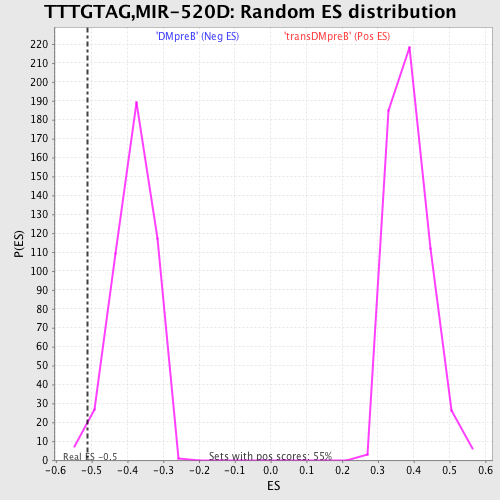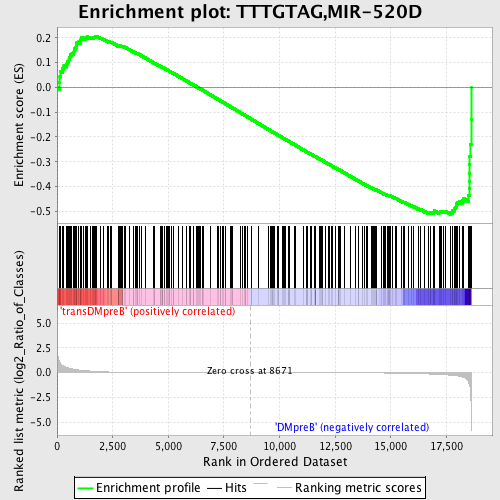
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMpreB_versus_DMpreB.cls#transDMpreB_versus_DMpreB_repos |
| Upregulated in class | DMpreB |
| GeneSet | TTTGTAG,MIR-520D |
| Enrichment Score (ES) | -0.5129275 |
| Normalized Enrichment Score (NES) | -1.3394123 |
| Nominal p-value | 0.02 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SFRS5 | 9808 2062 | 120 | 1.050 | 0.0195 | No | ||
| 2 | BRWD1 | 8222 1659 22532 | 126 | 1.017 | 0.0444 | No | ||
| 3 | GAD1 | 14993 2690 | 168 | 0.889 | 0.0642 | No | ||
| 4 | CUGBP2 | 4687 8923 | 254 | 0.703 | 0.0770 | No | ||
| 5 | RNF44 | 8364 | 306 | 0.631 | 0.0898 | No | ||
| 6 | PHF2 | 21470 | 441 | 0.507 | 0.0951 | No | ||
| 7 | MTA1 | 21128 | 477 | 0.483 | 0.1052 | No | ||
| 8 | HES1 | 22798 | 533 | 0.455 | 0.1135 | No | ||
| 9 | PRDM2 | 2522 4289 | 570 | 0.432 | 0.1222 | No | ||
| 10 | CCND1 | 4487 4488 8707 17535 | 581 | 0.427 | 0.1322 | No | ||
| 11 | MAP3K7IP2 | 19827 | 667 | 0.380 | 0.1370 | No | ||
| 12 | NXF1 | 11986 | 728 | 0.350 | 0.1424 | No | ||
| 13 | HNRPA0 | 13382 | 762 | 0.336 | 0.1489 | No | ||
| 14 | MEF2C | 3204 9378 | 787 | 0.327 | 0.1557 | No | ||
| 15 | ANKRD10 | 18935 | 840 | 0.310 | 0.1606 | No | ||
| 16 | ATP2A2 | 4421 3481 | 863 | 0.303 | 0.1669 | No | ||
| 17 | GTPBP1 | 4814 2255 | 884 | 0.298 | 0.1732 | No | ||
| 18 | PRPF39 | 21259 | 888 | 0.296 | 0.1803 | No | ||
| 19 | MECP2 | 5088 | 951 | 0.275 | 0.1838 | No | ||
| 20 | WASF2 | 6326 | 1054 | 0.248 | 0.1844 | No | ||
| 21 | PURB | 5335 | 1055 | 0.247 | 0.1905 | No | ||
| 22 | CUEDC1 | 8343 | 1070 | 0.242 | 0.1957 | No | ||
| 23 | VAMP2 | 20826 | 1082 | 0.237 | 0.2010 | No | ||
| 24 | FMN2 | 14037 | 1197 | 0.212 | 0.2000 | No | ||
| 25 | STK4 | 14743 | 1295 | 0.191 | 0.1995 | No | ||
| 26 | TAL1 | 16129 | 1309 | 0.187 | 0.2034 | No | ||
| 27 | CTCF | 18490 | 1361 | 0.179 | 0.2051 | No | ||
| 28 | ACACA | 309 4209 | 1489 | 0.160 | 0.2022 | No | ||
| 29 | RANBP2 | 20019 | 1569 | 0.149 | 0.2015 | No | ||
| 30 | LRRC8D | 16769 | 1656 | 0.136 | 0.2002 | No | ||
| 31 | PUM1 | 8160 | 1663 | 0.135 | 0.2032 | No | ||
| 32 | H3F3B | 4839 | 1708 | 0.128 | 0.2040 | No | ||
| 33 | NEDD9 | 3206 21483 | 1715 | 0.128 | 0.2068 | No | ||
| 34 | SNF1LK | 23033 | 1762 | 0.123 | 0.2074 | No | ||
| 35 | LUC7L | 47 1520 | 1969 | 0.096 | 0.1985 | No | ||
| 36 | ARRDC1 | 14669 | 2092 | 0.084 | 0.1940 | No | ||
| 37 | CRHBP | 8785 | 2275 | 0.069 | 0.1858 | No | ||
| 38 | NDEL1 | 20399 | 2330 | 0.066 | 0.1845 | No | ||
| 39 | ELF1 | 21948 | 2389 | 0.063 | 0.1829 | No | ||
| 40 | PHLPP | 14166 | 2422 | 0.061 | 0.1827 | No | ||
| 41 | MAP3K2 | 11165 | 2447 | 0.060 | 0.1828 | No | ||
| 42 | CUTL1 | 3507 3480 8820 3609 4577 | 2749 | 0.045 | 0.1676 | No | ||
| 43 | ETS1 | 10715 6230 3135 | 2762 | 0.044 | 0.1680 | No | ||
| 44 | RPS6KB1 | 7815 1207 13040 | 2778 | 0.044 | 0.1683 | No | ||
| 45 | CDH2 | 1963 8727 8726 4508 | 2780 | 0.044 | 0.1693 | No | ||
| 46 | PIP5K3 | 14234 | 2807 | 0.043 | 0.1689 | No | ||
| 47 | SORCS1 | 7184 12203 | 2870 | 0.041 | 0.1666 | No | ||
| 48 | USP47 | 6794 | 2879 | 0.041 | 0.1672 | No | ||
| 49 | SCN5A | 5412 | 2936 | 0.039 | 0.1651 | No | ||
| 50 | SFRS1 | 8492 | 2955 | 0.038 | 0.1650 | No | ||
| 51 | PCNP | 390 | 2956 | 0.038 | 0.1660 | No | ||
| 52 | RNF12 | 5384 | 3013 | 0.036 | 0.1638 | No | ||
| 53 | U2AF1 | 8431 | 3059 | 0.035 | 0.1623 | No | ||
| 54 | CCT8 | 22551 | 3240 | 0.031 | 0.1532 | No | ||
| 55 | MBNL2 | 21932 | 3427 | 0.026 | 0.1437 | No | ||
| 56 | STK17B | 13957 | 3443 | 0.026 | 0.1436 | No | ||
| 57 | SESTD1 | 5999 5998 | 3526 | 0.024 | 0.1397 | No | ||
| 58 | PDE4D | 10722 6235 | 3548 | 0.024 | 0.1392 | No | ||
| 59 | RFX1 | 18548 | 3611 | 0.023 | 0.1364 | No | ||
| 60 | SFRS6 | 14751 | 3612 | 0.023 | 0.1369 | No | ||
| 61 | CPEB2 | 6077 | 3705 | 0.022 | 0.1325 | No | ||
| 62 | PFN2 | 15337 1818 | 3810 | 0.020 | 0.1273 | No | ||
| 63 | SON | 5473 1657 1684 | 3979 | 0.018 | 0.1186 | No | ||
| 64 | HES5 | 15973 | 4316 | 0.015 | 0.1007 | No | ||
| 65 | NCAM1 | 5149 | 4368 | 0.014 | 0.0983 | No | ||
| 66 | HOXA7 | 4862 | 4398 | 0.014 | 0.0971 | No | ||
| 67 | RSBN1 | 6035 | 4638 | 0.012 | 0.0844 | No | ||
| 68 | BHLHB3 | 16935 | 4651 | 0.012 | 0.0840 | No | ||
| 69 | SULF2 | 12988 | 4670 | 0.012 | 0.0833 | No | ||
| 70 | DCAMKL1 | 8838 4600 | 4716 | 0.012 | 0.0812 | No | ||
| 71 | BTBD3 | 10457 2975 6007 6006 | 4719 | 0.012 | 0.0814 | No | ||
| 72 | FOSL1 | 23779 | 4732 | 0.012 | 0.0810 | No | ||
| 73 | DACH1 | 4595 3371 | 4821 | 0.011 | 0.0765 | No | ||
| 74 | TBC1D15 | 19625 | 4843 | 0.011 | 0.0756 | No | ||
| 75 | LPP | 5570 | 4898 | 0.011 | 0.0730 | No | ||
| 76 | MAFF | 22423 2243 | 4949 | 0.011 | 0.0705 | No | ||
| 77 | SEH1L | 13001 | 4986 | 0.010 | 0.0688 | No | ||
| 78 | DNAJC6 | 7828 13056 | 5044 | 0.010 | 0.0660 | No | ||
| 79 | SERTAD2 | 12200 7182 | 5049 | 0.010 | 0.0660 | No | ||
| 80 | IER5 | 13802 | 5119 | 0.010 | 0.0625 | No | ||
| 81 | JMJD1C | 11531 19996 | 5228 | 0.009 | 0.0568 | No | ||
| 82 | SNAP25 | 5466 | 5247 | 0.009 | 0.0561 | No | ||
| 83 | PTPN2 | 23414 | 5444 | 0.008 | 0.0456 | No | ||
| 84 | ABHD3 | 23491 | 5621 | 0.008 | 0.0362 | No | ||
| 85 | SOCS5 | 1619 23141 | 5821 | 0.007 | 0.0256 | No | ||
| 86 | EGR3 | 4656 | 5831 | 0.007 | 0.0252 | No | ||
| 87 | GABRA1 | 20492 | 5938 | 0.006 | 0.0196 | No | ||
| 88 | ABCA1 | 15881 | 5975 | 0.006 | 0.0178 | No | ||
| 89 | ZFHX4 | 8158 | 6002 | 0.006 | 0.0166 | No | ||
| 90 | ARL4A | 8627 | 6114 | 0.006 | 0.0107 | No | ||
| 91 | GRM3 | 16929 | 6126 | 0.006 | 0.0102 | No | ||
| 92 | SEMA4G | 11182 | 6276 | 0.005 | 0.0022 | No | ||
| 93 | LRRN1 | 17343 | 6290 | 0.005 | 0.0017 | No | ||
| 94 | HOXD3 | 9115 | 6372 | 0.005 | -0.0026 | No | ||
| 95 | TIPARP | 15576 | 6386 | 0.005 | -0.0032 | No | ||
| 96 | SYNPR | 22094 | 6465 | 0.005 | -0.0073 | No | ||
| 97 | CNTNAP2 | 17463 12406 | 6519 | 0.005 | -0.0101 | No | ||
| 98 | SBF2 | 11485 | 6531 | 0.005 | -0.0106 | No | ||
| 99 | CITED2 | 5118 14477 | 6561 | 0.005 | -0.0120 | No | ||
| 100 | SLC35F1 | 20027 | 6895 | 0.004 | -0.0300 | No | ||
| 101 | PDZRN3 | 17054 | 7188 | 0.003 | -0.0459 | No | ||
| 102 | KCNMA1 | 4947 | 7243 | 0.003 | -0.0487 | No | ||
| 103 | TOX | 15943 | 7326 | 0.003 | -0.0531 | No | ||
| 104 | LRCH4 | 10544 6090 | 7362 | 0.003 | -0.0550 | No | ||
| 105 | NDFIP1 | 23573 | 7412 | 0.003 | -0.0576 | No | ||
| 106 | KLF11 | 9700 | 7488 | 0.002 | -0.0616 | No | ||
| 107 | UBE2E3 | 5816 5815 | 7555 | 0.002 | -0.0651 | No | ||
| 108 | TEAD1 | 18121 | 7570 | 0.002 | -0.0658 | No | ||
| 109 | SDC2 | 9134 | 7813 | 0.002 | -0.0790 | No | ||
| 110 | UBQLN2 | 896 24223 897 | 7830 | 0.002 | -0.0798 | No | ||
| 111 | POU4F2 | 9603 5276 | 7871 | 0.002 | -0.0819 | No | ||
| 112 | SLC35F5 | 13171 | 7901 | 0.002 | -0.0835 | No | ||
| 113 | MTSS1 | 5585 22285 | 8257 | 0.001 | -0.1028 | No | ||
| 114 | PTN | 5323 | 8323 | 0.001 | -0.1063 | No | ||
| 115 | DAPK1 | 7632 | 8443 | 0.000 | -0.1127 | No | ||
| 116 | RKHD3 | 18199 | 8478 | 0.000 | -0.1146 | No | ||
| 117 | PHF15 | 20467 8050 | 8488 | 0.000 | -0.1151 | No | ||
| 118 | DKK2 | 15418 | 8555 | 0.000 | -0.1187 | No | ||
| 119 | PTP4A1 | 9658 | 8725 | -0.000 | -0.1278 | No | ||
| 120 | HDGFRP3 | 6621 | 8756 | -0.000 | -0.1295 | No | ||
| 121 | GIPC2 | 15137 | 9072 | -0.001 | -0.1466 | No | ||
| 122 | PURA | 9670 | 9497 | -0.002 | -0.1696 | No | ||
| 123 | POLR2B | 16817 | 9609 | -0.002 | -0.1756 | No | ||
| 124 | COCH | 21277 | 9631 | -0.002 | -0.1767 | No | ||
| 125 | NRXN3 | 2153 5196 | 9670 | -0.002 | -0.1787 | No | ||
| 126 | EIF2C1 | 10672 | 9701 | -0.002 | -0.1803 | No | ||
| 127 | SYT1 | 5565 | 9705 | -0.002 | -0.1804 | No | ||
| 128 | TRPC4 | 15593 | 9756 | -0.002 | -0.1831 | No | ||
| 129 | ENAH | 4665 8901 | 9896 | -0.003 | -0.1906 | No | ||
| 130 | NR5A2 | 4105 13817 | 9940 | -0.003 | -0.1928 | No | ||
| 131 | FBXO22 | 19442 | 9953 | -0.003 | -0.1934 | No | ||
| 132 | BIVM | 14256 | 10125 | -0.003 | -0.2026 | No | ||
| 133 | SP3 | 5483 | 10169 | -0.003 | -0.2049 | No | ||
| 134 | TCF7L2 | 10048 5646 | 10213 | -0.003 | -0.2072 | No | ||
| 135 | TMEM47 | 5315 | 10281 | -0.004 | -0.2107 | No | ||
| 136 | NOPE | 19407 | 10414 | -0.004 | -0.2178 | No | ||
| 137 | LIN28 | 15723 | 10416 | -0.004 | -0.2178 | No | ||
| 138 | CASP8AP2 | 16253 | 10454 | -0.004 | -0.2197 | No | ||
| 139 | ABCC12 | 18804 | 10460 | -0.004 | -0.2198 | No | ||
| 140 | SRC | 5507 | 10684 | -0.005 | -0.2319 | No | ||
| 141 | STX12 | 4140 | 10692 | -0.005 | -0.2321 | No | ||
| 142 | KLF12 | 4960 2637 21732 9228 | 11065 | -0.006 | -0.2522 | No | ||
| 143 | USP33 | 5036 | 11206 | -0.006 | -0.2597 | No | ||
| 144 | PRDM1 | 19775 3337 | 11218 | -0.006 | -0.2602 | No | ||
| 145 | NPAS4 | 23970 3683 | 11263 | -0.006 | -0.2624 | No | ||
| 146 | KDELR1 | 18241 | 11267 | -0.006 | -0.2624 | No | ||
| 147 | SOX11 | 5477 | 11376 | -0.006 | -0.2681 | No | ||
| 148 | CHES1 | 21007 11200 | 11432 | -0.007 | -0.2710 | No | ||
| 149 | CBX4 | 20128 | 11433 | -0.007 | -0.2708 | No | ||
| 150 | GADD45A | 17129 | 11554 | -0.007 | -0.2772 | No | ||
| 151 | PRPF4B | 3208 5295 | 11605 | -0.007 | -0.2797 | No | ||
| 152 | SLC1A3 | 22331 | 11606 | -0.007 | -0.2795 | No | ||
| 153 | SMOC2 | 23378 | 11608 | -0.007 | -0.2794 | No | ||
| 154 | PDCL | 14602 | 11627 | -0.007 | -0.2802 | No | ||
| 155 | TLOC1 | 12787 | 11634 | -0.007 | -0.2804 | No | ||
| 156 | GNAO1 | 4785 3829 | 11779 | -0.008 | -0.2880 | No | ||
| 157 | SLITRK3 | 11816 | 11829 | -0.008 | -0.2905 | No | ||
| 158 | HOXB5 | 20687 | 11882 | -0.008 | -0.2931 | No | ||
| 159 | SOX1 | 18685 | 11935 | -0.008 | -0.2957 | No | ||
| 160 | MDM1 | 3306 3370 3316 3405 3376 19870 3429 3398 3406 3428 3346 355 | 11949 | -0.008 | -0.2962 | No | ||
| 161 | IGF1 | 3352 9156 3409 | 12045 | -0.009 | -0.3012 | No | ||
| 162 | CBX3 | 8705 4484 | 12183 | -0.009 | -0.3084 | No | ||
| 163 | PPP1CB | 5282 3529 3504 | 12226 | -0.010 | -0.3104 | No | ||
| 164 | H3F3A | 4838 | 12342 | -0.010 | -0.3165 | No | ||
| 165 | ODC1 | 9500 | 12373 | -0.010 | -0.3178 | No | ||
| 166 | NEBL | 14678 2691 | 12493 | -0.011 | -0.3240 | No | ||
| 167 | MLLT10 | 9393 5104 | 12647 | -0.012 | -0.3321 | No | ||
| 168 | OSBPL10 | 13213 | 12661 | -0.012 | -0.3325 | No | ||
| 169 | GPM6A | 3834 10618 | 12666 | -0.012 | -0.3324 | No | ||
| 170 | KIF11 | 23873 | 12700 | -0.012 | -0.3339 | No | ||
| 171 | NHLH2 | 9462 | 12748 | -0.012 | -0.3362 | No | ||
| 172 | QKI | 23128 5340 | 12898 | -0.013 | -0.3440 | No | ||
| 173 | ATXN2 | 9780 3506 | 13203 | -0.015 | -0.3601 | No | ||
| 174 | ADD1 | 3615 4350 3590 | 13428 | -0.017 | -0.3719 | No | ||
| 175 | RFX4 | 7721 3410 | 13532 | -0.018 | -0.3771 | No | ||
| 176 | RFX3 | 9722 5377 3712 3762 | 13568 | -0.018 | -0.3785 | No | ||
| 177 | SLITRK2 | 24322 | 13737 | -0.020 | -0.3872 | No | ||
| 178 | FBN2 | 23433 | 13831 | -0.021 | -0.3917 | No | ||
| 179 | ZZZ3 | 15389 8445 4253 8444 1761 | 13916 | -0.022 | -0.3958 | No | ||
| 180 | ZNF22 | 12497 7417 | 13965 | -0.022 | -0.3978 | No | ||
| 181 | ETV5 | 22630 | 14115 | -0.024 | -0.4054 | No | ||
| 182 | PPARGC1A | 16533 | 14181 | -0.025 | -0.4083 | No | ||
| 183 | ATXN1 | 21477 | 14221 | -0.025 | -0.4098 | No | ||
| 184 | ADCYAP1 | 4348 | 14257 | -0.026 | -0.4110 | No | ||
| 185 | HOXB6 | 20688 | 14311 | -0.027 | -0.4133 | No | ||
| 186 | NFYB | 5173 | 14334 | -0.027 | -0.4138 | No | ||
| 187 | DDIT4L | 15410 | 14341 | -0.027 | -0.4134 | No | ||
| 188 | RAI14 | 22327 2191 | 14368 | -0.028 | -0.4142 | No | ||
| 189 | HNRPDL | 11947 3551 6954 | 14582 | -0.032 | -0.4250 | No | ||
| 190 | HTR7 | 23690 | 14690 | -0.034 | -0.4300 | No | ||
| 191 | XPO5 | 7802 | 14729 | -0.035 | -0.4312 | No | ||
| 192 | PIWIL4 | 142 | 14780 | -0.036 | -0.4330 | No | ||
| 193 | SP4 | 20972 | 14839 | -0.037 | -0.4352 | No | ||
| 194 | ELAVL1 | 4888 | 14882 | -0.038 | -0.4366 | No | ||
| 195 | NEUROG1 | 21446 | 14906 | -0.039 | -0.4369 | No | ||
| 196 | SERPINF2 | 9589 20348 | 14936 | -0.040 | -0.4375 | No | ||
| 197 | EIF4G2 | 1908 8892 | 14938 | -0.040 | -0.4365 | No | ||
| 198 | NPTX1 | 20126 | 14973 | -0.040 | -0.4374 | No | ||
| 199 | PDCD4 | 5232 23816 | 15088 | -0.043 | -0.4425 | No | ||
| 200 | NEUROG2 | 15426 | 15191 | -0.046 | -0.4469 | No | ||
| 201 | UBE2L3 | 5818 22647 | 15237 | -0.048 | -0.4482 | No | ||
| 202 | GPM6B | 9034 | 15473 | -0.055 | -0.4596 | No | ||
| 203 | MTF2 | 5129 | 15581 | -0.060 | -0.4639 | No | ||
| 204 | BZW1 | 7356 | 15594 | -0.061 | -0.4631 | No | ||
| 205 | SCN3A | 14573 | 15793 | -0.069 | -0.4721 | No | ||
| 206 | UBE3A | 1209 1084 1830 | 15942 | -0.077 | -0.4783 | No | ||
| 207 | TUSC2 | 19321 | 16030 | -0.081 | -0.4810 | No | ||
| 208 | LRBA | 15562 | 16234 | -0.095 | -0.4897 | No | ||
| 209 | ADIPOR1 | 14126 | 16342 | -0.101 | -0.4930 | No | ||
| 210 | TRAM1 | 13016 14000 | 16533 | -0.115 | -0.5005 | No | ||
| 211 | CDC42BPB | 20982 | 16673 | -0.129 | -0.5049 | No | ||
| 212 | UBE3C | 16896 | 16767 | -0.137 | -0.5065 | No | ||
| 213 | ARL8B | 7396 | 16782 | -0.139 | -0.5038 | No | ||
| 214 | TMPO | 5778 | 16908 | -0.155 | -0.5068 | Yes | ||
| 215 | RCE1 | 23965 | 16936 | -0.157 | -0.5044 | Yes | ||
| 216 | PPM1D | 20721 | 16943 | -0.158 | -0.5008 | Yes | ||
| 217 | SIRT1 | 19745 | 16953 | -0.159 | -0.4974 | Yes | ||
| 218 | FKBP1A | 2801 | 17167 | -0.182 | -0.5044 | Yes | ||
| 219 | CSTF3 | 10449 6002 | 17213 | -0.186 | -0.5023 | Yes | ||
| 220 | PPP2CA | 20890 | 17280 | -0.195 | -0.5010 | Yes | ||
| 221 | FNTA | 18904 | 17356 | -0.204 | -0.5001 | Yes | ||
| 222 | SCD | 23662 | 17470 | -0.221 | -0.5007 | Yes | ||
| 223 | TXN2 | 22227 | 17695 | -0.252 | -0.5067 | Yes | ||
| 224 | VGLL4 | 10558 | 17755 | -0.265 | -0.5033 | Yes | ||
| 225 | TPST1 | 10214 | 17781 | -0.271 | -0.4980 | Yes | ||
| 226 | MAPKAP1 | 15030 5993 | 17841 | -0.284 | -0.4942 | Yes | ||
| 227 | CLNS1A | 4526 4525 | 17871 | -0.293 | -0.4885 | Yes | ||
| 228 | CMTM4 | 3786 18777 | 17907 | -0.300 | -0.4830 | Yes | ||
| 229 | ATF4 | 22417 2239 | 17935 | -0.307 | -0.4768 | Yes | ||
| 230 | ZFX | 5984 | 17959 | -0.312 | -0.4703 | Yes | ||
| 231 | DHX32 | 17587 | 18011 | -0.327 | -0.4650 | Yes | ||
| 232 | MOCS2 | 21558 | 18085 | -0.351 | -0.4603 | Yes | ||
| 233 | LRP11 | 20095 | 18205 | -0.424 | -0.4563 | Yes | ||
| 234 | TRPS1 | 8195 | 18282 | -0.473 | -0.4487 | Yes | ||
| 235 | EIF3S10 | 4659 8887 | 18500 | -0.972 | -0.4364 | Yes | ||
| 236 | SMOC1 | 21221 | 18518 | -1.131 | -0.4093 | Yes | ||
| 237 | KCNK1 | 18415 | 18529 | -1.258 | -0.3787 | Yes | ||
| 238 | ACTL6A | 12136 13068 16662 3457 3554 | 18535 | -1.332 | -0.3460 | Yes | ||
| 239 | RDX | 9712 5371 | 18544 | -1.380 | -0.3122 | Yes | ||
| 240 | PPP3CA | 1863 5284 | 18554 | -1.410 | -0.2778 | Yes | ||
| 241 | PHF6 | 2648 24340 | 18581 | -2.048 | -0.2284 | Yes | ||
| 242 | HNRPM | 1511 13370 | 18607 | -4.087 | -0.1286 | Yes | ||
| 243 | UHRF1 | 5185 9477 | 18614 | -5.207 | 0.0001 | Yes |