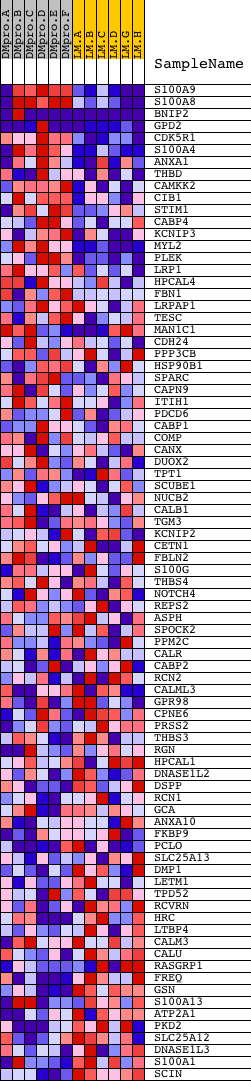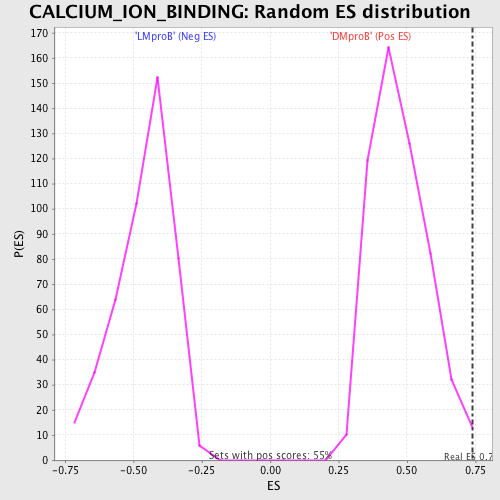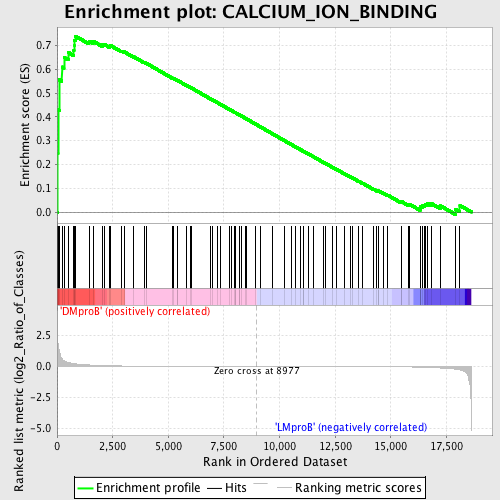
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_DMproB_versus_LMproB.phenotype_DMproB_versus_LMproB.cls #DMproB_versus_LMproB |
| Phenotype | phenotype_DMproB_versus_LMproB.cls#DMproB_versus_LMproB |
| Upregulated in class | DMproB |
| GeneSet | CALCIUM_ION_BINDING |
| Enrichment Score (ES) | 0.73918915 |
| Normalized Enrichment Score (NES) | 1.561031 |
| Nominal p-value | 0.0036630037 |
| FDR q-value | 0.37769368 |
| FWER p-Value | 0.988 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | S100A9 | 9774 | 22 | 2.490 | 0.2454 | Yes | ||
| 2 | S100A8 | 15527 | 40 | 1.871 | 0.4297 | Yes | ||
| 3 | BNIP2 | 179 19388 | 95 | 1.304 | 0.5559 | Yes | ||
| 4 | GPD2 | 4773 9014 | 219 | 0.608 | 0.6095 | Yes | ||
| 5 | CDK5R1 | 20745 | 310 | 0.455 | 0.6497 | Yes | ||
| 6 | S100A4 | 9773 | 501 | 0.313 | 0.6704 | Yes | ||
| 7 | ANXA1 | 9282 | 752 | 0.223 | 0.6791 | Yes | ||
| 8 | THBD | 14404 | 761 | 0.220 | 0.7004 | Yes | ||
| 9 | CAMKK2 | 9870 | 768 | 0.217 | 0.7216 | Yes | ||
| 10 | CIB1 | 17780 | 823 | 0.207 | 0.7392 | Yes | ||
| 11 | STIM1 | 18164 | 1434 | 0.118 | 0.7179 | No | ||
| 12 | CABP4 | 13125 | 1619 | 0.100 | 0.7179 | No | ||
| 13 | KCNIP3 | 14441 | 2028 | 0.070 | 0.7028 | No | ||
| 14 | MYL2 | 16713 | 2114 | 0.064 | 0.7046 | No | ||
| 15 | PLEK | 7072 | 2374 | 0.053 | 0.6959 | No | ||
| 16 | LRP1 | 9284 | 2391 | 0.052 | 0.7001 | No | ||
| 17 | HPCAL4 | 16096 | 2892 | 0.034 | 0.6765 | No | ||
| 18 | FBN1 | 14449 | 3013 | 0.031 | 0.6731 | No | ||
| 19 | LRPAP1 | 5005 | 3434 | 0.023 | 0.6528 | No | ||
| 20 | TESC | 16725 | 3926 | 0.017 | 0.6280 | No | ||
| 21 | MAN1C1 | 10514 6064 15714 | 4020 | 0.016 | 0.6245 | No | ||
| 22 | CDH24 | 10726 | 5173 | 0.009 | 0.5633 | No | ||
| 23 | PPP3CB | 5285 | 5226 | 0.009 | 0.5613 | No | ||
| 24 | HSP90B1 | 19657 | 5409 | 0.008 | 0.5523 | No | ||
| 25 | SPARC | 20444 | 5422 | 0.008 | 0.5525 | No | ||
| 26 | CAPN9 | 3772 18422 | 5822 | 0.007 | 0.5316 | No | ||
| 27 | ITIH1 | 21895 | 5989 | 0.006 | 0.5233 | No | ||
| 28 | PDCD6 | 21404 | 6046 | 0.006 | 0.5209 | No | ||
| 29 | CABP1 | 16412 | 6896 | 0.004 | 0.4754 | No | ||
| 30 | COMP | 18589 | 6991 | 0.004 | 0.4707 | No | ||
| 31 | CANX | 4474 | 7200 | 0.003 | 0.4598 | No | ||
| 32 | DUOX2 | 14453 | 7323 | 0.003 | 0.4536 | No | ||
| 33 | TPT1 | 5799 | 7741 | 0.002 | 0.4313 | No | ||
| 34 | SCUBE1 | 7231 | 7831 | 0.002 | 0.4267 | No | ||
| 35 | NUCB2 | 18114 | 7983 | 0.002 | 0.4187 | No | ||
| 36 | CALB1 | 4469 | 8012 | 0.002 | 0.4174 | No | ||
| 37 | TGM3 | 10168 | 8035 | 0.002 | 0.4164 | No | ||
| 38 | KCNIP2 | 23655 3725 | 8206 | 0.001 | 0.4073 | No | ||
| 39 | CETN1 | 6443 | 8208 | 0.001 | 0.4074 | No | ||
| 40 | FBLN2 | 994 8956 | 8265 | 0.001 | 0.4045 | No | ||
| 41 | S100G | 24016 | 8475 | 0.001 | 0.3933 | No | ||
| 42 | THBS4 | 21393 | 8494 | 0.001 | 0.3924 | No | ||
| 43 | NOTCH4 | 23277 9475 | 8924 | 0.000 | 0.3693 | No | ||
| 44 | REPS2 | 24018 | 9121 | -0.000 | 0.3587 | No | ||
| 45 | ASPH | 2498 2460 2396 2370 2357 2502 2516 2375 2337 12276 2381 2454 7246 2480 2393 | 9691 | -0.001 | 0.3281 | No | ||
| 46 | SPOCK2 | 13585 | 10212 | -0.002 | 0.3003 | No | ||
| 47 | PPM2C | 491 | 10534 | -0.003 | 0.2832 | No | ||
| 48 | CALR | 18814 | 10699 | -0.003 | 0.2747 | No | ||
| 49 | CABP2 | 23765 | 10961 | -0.003 | 0.2609 | No | ||
| 50 | RCN2 | 19438 | 11086 | -0.004 | 0.2546 | No | ||
| 51 | CALML3 | 21553 | 11091 | -0.004 | 0.2547 | No | ||
| 52 | GPR98 | 21400 | 11298 | -0.004 | 0.2440 | No | ||
| 53 | CPNE6 | 22010 | 11511 | -0.005 | 0.2330 | No | ||
| 54 | PRSS2 | 17474 | 11982 | -0.006 | 0.2082 | No | ||
| 55 | THBS3 | 15542 1796 | 12082 | -0.006 | 0.2035 | No | ||
| 56 | RGN | 24371 | 12375 | -0.007 | 0.1884 | No | ||
| 57 | HPCAL1 | 21307 2123 | 12553 | -0.007 | 0.1795 | No | ||
| 58 | DNASE1L2 | 960 23092 | 12918 | -0.008 | 0.1607 | No | ||
| 59 | DSPP | 8867 | 13172 | -0.009 | 0.1480 | No | ||
| 60 | RCN1 | 9710 | 13263 | -0.010 | 0.1441 | No | ||
| 61 | GCA | 2813 15003 10720 7830 | 13544 | -0.011 | 0.1300 | No | ||
| 62 | ANXA10 | 18869 | 13720 | -0.012 | 0.1218 | No | ||
| 63 | FKBP9 | 17433 | 14230 | -0.015 | 0.0958 | No | ||
| 64 | PCLO | 11207 6495 | 14375 | -0.016 | 0.0897 | No | ||
| 65 | SLC25A13 | 17221 1159 | 14428 | -0.017 | 0.0885 | No | ||
| 66 | DMP1 | 16776 | 14457 | -0.017 | 0.0887 | No | ||
| 67 | LETM1 | 7108 16563 12114 | 14692 | -0.019 | 0.0780 | No | ||
| 68 | TPD52 | 18830 5794 | 14854 | -0.021 | 0.0713 | No | ||
| 69 | RCVRN | 20835 | 15462 | -0.031 | 0.0416 | No | ||
| 70 | HRC | 18251 | 15498 | -0.032 | 0.0429 | No | ||
| 71 | LTBP4 | 1739 17918 | 15791 | -0.040 | 0.0311 | No | ||
| 72 | CALM3 | 8682 | 15850 | -0.041 | 0.0321 | No | ||
| 73 | CALU | 1124 1149 8683 1162 4472 | 16315 | -0.060 | 0.0130 | No | ||
| 74 | RASGRP1 | 5360 14476 9694 | 16333 | -0.061 | 0.0182 | No | ||
| 75 | FREQ | 4734 8983 15046 | 16339 | -0.062 | 0.0240 | No | ||
| 76 | GSN | 2784 | 16427 | -0.065 | 0.0258 | No | ||
| 77 | S100A13 | 15532 | 16514 | -0.070 | 0.0281 | No | ||
| 78 | ATP2A1 | 17639 | 16560 | -0.072 | 0.0328 | No | ||
| 79 | PKD2 | 16772 | 16641 | -0.077 | 0.0362 | No | ||
| 80 | SLC25A12 | 14562 | 16829 | -0.092 | 0.0352 | No | ||
| 81 | DNASE1L3 | 21925 | 17234 | -0.126 | 0.0259 | No | ||
| 82 | S100A1 | 15271 | 17906 | -0.215 | 0.0109 | No | ||
| 83 | SCIN | 21079 | 18107 | -0.276 | 0.0275 | No |