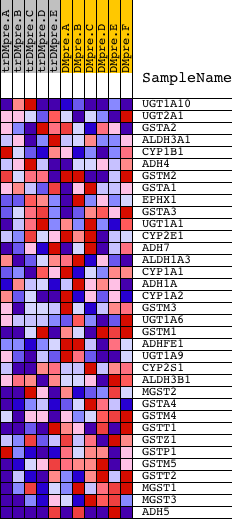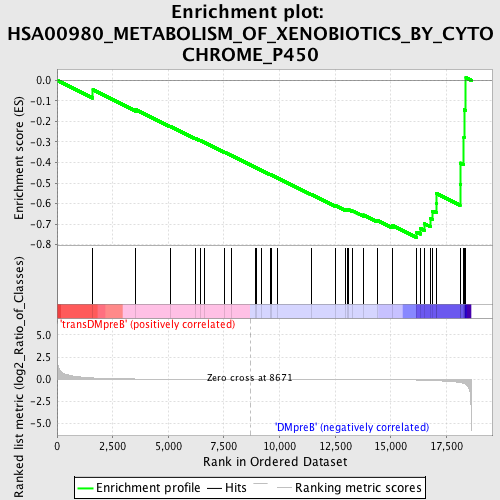
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMpreB_versus_DMpreB.cls#transDMpreB_versus_DMpreB_repos |
| Upregulated in class | DMpreB |
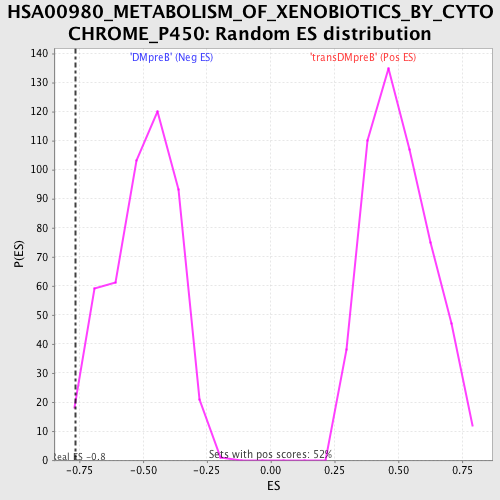
| GeneSet | HSA00980_METABOLISM_OF_XENOBIOTICS_BY_CYTOCHROME_P450 |
| Enrichment Score (ES) | -0.76481557 |
| Normalized Enrichment Score (NES) | -1.5310328 |
| Nominal p-value | 0.008403362 |
| FDR q-value | 0.53111356 |
| FWER p-Value | 0.975 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | UGT1A10 | 6908 | 1612 | 0.143 | -0.0467 | No | ||
| 2 | UGT2A1 | 8247 | 3539 | 0.024 | -0.1436 | No | ||
| 3 | GSTA2 | 4808 6173 | 5094 | 0.010 | -0.2245 | No | ||
| 4 | ALDH3A1 | 20854 | 6219 | 0.006 | -0.2834 | No | ||
| 5 | CYP1B1 | 4581 | 6441 | 0.005 | -0.2939 | No | ||
| 6 | ADH4 | 15405 | 6645 | 0.004 | -0.3036 | No | ||
| 7 | GSTM2 | 9049 4810 | 7506 | 0.002 | -0.3492 | No | ||
| 8 | GSTA1 | 4807 | 7854 | 0.002 | -0.3674 | No | ||
| 9 | EPHX1 | 13734 | 8898 | -0.000 | -0.4234 | No | ||
| 10 | GSTA3 | 14287 | 8943 | -0.001 | -0.4257 | No | ||
| 11 | UGT1A1 | 11851 | 9203 | -0.001 | -0.4393 | No | ||
| 12 | CYP2E1 | 18025 | 9584 | -0.002 | -0.4592 | No | ||
| 13 | ADH7 | 15408 | 9627 | -0.002 | -0.4609 | No | ||
| 14 | ALDH1A3 | 17802 | 9916 | -0.003 | -0.4756 | No | ||
| 15 | CYP1A1 | 19428 | 11445 | -0.007 | -0.5560 | No | ||
| 16 | ADH1A | 15406 | 12529 | -0.011 | -0.6112 | No | ||
| 17 | CYP1A2 | 19097 | 12956 | -0.014 | -0.6303 | No | ||
| 18 | GSTM3 | 9050 | 13053 | -0.014 | -0.6315 | No | ||
| 19 | UGT1A6 | 3969 4079 6911 13591 | 13108 | -0.014 | -0.6304 | No | ||
| 20 | GSTM1 | 4809 9048 | 13294 | -0.016 | -0.6360 | No | ||
| 21 | ADHFE1 | 4049 8008 | 13760 | -0.020 | -0.6554 | No | ||
| 22 | UGT1A9 | 11849 11850 | 14380 | -0.028 | -0.6809 | No | ||
| 23 | CYP2S1 | 17921 | 15083 | -0.043 | -0.7066 | No | ||
| 24 | ALDH3B1 | 12569 23949 | 16166 | -0.090 | -0.7397 | Yes | ||
| 25 | MGST2 | 15597 | 16326 | -0.100 | -0.7202 | Yes | ||
| 26 | GSTA4 | 19371 | 16524 | -0.115 | -0.6988 | Yes | ||
| 27 | GSTM4 | 15199 | 16783 | -0.140 | -0.6737 | Yes | ||
| 28 | GSTT1 | 19730 | 16872 | -0.151 | -0.6362 | Yes | ||
| 29 | GSTZ1 | 21193 | 17043 | -0.169 | -0.5981 | Yes | ||
| 30 | GSTP1 | 9051 4812 4811 | 17055 | -0.171 | -0.5509 | Yes | ||
| 31 | GSTM5 | 15455 | 18113 | -0.361 | -0.5070 | Yes | ||
| 32 | GSTT2 | 9052 | 18137 | -0.378 | -0.4026 | Yes | ||
| 33 | MGST1 | 17254 | 18284 | -0.474 | -0.2779 | Yes | ||
| 34 | MGST3 | 13771 12349 | 18298 | -0.481 | -0.1441 | Yes | ||
| 35 | ADH5 | 8554 15404 | 18377 | -0.577 | 0.0129 | Yes |