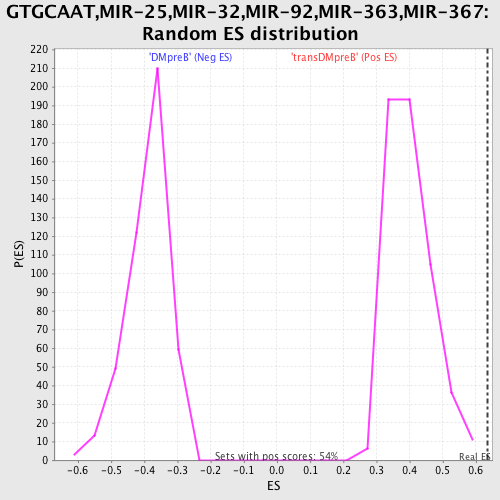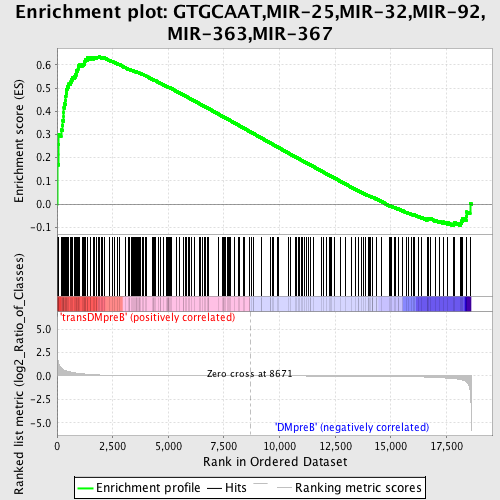
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMpreB_versus_DMpreB.cls#transDMpreB_versus_DMpreB_repos |
| Upregulated in class | transDMpreB |
| GeneSet | GTGCAAT,MIR-25,MIR-32,MIR-92,MIR-363,MIR-367 |
| Enrichment Score (ES) | 0.6344358 |
| Normalized Enrichment Score (NES) | 1.5879751 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.32756147 |
| FWER p-Value | 0.62 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | MYH9 | 2252 2244 | 1 | 5.596 | 0.1717 | Yes | ||
| 2 | FBXW7 | 1805 11928 | 65 | 1.463 | 0.2132 | Yes | ||
| 3 | BTBD12 | 22684 | 66 | 1.450 | 0.2577 | Yes | ||
| 4 | TRAF3 | 21147 | 76 | 1.340 | 0.2984 | Yes | ||
| 5 | FNBP4 | 14956 2671 | 177 | 0.869 | 0.3196 | Yes | ||
| 6 | ITPR1 | 17341 | 237 | 0.732 | 0.3389 | Yes | ||
| 7 | HERC2 | 9085 | 257 | 0.697 | 0.3593 | Yes | ||
| 8 | VPS54 | 20941 | 296 | 0.640 | 0.3769 | Yes | ||
| 9 | OGT | 4241 24274 | 300 | 0.635 | 0.3962 | Yes | ||
| 10 | RNF44 | 8364 | 306 | 0.631 | 0.4153 | Yes | ||
| 11 | KLF2 | 9229 | 336 | 0.596 | 0.4320 | Yes | ||
| 12 | BCL11A | 4691 | 380 | 0.557 | 0.4468 | Yes | ||
| 13 | CIC | 18338 | 388 | 0.550 | 0.4633 | Yes | ||
| 14 | DOCK9 | 8369 | 401 | 0.537 | 0.4791 | Yes | ||
| 15 | SLC9A1 | 16053 | 425 | 0.520 | 0.4938 | Yes | ||
| 16 | REV3L | 20050 | 451 | 0.499 | 0.5078 | Yes | ||
| 17 | PCTK1 | 24369 | 529 | 0.457 | 0.5176 | Yes | ||
| 18 | BRMS1L | 21268 | 603 | 0.410 | 0.5263 | Yes | ||
| 19 | NFIX | 9458 | 630 | 0.396 | 0.5370 | Yes | ||
| 20 | RAB14 | 12685 | 678 | 0.374 | 0.5459 | Yes | ||
| 21 | MYO18A | 20766 | 790 | 0.326 | 0.5499 | Yes | ||
| 22 | NLK | 5179 5178 | 838 | 0.311 | 0.5569 | Yes | ||
| 23 | MCL1 | 15502 | 854 | 0.306 | 0.5655 | Yes | ||
| 24 | ATP2A2 | 4421 3481 | 863 | 0.303 | 0.5743 | Yes | ||
| 25 | PPP1R12A | 3354 19886 | 936 | 0.281 | 0.5790 | Yes | ||
| 26 | TOB1 | 20703 | 942 | 0.278 | 0.5873 | Yes | ||
| 27 | BAZ2A | 4371 | 950 | 0.276 | 0.5954 | Yes | ||
| 28 | POLS | 9963 | 1017 | 0.256 | 0.5997 | Yes | ||
| 29 | MAN2A1 | 23170 | 1124 | 0.227 | 0.6009 | Yes | ||
| 30 | FMN2 | 14037 | 1197 | 0.212 | 0.6035 | Yes | ||
| 31 | ADAM19 | 8548 | 1225 | 0.206 | 0.6083 | Yes | ||
| 32 | POLK | 21384 | 1246 | 0.203 | 0.6135 | Yes | ||
| 33 | RPS6KA4 | 23804 | 1261 | 0.199 | 0.6188 | Yes | ||
| 34 | NELF | 12174 | 1272 | 0.197 | 0.6243 | Yes | ||
| 35 | NSMAF | 15944 | 1354 | 0.179 | 0.6254 | Yes | ||
| 36 | SRPR | 12525 7433 | 1367 | 0.178 | 0.6303 | Yes | ||
| 37 | TOB2 | 7162 | 1500 | 0.159 | 0.6280 | Yes | ||
| 38 | ADM | 18125 | 1508 | 0.157 | 0.6324 | Yes | ||
| 39 | P2RY13 | 15332 | 1648 | 0.136 | 0.6290 | Yes | ||
| 40 | SGPP1 | 21039 | 1688 | 0.131 | 0.6309 | Yes | ||
| 41 | SNF1LK | 23033 | 1762 | 0.123 | 0.6307 | Yes | ||
| 42 | TSGA14 | 17194 | 1783 | 0.119 | 0.6333 | Yes | ||
| 43 | APRIN | 4145 | 1853 | 0.110 | 0.6329 | Yes | ||
| 44 | PHTF2 | 16599 | 1886 | 0.106 | 0.6344 | Yes | ||
| 45 | E2F3 | 21500 8874 | 2002 | 0.092 | 0.6310 | No | ||
| 46 | DDX3X | 24379 | 2034 | 0.089 | 0.6321 | No | ||
| 47 | PITPNC1 | 7759 | 2112 | 0.082 | 0.6304 | No | ||
| 48 | GATA2 | 17370 | 2362 | 0.064 | 0.6188 | No | ||
| 49 | BCAT2 | 18244 | 2479 | 0.057 | 0.6143 | No | ||
| 50 | IQWD1 | 13778 | 2574 | 0.052 | 0.6108 | No | ||
| 51 | NRF1 | 17499 | 2724 | 0.046 | 0.6041 | No | ||
| 52 | PIP5K3 | 14234 | 2807 | 0.043 | 0.6010 | No | ||
| 53 | IXL | 17910 | 3070 | 0.035 | 0.5878 | No | ||
| 54 | ROBO2 | 6501 | 3210 | 0.031 | 0.5812 | No | ||
| 55 | TSC1 | 7235 | 3237 | 0.031 | 0.5807 | No | ||
| 56 | TRAK2 | 12903 13946 | 3243 | 0.030 | 0.5814 | No | ||
| 57 | HCN2 | 19965 | 3360 | 0.028 | 0.5759 | No | ||
| 58 | SSFA2 | 7692 | 3382 | 0.027 | 0.5756 | No | ||
| 59 | TACC2 | 18052 2534 2234 2435 2350 | 3445 | 0.026 | 0.5731 | No | ||
| 60 | WBSCR18 | 7254 12289 | 3497 | 0.025 | 0.5711 | No | ||
| 61 | PTPRO | 1093 17256 | 3535 | 0.024 | 0.5698 | No | ||
| 62 | HIVEP1 | 8486 | 3551 | 0.024 | 0.5697 | No | ||
| 63 | NFAT5 | 3921 7037 12036 | 3565 | 0.024 | 0.5697 | No | ||
| 64 | RFX1 | 18548 | 3611 | 0.023 | 0.5680 | No | ||
| 65 | CSMD3 | 10735 7938 | 3666 | 0.022 | 0.5658 | No | ||
| 66 | CPEB2 | 6077 | 3705 | 0.022 | 0.5644 | No | ||
| 67 | DSCAML1 | 4338 13224 | 3724 | 0.022 | 0.5641 | No | ||
| 68 | RNF38 | 7862 2328 13115 | 3736 | 0.021 | 0.5641 | No | ||
| 69 | KLHL18 | 11291 | 3849 | 0.020 | 0.5586 | No | ||
| 70 | PIP5K1C | 5250 19929 | 3871 | 0.020 | 0.5581 | No | ||
| 71 | CREB1 | 3990 8782 4558 4093 | 3985 | 0.018 | 0.5525 | No | ||
| 72 | LYST | 9325 5040 | 4011 | 0.018 | 0.5517 | No | ||
| 73 | PAX3 | 13910 | 4275 | 0.015 | 0.5379 | No | ||
| 74 | ARMC1 | 7907 | 4343 | 0.015 | 0.5347 | No | ||
| 75 | SLC32A1 | 14757 | 4349 | 0.014 | 0.5349 | No | ||
| 76 | ARRDC4 | 12346 | 4379 | 0.014 | 0.5337 | No | ||
| 77 | SLC38A2 | 22149 2219 | 4432 | 0.014 | 0.5313 | No | ||
| 78 | ATRX | 10350 5922 2628 | 4574 | 0.013 | 0.5241 | No | ||
| 79 | CD2AP | 22975 | 4626 | 0.013 | 0.5217 | No | ||
| 80 | RSBN1 | 6035 | 4638 | 0.012 | 0.5215 | No | ||
| 81 | DDX3Y | 6507 4593 | 4802 | 0.011 | 0.5130 | No | ||
| 82 | CREB3L2 | 17182 | 4934 | 0.011 | 0.5062 | No | ||
| 83 | KLF4 | 9230 4962 4963 | 4951 | 0.011 | 0.5056 | No | ||
| 84 | ITGA5 | 22105 | 4968 | 0.010 | 0.5051 | No | ||
| 85 | RBPMS2 | 19399 | 4996 | 0.010 | 0.5039 | No | ||
| 86 | OAZ3 | 15255 | 5010 | 0.010 | 0.5035 | No | ||
| 87 | USP28 | 19465 | 5022 | 0.010 | 0.5033 | No | ||
| 88 | SERTAD2 | 12200 7182 | 5049 | 0.010 | 0.5022 | No | ||
| 89 | RGS17 | 7133 | 5103 | 0.010 | 0.4996 | No | ||
| 90 | DLGAP2 | 10856 6350 | 5138 | 0.010 | 0.4980 | No | ||
| 91 | CPEB3 | 5539 | 5367 | 0.009 | 0.4859 | No | ||
| 92 | TCF21 | 3437 | 5497 | 0.008 | 0.4791 | No | ||
| 93 | PTGER4 | 9649 | 5665 | 0.007 | 0.4703 | No | ||
| 94 | BMPR2 | 14241 4081 | 5701 | 0.007 | 0.4686 | No | ||
| 95 | MYLIP | 21652 3189 | 5777 | 0.007 | 0.4648 | No | ||
| 96 | SOCS5 | 1619 23141 | 5821 | 0.007 | 0.4626 | No | ||
| 97 | LUZP1 | 6533 11257 2503 | 5912 | 0.007 | 0.4580 | No | ||
| 98 | GATA6 | 96 | 5946 | 0.006 | 0.4564 | No | ||
| 99 | ITGAV | 4931 | 6031 | 0.006 | 0.4520 | No | ||
| 100 | HAND2 | 18615 | 6162 | 0.006 | 0.4451 | No | ||
| 101 | EGR2 | 8886 | 6388 | 0.005 | 0.4330 | No | ||
| 102 | PIK3AP1 | 23679 | 6452 | 0.005 | 0.4298 | No | ||
| 103 | CNNM4 | 14273 | 6551 | 0.005 | 0.4246 | No | ||
| 104 | CNTN4 | 11265 1088 17345 | 6610 | 0.005 | 0.4216 | No | ||
| 105 | NR4A3 | 9473 16212 5183 | 6652 | 0.004 | 0.4195 | No | ||
| 106 | CXCL5 | 16802 | 6750 | 0.004 | 0.4144 | No | ||
| 107 | MEF2D | 9379 | 6793 | 0.004 | 0.4122 | No | ||
| 108 | GDF11 | 9011 | 6809 | 0.004 | 0.4115 | No | ||
| 109 | HOXB8 | 9111 | 7274 | 0.003 | 0.3864 | No | ||
| 110 | FMR1 | 24321 | 7414 | 0.003 | 0.3789 | No | ||
| 111 | SMAD7 | 23513 | 7464 | 0.002 | 0.3763 | No | ||
| 112 | PCDH11X | 2559 | 7492 | 0.002 | 0.3749 | No | ||
| 113 | DAB2IP | 12808 2692 | 7511 | 0.002 | 0.3740 | No | ||
| 114 | TEAD1 | 18121 | 7570 | 0.002 | 0.3710 | No | ||
| 115 | RHPN2 | 18293 | 7680 | 0.002 | 0.3651 | No | ||
| 116 | PPP1R12C | 6115 17987 | 7694 | 0.002 | 0.3644 | No | ||
| 117 | GOLGA4 | 12022 | 7726 | 0.002 | 0.3628 | No | ||
| 118 | SDC2 | 9134 | 7813 | 0.002 | 0.3582 | No | ||
| 119 | FZD10 | 13565 16693 | 7982 | 0.001 | 0.3491 | No | ||
| 120 | DNAJB9 | 6565 | 8167 | 0.001 | 0.3392 | No | ||
| 121 | HAS3 | 18477 | 8177 | 0.001 | 0.3387 | No | ||
| 122 | EDG1 | 15187 | 8398 | 0.001 | 0.3268 | No | ||
| 123 | PCAF | 5226 9532 5225 | 8442 | 0.000 | 0.3244 | No | ||
| 124 | SIM2 | 22693 | 8625 | 0.000 | 0.3146 | No | ||
| 125 | SESN3 | 19561 | 8626 | 0.000 | 0.3146 | No | ||
| 126 | COL12A1 | 3143 19054 | 8716 | -0.000 | 0.3097 | No | ||
| 127 | EPHA8 | 15705 | 8807 | -0.000 | 0.3049 | No | ||
| 128 | WRNIP1 | 21675 3161 | 8833 | -0.000 | 0.3035 | No | ||
| 129 | PRKCE | 9575 | 9168 | -0.001 | 0.2854 | No | ||
| 130 | FASLG | 13789 | 9169 | -0.001 | 0.2854 | No | ||
| 131 | ADAM10 | 4336 | 9582 | -0.002 | 0.2631 | No | ||
| 132 | MYO1B | 13962 | 9592 | -0.002 | 0.2627 | No | ||
| 133 | HHIP | 4849 | 9672 | -0.002 | 0.2585 | No | ||
| 134 | TCF2 | 1395 20727 | 9721 | -0.002 | 0.2559 | No | ||
| 135 | DSC2 | 8865 | 9895 | -0.003 | 0.2466 | No | ||
| 136 | KIF5B | 4955 2031 | 9960 | -0.003 | 0.2432 | No | ||
| 137 | FHL2 | 13966 | 10410 | -0.004 | 0.2189 | No | ||
| 138 | BTG2 | 13832 | 10489 | -0.004 | 0.2148 | No | ||
| 139 | SLC12A5 | 14731 | 10708 | -0.005 | 0.2031 | No | ||
| 140 | COL1A2 | 8772 | 10779 | -0.005 | 0.1995 | No | ||
| 141 | PLEKHA6 | 10786 6288 | 10830 | -0.005 | 0.1969 | No | ||
| 142 | BSN | 18997 | 10877 | -0.005 | 0.1946 | No | ||
| 143 | NOX4 | 18196 | 10969 | -0.005 | 0.1898 | No | ||
| 144 | SYN2 | 5559 | 11008 | -0.005 | 0.1879 | No | ||
| 145 | CXXC5 | 12523 | 11128 | -0.006 | 0.1816 | No | ||
| 146 | ATXN3 | 21001 2051 | 11195 | -0.006 | 0.1782 | No | ||
| 147 | NFIA | 16172 5170 | 11299 | -0.006 | 0.1728 | No | ||
| 148 | HAND1 | 20440 | 11367 | -0.006 | 0.1694 | No | ||
| 149 | SOX11 | 5477 | 11376 | -0.006 | 0.1691 | No | ||
| 150 | EXOC5 | 8370 | 11507 | -0.007 | 0.1623 | No | ||
| 151 | DPP10 | 13850 | 11861 | -0.008 | 0.1434 | No | ||
| 152 | TEF | 2287 22414 2192 | 11992 | -0.009 | 0.1366 | No | ||
| 153 | SLC17A6 | 18226 | 12125 | -0.009 | 0.1297 | No | ||
| 154 | CBLN4 | 14326 | 12233 | -0.010 | 0.1241 | No | ||
| 155 | FBN1 | 14449 | 12298 | -0.010 | 0.1210 | No | ||
| 156 | MOAP1 | 12252 | 12339 | -0.010 | 0.1191 | No | ||
| 157 | ARHGEF17 | 17737 | 12483 | -0.011 | 0.1117 | No | ||
| 158 | GAP43 | 9001 | 12726 | -0.012 | 0.0989 | No | ||
| 159 | TRAM2 | 13990 | 12955 | -0.014 | 0.0869 | No | ||
| 160 | EDEM1 | 17339 938 | 13249 | -0.015 | 0.0715 | No | ||
| 161 | CEBPA | 4514 | 13397 | -0.017 | 0.0640 | No | ||
| 162 | GRIA3 | 24349 11996 2649 | 13541 | -0.018 | 0.0568 | No | ||
| 163 | GFPT2 | 4775 | 13543 | -0.018 | 0.0573 | No | ||
| 164 | SLC24A3 | 14818 | 13703 | -0.019 | 0.0492 | No | ||
| 165 | INSIG1 | 6073 10525 10526 | 13780 | -0.020 | 0.0457 | No | ||
| 166 | ANP32E | 15500 | 13882 | -0.021 | 0.0409 | No | ||
| 167 | LBX1 | 98 23659 | 13974 | -0.022 | 0.0366 | No | ||
| 168 | LHFPL2 | 21584 | 14006 | -0.023 | 0.0357 | No | ||
| 169 | MARK1 | 5955 | 14035 | -0.023 | 0.0348 | No | ||
| 170 | LATS2 | 11923 6928 | 14069 | -0.023 | 0.0338 | No | ||
| 171 | DUSP10 | 4003 14016 | 14161 | -0.024 | 0.0296 | No | ||
| 172 | PITPNA | 20778 | 14177 | -0.025 | 0.0295 | No | ||
| 173 | PIK3R3 | 5248 | 14193 | -0.025 | 0.0295 | No | ||
| 174 | RGS3 | 2434 2474 6937 11935 2374 2332 2445 2390 | 14351 | -0.027 | 0.0218 | No | ||
| 175 | BCL2L11 | 2790 14861 | 14353 | -0.027 | 0.0226 | No | ||
| 176 | STK39 | 14570 | 14588 | -0.032 | 0.0108 | No | ||
| 177 | PER2 | 3940 13881 | 14951 | -0.040 | -0.0076 | No | ||
| 178 | NPTX1 | 20126 | 14973 | -0.040 | -0.0075 | No | ||
| 179 | ADCY3 | 21329 894 | 15046 | -0.042 | -0.0101 | No | ||
| 180 | PRDM13 | 15933 | 15162 | -0.045 | -0.0150 | No | ||
| 181 | RNF4 | 5385 9729 | 15206 | -0.047 | -0.0159 | No | ||
| 182 | RAB23 | 14282 | 15332 | -0.050 | -0.0211 | No | ||
| 183 | EOMES | 4669 | 15345 | -0.050 | -0.0202 | No | ||
| 184 | RAD21 | 22298 | 15539 | -0.058 | -0.0289 | No | ||
| 185 | XYLT2 | 20288 | 15688 | -0.065 | -0.0350 | No | ||
| 186 | SCN3A | 14573 | 15793 | -0.069 | -0.0385 | No | ||
| 187 | SMAD6 | 19083 | 15935 | -0.076 | -0.0438 | No | ||
| 188 | NFIB | 15855 | 15999 | -0.079 | -0.0448 | No | ||
| 189 | SOX4 | 5479 | 16049 | -0.082 | -0.0449 | No | ||
| 190 | DNAJB12 | 7146 | 16248 | -0.096 | -0.0528 | No | ||
| 191 | ZDHHC5 | 14547 | 16390 | -0.105 | -0.0572 | No | ||
| 192 | IBTK | 19047 | 16630 | -0.125 | -0.0664 | No | ||
| 193 | UGP2 | 20518 | 16654 | -0.127 | -0.0637 | No | ||
| 194 | MAP2K4 | 20405 | 16690 | -0.131 | -0.0616 | No | ||
| 195 | RAP1B | 10083 | 16790 | -0.140 | -0.0626 | No | ||
| 196 | PGAM1 | 9556 | 16992 | -0.164 | -0.0685 | No | ||
| 197 | RNF141 | 12478 7392 | 17196 | -0.185 | -0.0739 | No | ||
| 198 | CCNE2 | 4491 | 17367 | -0.205 | -0.0768 | No | ||
| 199 | CCNC | 16263 | 17557 | -0.232 | -0.0800 | No | ||
| 200 | SERTAD3 | 18327 | 17820 | -0.280 | -0.0856 | No | ||
| 201 | ARRDC3 | 4191 | 17852 | -0.287 | -0.0785 | No | ||
| 202 | DYNLT3 | 24187 | 18124 | -0.369 | -0.0819 | No | ||
| 203 | TULP4 | 12740 | 18186 | -0.413 | -0.0725 | No | ||
| 204 | CDKN1C | 17546 | 18217 | -0.433 | -0.0609 | No | ||
| 205 | CD69 | 16979 8717 | 18386 | -0.589 | -0.0519 | No | ||
| 206 | ELOVL4 | 19048 | 18416 | -0.643 | -0.0338 | No | ||
| 207 | PAFAH1B1 | 1340 5220 9524 | 18560 | -1.453 | 0.0030 | No |