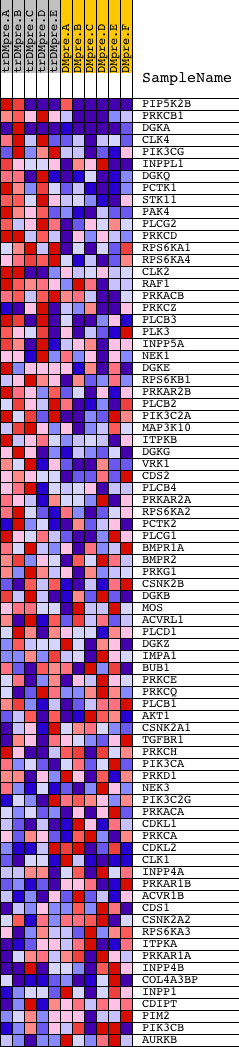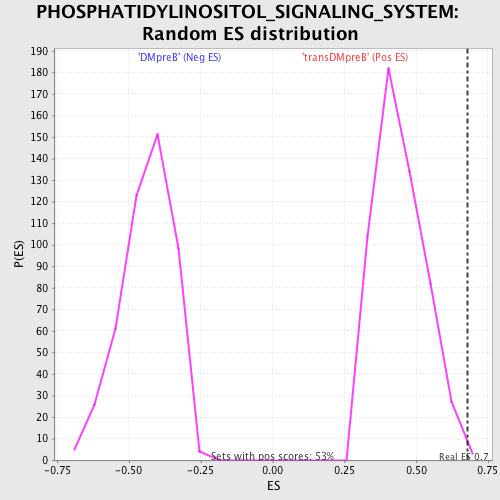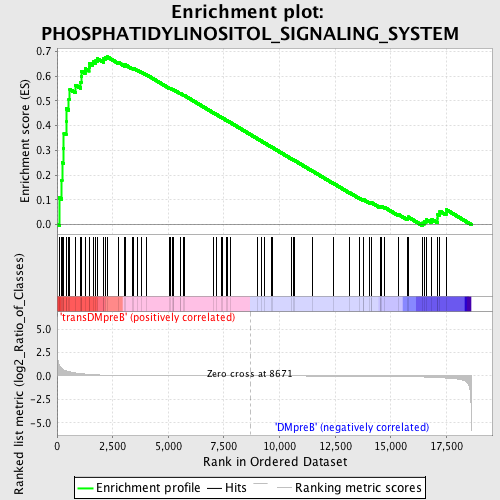
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMpreB_versus_DMpreB.cls#transDMpreB_versus_DMpreB_repos |
| Upregulated in class | transDMpreB |
| GeneSet | PHOSPHATIDYLINOSITOL_SIGNALING_SYSTEM |
| Enrichment Score (ES) | 0.6777191 |
| Normalized Enrichment Score (NES) | 1.5344516 |
| Nominal p-value | 0.0018796993 |
| FDR q-value | 0.7678623 |
| FWER p-Value | 0.95 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PIP5K2B | 20269 1254 | 98 | 1.165 | 0.1083 | Yes | ||
| 2 | PRKCB1 | 1693 9574 | 210 | 0.777 | 0.1781 | Yes | ||
| 3 | DGKA | 3359 19589 | 222 | 0.756 | 0.2512 | Yes | ||
| 4 | CLK4 | 20895 1292 | 298 | 0.637 | 0.3093 | Yes | ||
| 5 | PIK3CG | 6635 | 307 | 0.628 | 0.3701 | Yes | ||
| 6 | INPPL1 | 17733 3775 | 405 | 0.535 | 0.4171 | Yes | ||
| 7 | DGKQ | 4287 | 412 | 0.530 | 0.4684 | Yes | ||
| 8 | PCTK1 | 24369 | 529 | 0.457 | 0.5067 | Yes | ||
| 9 | STK11 | 19950 | 574 | 0.431 | 0.5464 | Yes | ||
| 10 | PAK4 | 17909 | 822 | 0.317 | 0.5640 | Yes | ||
| 11 | PLCG2 | 18453 | 1057 | 0.247 | 0.5754 | Yes | ||
| 12 | PRKCD | 21897 | 1081 | 0.237 | 0.5972 | Yes | ||
| 13 | RPS6KA1 | 15725 | 1096 | 0.233 | 0.6192 | Yes | ||
| 14 | RPS6KA4 | 23804 | 1261 | 0.199 | 0.6298 | Yes | ||
| 15 | CLK2 | 1883 15544 | 1434 | 0.169 | 0.6370 | Yes | ||
| 16 | RAF1 | 17035 | 1454 | 0.166 | 0.6522 | Yes | ||
| 17 | PRKACB | 15140 | 1613 | 0.143 | 0.6576 | Yes | ||
| 18 | PRKCZ | 5260 | 1742 | 0.124 | 0.6628 | Yes | ||
| 19 | PLCB3 | 23799 | 1806 | 0.116 | 0.6707 | Yes | ||
| 20 | PLK3 | 15790 | 2077 | 0.085 | 0.6645 | Yes | ||
| 21 | INPP5A | 10003 | 2080 | 0.085 | 0.6727 | Yes | ||
| 22 | NEK1 | 18610 | 2170 | 0.077 | 0.6754 | Yes | ||
| 23 | DGKE | 7069 | 2256 | 0.071 | 0.6777 | Yes | ||
| 24 | RPS6KB1 | 7815 1207 13040 | 2778 | 0.044 | 0.6539 | No | ||
| 25 | PRKAR2B | 5288 2107 | 3033 | 0.036 | 0.6437 | No | ||
| 26 | PLCB2 | 5262 | 3051 | 0.035 | 0.6462 | No | ||
| 27 | PIK3C2A | 17667 | 3377 | 0.027 | 0.6313 | No | ||
| 28 | MAP3K10 | 3831 17916 | 3450 | 0.026 | 0.6299 | No | ||
| 29 | ITPKB | 3963 14028 | 3605 | 0.023 | 0.6239 | No | ||
| 30 | DGKG | 4282 | 3800 | 0.020 | 0.6154 | No | ||
| 31 | VRK1 | 5862 2049 | 4017 | 0.018 | 0.6055 | No | ||
| 32 | CDS2 | 14837 | 5041 | 0.010 | 0.5513 | No | ||
| 33 | PLCB4 | 9586 | 5101 | 0.010 | 0.5491 | No | ||
| 34 | PRKAR2A | 5287 | 5178 | 0.009 | 0.5459 | No | ||
| 35 | RPS6KA2 | 9759 9758 | 5211 | 0.009 | 0.5451 | No | ||
| 36 | PCTK2 | 19907 | 5238 | 0.009 | 0.5446 | No | ||
| 37 | PLCG1 | 14753 | 5548 | 0.008 | 0.5287 | No | ||
| 38 | BMPR1A | 8660 4450 | 5566 | 0.008 | 0.5286 | No | ||
| 39 | BMPR2 | 14241 4081 | 5701 | 0.007 | 0.5220 | No | ||
| 40 | PRKG1 | 5289 | 5731 | 0.007 | 0.5212 | No | ||
| 41 | CSNK2B | 23008 | 7011 | 0.004 | 0.4525 | No | ||
| 42 | DGKB | 5722 | 7150 | 0.003 | 0.4454 | No | ||
| 43 | MOS | 16254 | 7390 | 0.003 | 0.4328 | No | ||
| 44 | ACVRL1 | 22354 | 7452 | 0.003 | 0.4297 | No | ||
| 45 | PLCD1 | 18972 | 7594 | 0.002 | 0.4223 | No | ||
| 46 | DGKZ | 2836 14522 | 7645 | 0.002 | 0.4198 | No | ||
| 47 | IMPA1 | 12066 | 7812 | 0.002 | 0.4110 | No | ||
| 48 | BUB1 | 8665 | 9007 | -0.001 | 0.3467 | No | ||
| 49 | PRKCE | 9575 | 9168 | -0.001 | 0.3382 | No | ||
| 50 | PRKCQ | 2873 2831 | 9311 | -0.001 | 0.3306 | No | ||
| 51 | PLCB1 | 14832 2821 | 9655 | -0.002 | 0.3123 | No | ||
| 52 | AKT1 | 8568 | 9661 | -0.002 | 0.3123 | No | ||
| 53 | CSNK2A1 | 14797 | 10540 | -0.004 | 0.2653 | No | ||
| 54 | TGFBR1 | 16213 10165 5745 | 10618 | -0.004 | 0.2616 | No | ||
| 55 | PRKCH | 21246 | 10681 | -0.005 | 0.2587 | No | ||
| 56 | PIK3CA | 9562 | 11488 | -0.007 | 0.2159 | No | ||
| 57 | PRKD1 | 2074 21075 | 12406 | -0.010 | 0.1674 | No | ||
| 58 | NEK3 | 3778 18916 | 13154 | -0.015 | 0.1286 | No | ||
| 59 | PIK3C2G | 17253 | 13590 | -0.018 | 0.1069 | No | ||
| 60 | PRKACA | 18549 3844 | 13785 | -0.020 | 0.0984 | No | ||
| 61 | CDKL1 | 12914 | 13787 | -0.020 | 0.1003 | No | ||
| 62 | PRKCA | 20174 | 14043 | -0.023 | 0.0888 | No | ||
| 63 | CDKL2 | 16482 3473 12001 3464 | 14126 | -0.024 | 0.0868 | No | ||
| 64 | CLK1 | 13951 4127 4529 8748 | 14136 | -0.024 | 0.0886 | No | ||
| 65 | INPP4A | 3985 3935 11237 4085 | 14514 | -0.030 | 0.0712 | No | ||
| 66 | PRKAR1B | 16320 | 14524 | -0.031 | 0.0737 | No | ||
| 67 | ACVR1B | 4335 22353 | 14565 | -0.031 | 0.0746 | No | ||
| 68 | CDS1 | 16779 | 14709 | -0.034 | 0.0703 | No | ||
| 69 | CSNK2A2 | 4567 8808 | 15352 | -0.051 | 0.0406 | No | ||
| 70 | RPS6KA3 | 8490 | 15744 | -0.067 | 0.0261 | No | ||
| 71 | ITPKA | 14898 | 15774 | -0.069 | 0.0312 | No | ||
| 72 | PRKAR1A | 20615 | 16423 | -0.108 | 0.0067 | No | ||
| 73 | INPP4B | 18557 | 16500 | -0.113 | 0.0136 | No | ||
| 74 | COL4A3BP | 3221 7522 | 16583 | -0.120 | 0.0209 | No | ||
| 75 | INPP1 | 13961 | 16843 | -0.147 | 0.0214 | No | ||
| 76 | CDIPT | 18071 | 17110 | -0.176 | 0.0241 | No | ||
| 77 | PIM2 | 5249 9564 9565 | 17112 | -0.176 | 0.0412 | No | ||
| 78 | PIK3CB | 19030 | 17205 | -0.186 | 0.0544 | No | ||
| 79 | AURKB | 20832 | 17487 | -0.223 | 0.0609 | No |