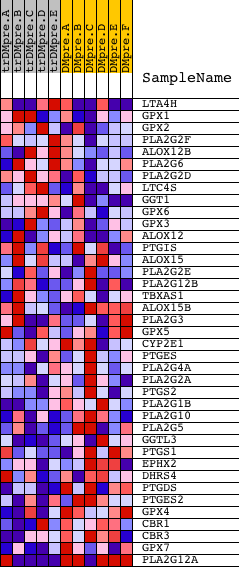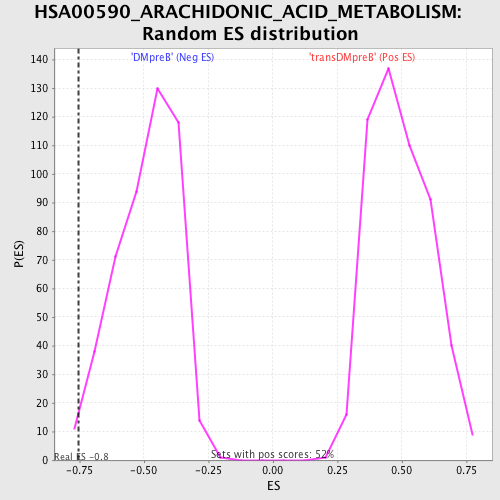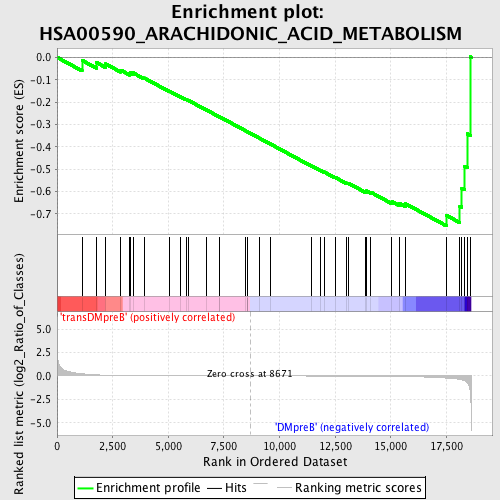
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMpreB_versus_DMpreB.cls#transDMpreB_versus_DMpreB_repos |
| Upregulated in class | DMpreB |
| GeneSet | HSA00590_ARACHIDONIC_ACID_METABOLISM |
| Enrichment Score (ES) | -0.75436723 |
| Normalized Enrichment Score (NES) | -1.5443896 |
| Nominal p-value | 0.01048218 |
| FDR q-value | 0.53964823 |
| FWER p-Value | 0.947 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | LTA4H | 19906 3374 | 1123 | 0.227 | -0.0135 | No | ||
| 2 | GPX1 | 19310 | 1767 | 0.122 | -0.0230 | No | ||
| 3 | GPX2 | 9037 | 2179 | 0.077 | -0.0293 | No | ||
| 4 | PLA2G2F | 15700 | 2857 | 0.041 | -0.0572 | No | ||
| 5 | ALOX12B | 20825 | 3268 | 0.030 | -0.0731 | No | ||
| 6 | PLA2G6 | 22209 | 3302 | 0.029 | -0.0689 | No | ||
| 7 | PLA2G2D | 16018 | 3413 | 0.026 | -0.0694 | No | ||
| 8 | LTC4S | 20478 | 3911 | 0.019 | -0.0922 | No | ||
| 9 | GGT1 | 19987 | 5043 | 0.010 | -0.1510 | No | ||
| 10 | GPX6 | 21698 | 5557 | 0.008 | -0.1770 | No | ||
| 11 | GPX3 | 20880 | 5803 | 0.007 | -0.1887 | No | ||
| 12 | ALOX12 | 20372 | 5900 | 0.007 | -0.1925 | No | ||
| 13 | PTGIS | 9653 | 6712 | 0.004 | -0.2353 | No | ||
| 14 | ALOX15 | 20370 | 7316 | 0.003 | -0.2672 | No | ||
| 15 | PLA2G2E | 16016 | 7319 | 0.003 | -0.2667 | No | ||
| 16 | PLA2G12B | 20016 | 8454 | 0.000 | -0.3277 | No | ||
| 17 | TBXAS1 | 17480 1105 | 8565 | 0.000 | -0.3335 | No | ||
| 18 | ALOX15B | 20397 | 8571 | 0.000 | -0.3338 | No | ||
| 19 | PLA2G3 | 20968 | 8577 | 0.000 | -0.3340 | No | ||
| 20 | GPX5 | 21532 3242 | 9105 | -0.001 | -0.3622 | No | ||
| 21 | CYP2E1 | 18025 | 9584 | -0.002 | -0.3875 | No | ||
| 22 | PTGES | 7224 | 11440 | -0.007 | -0.4860 | No | ||
| 23 | PLA2G4A | 13809 | 11823 | -0.008 | -0.5049 | No | ||
| 24 | PLA2G2A | 16017 | 12003 | -0.009 | -0.5127 | No | ||
| 25 | PTGS2 | 5317 9655 | 12502 | -0.011 | -0.5373 | No | ||
| 26 | PLA2G1B | 9581 | 13015 | -0.014 | -0.5620 | No | ||
| 27 | PLA2G10 | 22658 | 13082 | -0.014 | -0.5626 | No | ||
| 28 | PLA2G5 | 9583 | 13860 | -0.021 | -0.6001 | No | ||
| 29 | GGTL3 | 14381 | 13895 | -0.021 | -0.5975 | No | ||
| 30 | PTGS1 | 2726 5316 15028 | 14095 | -0.024 | -0.6034 | No | ||
| 31 | EPHX2 | 21778 | 15051 | -0.042 | -0.6460 | No | ||
| 32 | DHRS4 | 6609 | 15391 | -0.052 | -0.6535 | No | ||
| 33 | PTGDS | 14658 | 15653 | -0.063 | -0.6545 | No | ||
| 34 | PTGES2 | 15038 | 17509 | -0.226 | -0.7076 | Yes | ||
| 35 | GPX4 | 9038 | 18102 | -0.357 | -0.6657 | Yes | ||
| 36 | CBR1 | 22697 | 18190 | -0.415 | -0.5847 | Yes | ||
| 37 | CBR3 | 22696 | 18324 | -0.504 | -0.4878 | Yes | ||
| 38 | GPX7 | 15809 | 18449 | -0.736 | -0.3423 | Yes | ||
| 39 | PLA2G12A | 1782 7281 1839 | 18573 | -1.700 | 0.0023 | Yes |