||
| 130 | FLT1 | 3483 16287 | 12654 | -0.012 | 0.1032 | No | ||
| 131 | PAK1IP1 | 21660 | 12676 | -0.012 | 0.1028 | No | ||
| 132 | KLK9 | 18267 | 12775 | -0.012 | 0.0982 | No | ||
| 133 | FES | 17779 3949 | 12817 | -0.013 | 0.0967 | No | ||
| 134 | BCL6 | 22624 | 13018 | -0.014 | 0.0866 | No | ||
| 135 | PLA2G10 | 22658 | 13082 | -0.014 | 0.0841 | No | ||
| 136 | PAX7 | 15697 | 13133 | -0.015 | 0.0822 | No | ||
| 137 | KCNAB1 | 1791 15577 | 13371 | -0.016 | 0.0703 | No | ||
| 138 | RFX4 | 7721 3410 | 13532 | -0.018 | 0.0627 | No | ||
| 139 | PLEC1 | 2273 2198 2205 2178 2230 22247 2213 2231 2295 2172 5263 2266 | 13549 | -0.018 | 0.0629 | No | ||
| 140 | ZNF385 | 22106 2297 | 13572 | -0.018 | 0.0627 | No | ||
| 141 | EFNA3 | 15278 | 13752 | -0.020 | 0.0542 | No | ||
| 142 | HOXC6 | 22339 2302 | 13875 | -0.021 | 0.0488 | No | ||
| 143 | PDLIM7 | 7434 | 14000 | -0.023 | 0.0434 | No | ||
| 144 | DLGAP4 | 2950 10461 | 14132 | -0.024 | 0.0377 | No | ||
| 145 | CLTC | 14610 | 14182 | -0.025 | 0.0365 | No | ||
| 146 | M6PR | 1053 5044 9328 | 14303 | -0.027 | 0.0315 | No | ||
| 147 | DLX1 | 14988 | 14308 | -0.027 | 0.0329 | No | ||
| 148 | NRGN | 19176 | 14447 | -0.029 | 0.0271 | No | ||
| 149 | CACNA1G | 1237 940 20292 | 14502 | -0.030 | 0.0259 | No | ||
| 150 | NCDN | 15756 2487 6471 | 14856 | -0.038 | 0.0090 | No | ||
| 151 | NEUROG1 | 21446 | 14906 | -0.039 | 0.0086 | No | ||
| 152 | GSK3B | 22761 | 14935 | -0.040 | 0.0094 | No | ||
| 153 | ROM1 | 23751 | 15367 | -0.051 | -0.0110 | No | ||
| 154 | HOXC10 | 22342 | 15474 | -0.055 | -0.0135 | No | ||
| 155 | RAB4B | 5345 | 15636 | -0.062 | -0.0186 | No | ||
| 156 | GPR3 | 15729 | 15746 | -0.067 | -0.0206 | No | ||
| 157 | NPC2 | 21023 | 15756 | -0.068 | -0.0171 | No | ||
| 158 | PDYN | 14429 | 15842 | -0.072 | -0.0175 | No | ||
| 159 | SLC13A5 | 133 | 15855 | -0.072 | -0.0139 | No | ||
| 160 | VASP | 5847 | 15954 | -0.078 | -0.0147 | No | ||
| 161 | LZTS2 | 10362 | 16068 | -0.083 | -0.0160 | No | ||
| 162 | ARID4A | 2080 6215 | 16131 | -0.087 | -0.0142 | No | ||
| 163 | FKBP2 | 8972 | 16221 | -0.093 | -0.0136 | No | ||
| 164 | TBR1 | 83 | 16355 | -0.102 | -0.0149 | No | ||
| 165 | TRIP10 | 23174 | 16595 | -0.122 | -0.0207 | No | ||
| 166 | SLC25A29 | 20985 | 16734 | -0.134 | -0.0203 | No | ||
| 167 | GABARAP | 20815 | 17152 | -0.180 | -0.0324 | No | ||
| 168 | NDUFS2 | 4089 13760 | 17185 | -0.184 | -0.0234 | No | ||
| 169 | AKT3 | 13739 982 | 17357 | -0.204 | -0.0208 | No | ||
| 170 | NACA | 9444 5147 | 17428 | -0.214 | -0.0121 | No | ||
| 171 | MRPL40 | 22641 | 17430 | -0.214 | 0.0004 | No | ||
| 172 | BAX | 17832 | 17465 | -0.220 | 0.0114 | No | ||
| 173 | CXCR4 | 13844 | 17811 | -0.278 | 0.0090 | No | ||
| 174 | MCRS1 | 6961 6960 6962 | 17814 | -0.279 | 0.0251 | No | ||
| 175 | HEBP1 | 16961 | 17962 | -0.313 | 0.0355 | No |

