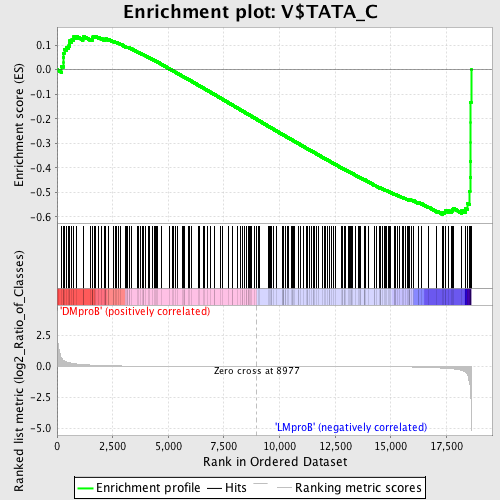
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_DMproB_versus_LMproB.phenotype_DMproB_versus_LMproB.cls #DMproB_versus_LMproB |
| Phenotype | phenotype_DMproB_versus_LMproB.cls#DMproB_versus_LMproB |
| Upregulated in class | LMproB |
| GeneSet | V$TATA_C |
| Enrichment Score (ES) | -0.590148 |
| Normalized Enrichment Score (NES) | -1.4433217 |
| Nominal p-value | 0.010460251 |
| FDR q-value | 0.85027444 |
| FWER p-Value | 0.999 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SSBP3 | 2439 7814 2431 | 197 | 0.657 | 0.0149 | No | ||
| 2 | MYLK | 22778 4213 | 277 | 0.494 | 0.0299 | No | ||
| 3 | LMO2 | 9281 | 280 | 0.490 | 0.0490 | No | ||
| 4 | CHD2 | 10847 | 290 | 0.478 | 0.0671 | No | ||
| 5 | IL1B | 14431 | 316 | 0.447 | 0.0832 | No | ||
| 6 | WSB2 | 7205 | 414 | 0.362 | 0.0921 | No | ||
| 7 | GCH1 | 9008 4762 | 531 | 0.296 | 0.0974 | No | ||
| 8 | CCR7 | 20254 | 561 | 0.284 | 0.1069 | No | ||
| 9 | GADD45G | 21637 | 573 | 0.279 | 0.1172 | No | ||
| 10 | CTDSP1 | 4121 10407 | 666 | 0.249 | 0.1219 | No | ||
| 11 | ANXA1 | 9282 | 752 | 0.223 | 0.1260 | No | ||
| 12 | H2AFX | 19477 | 753 | 0.223 | 0.1347 | No | ||
| 13 | SATB1 | 5408 | 886 | 0.195 | 0.1352 | No | ||
| 14 | LAPTM5 | 16061 | 1164 | 0.151 | 0.1260 | No | ||
| 15 | RANBP10 | 18764 | 1171 | 0.150 | 0.1315 | No | ||
| 16 | FOXP1 | 4242 | 1181 | 0.148 | 0.1368 | No | ||
| 17 | PPP1R12A | 3354 19886 | 1483 | 0.112 | 0.1248 | No | ||
| 18 | EGF | 15169 | 1577 | 0.103 | 0.1238 | No | ||
| 19 | HMGN2 | 9095 | 1595 | 0.102 | 0.1269 | No | ||
| 20 | TOB1 | 20703 | 1605 | 0.101 | 0.1304 | No | ||
| 21 | TCF4 | 5642 | 1612 | 0.100 | 0.1339 | No | ||
| 22 | PLA2G4B | 14896 | 1658 | 0.096 | 0.1353 | No | ||
| 23 | BACH1 | 8647 | 1733 | 0.090 | 0.1348 | No | ||
| 24 | BMI1 | 4448 15113 | 1873 | 0.081 | 0.1304 | No | ||
| 25 | CUEDC2 | 12469 | 2006 | 0.072 | 0.1260 | No | ||
| 26 | MYL2 | 16713 | 2114 | 0.064 | 0.1227 | No | ||
| 27 | HIVEP3 | 4973 | 2153 | 0.063 | 0.1231 | No | ||
| 28 | AGPAT4 | 23384 | 2164 | 0.062 | 0.1250 | No | ||
| 29 | HIVEP1 | 8486 | 2193 | 0.061 | 0.1258 | No | ||
| 30 | MKNK1 | 2504 | 2293 | 0.056 | 0.1227 | No | ||
| 31 | ATP5G2 | 12610 | 2308 | 0.056 | 0.1241 | No | ||
| 32 | OTUB2 | 7537 12654 | 2518 | 0.046 | 0.1145 | No | ||
| 33 | SOX5 | 9850 1044 5480 16937 | 2527 | 0.046 | 0.1159 | No | ||
| 34 | H3F3B | 4839 | 2645 | 0.042 | 0.1112 | No | ||
| 35 | TLE3 | 5767 19416 | 2668 | 0.041 | 0.1116 | No | ||
| 36 | HSD17B8 | 9061 | 2760 | 0.038 | 0.1081 | No | ||
| 37 | RAB33A | 358 | 2858 | 0.035 | 0.1042 | No | ||
| 38 | MYST3 | 18651 | 3086 | 0.030 | 0.0930 | No | ||
| 39 | STK39 | 14570 | 3133 | 0.029 | 0.0916 | No | ||
| 40 | KCNK12 | 9978 | 3172 | 0.028 | 0.0906 | No | ||
| 41 | CYB561D2 | 19001 | 3177 | 0.028 | 0.0915 | No | ||
| 42 | BIN1 | 6634 2017 | 3240 | 0.027 | 0.0892 | No | ||
| 43 | PKIA | 15639 | 3272 | 0.026 | 0.0885 | No | ||
| 44 | LOX | 23438 | 3360 | 0.024 | 0.0847 | No | ||
| 45 | MMP10 | 19571 | 3631 | 0.020 | 0.0709 | No | ||
| 46 | KCNA3 | 15460 | 3670 | 0.020 | 0.0696 | No | ||
| 47 | FBXO36 | 14205 | 3756 | 0.019 | 0.0657 | No | ||
| 48 | ALDOB | 10494 | 3852 | 0.018 | 0.0612 | No | ||
| 49 | SDPR | 14252 | 3863 | 0.018 | 0.0613 | No | ||
| 50 | HOXB9 | 20689 | 3875 | 0.017 | 0.0614 | No | ||
| 51 | CDC42EP2 | 4180 14771 | 3970 | 0.016 | 0.0570 | No | ||
| 52 | OTP | 21582 | 4085 | 0.015 | 0.0514 | No | ||
| 53 | MMP13 | 19573 3007 | 4141 | 0.015 | 0.0490 | No | ||
| 54 | MAPK10 | 11169 | 4304 | 0.014 | 0.0407 | No | ||
| 55 | HOXC6 | 22339 2302 | 4391 | 0.013 | 0.0365 | No | ||
| 56 | NR5A2 | 4105 13817 | 4432 | 0.013 | 0.0348 | No | ||
| 57 | PDGFRB | 23539 | 4486 | 0.012 | 0.0325 | No | ||
| 58 | KCTD15 | 17864 | 4498 | 0.012 | 0.0323 | No | ||
| 59 | LPL | 9283 5003 | 4683 | 0.011 | 0.0228 | No | ||
| 60 | MYH11 | 22656 | 5062 | 0.010 | 0.0026 | No | ||
| 61 | PHEX | 24024 | 5197 | 0.009 | -0.0043 | No | ||
| 62 | NFATC2 | 5168 2866 | 5215 | 0.009 | -0.0049 | No | ||
| 63 | ASB5 | 13322 | 5322 | 0.008 | -0.0103 | No | ||
| 64 | HOXA10 | 17145 989 | 5413 | 0.008 | -0.0149 | No | ||
| 65 | ELAVL2 | 15839 | 5650 | 0.007 | -0.0275 | No | ||
| 66 | FOXP3 | 9804 5426 | 5660 | 0.007 | -0.0277 | No | ||
| 67 | OTX2 | 9519 | 5727 | 0.007 | -0.0310 | No | ||
| 68 | FABP4 | 8601 | 5743 | 0.007 | -0.0315 | No | ||
| 69 | GNAZ | 71 | 5922 | 0.006 | -0.0409 | No | ||
| 70 | METAP1 | 15154 | 5953 | 0.006 | -0.0423 | No | ||
| 71 | KITLG | 19889 3342 | 6057 | 0.006 | -0.0477 | No | ||
| 72 | CAPN6 | 24041 | 6362 | 0.005 | -0.0640 | No | ||
| 73 | STC1 | 21974 | 6380 | 0.005 | -0.0647 | No | ||
| 74 | GTF3C1 | 10607 17646 | 6587 | 0.005 | -0.0757 | No | ||
| 75 | CFL2 | 2155 21069 | 6622 | 0.005 | -0.0774 | No | ||
| 76 | NPPA | 298 | 6741 | 0.004 | -0.0837 | No | ||
| 77 | PRDM16 | 12893 | 6904 | 0.004 | -0.0923 | No | ||
| 78 | BDNF | 14926 2797 | 7067 | 0.004 | -0.1010 | No | ||
| 79 | GPR50 | 24318 | 7072 | 0.004 | -0.1011 | No | ||
| 80 | DUOX2 | 14453 | 7323 | 0.003 | -0.1145 | No | ||
| 81 | SVIL | 1990 1942 10334 1976 23626 1946 1945 2041 | 7418 | 0.003 | -0.1195 | No | ||
| 82 | COL1A1 | 8771 | 7716 | 0.002 | -0.1356 | No | ||
| 83 | HOXD10 | 14983 | 7863 | 0.002 | -0.1434 | No | ||
| 84 | NEDD4 | 19384 3101 | 7900 | 0.002 | -0.1453 | No | ||
| 85 | PBX1 | 5224 | 8126 | 0.001 | -0.1575 | No | ||
| 86 | HAS2 | 22293 | 8227 | 0.001 | -0.1629 | No | ||
| 87 | PLUNC | 9593 | 8313 | 0.001 | -0.1674 | No | ||
| 88 | NRAP | 23642 | 8325 | 0.001 | -0.1680 | No | ||
| 89 | SHANK2 | 17998 | 8441 | 0.001 | -0.1742 | No | ||
| 90 | MYH3 | 20839 | 8513 | 0.001 | -0.1780 | No | ||
| 91 | PRDM1 | 19775 3337 | 8528 | 0.001 | -0.1788 | No | ||
| 92 | FSHB | 14500 | 8593 | 0.001 | -0.1822 | No | ||
| 93 | DDX5 | 4606 8843 4607 | 8604 | 0.001 | -0.1828 | No | ||
| 94 | HOXC5 | 22340 | 8636 | 0.001 | -0.1844 | No | ||
| 95 | NPY | 17449 | 8670 | 0.001 | -0.1862 | No | ||
| 96 | TAGLN | 19136 | 8748 | 0.000 | -0.1904 | No | ||
| 97 | CRABP2 | 15554 | 8886 | 0.000 | -0.1978 | No | ||
| 98 | SEMA3A | 16919 | 8963 | 0.000 | -0.2019 | No | ||
| 99 | ATP6V1G3 | 14107 | 9046 | -0.000 | -0.2064 | No | ||
| 100 | HAMP | 17878 | 9092 | -0.000 | -0.2088 | No | ||
| 101 | RFX4 | 7721 3410 | 9495 | -0.001 | -0.2306 | No | ||
| 102 | WDR20 | 21258 | 9544 | -0.001 | -0.2332 | No | ||
| 103 | RHOA | 8624 4409 4410 | 9583 | -0.001 | -0.2352 | No | ||
| 104 | EPAS1 | 23144 | 9649 | -0.001 | -0.2387 | No | ||
| 105 | JARID2 | 9200 | 9651 | -0.001 | -0.2387 | No | ||
| 106 | CDX1 | 23429 | 9719 | -0.001 | -0.2423 | No | ||
| 107 | PDHA2 | 15153 | 9872 | -0.001 | -0.2505 | No | ||
| 108 | AMY2A | 8587 | 10118 | -0.002 | -0.2638 | No | ||
| 109 | IL17F | 13992 | 10135 | -0.002 | -0.2646 | No | ||
| 110 | NOX3 | 10322 | 10154 | -0.002 | -0.2655 | No | ||
| 111 | HOXB8 | 9111 | 10171 | -0.002 | -0.2663 | No | ||
| 112 | MYH4 | 20837 | 10175 | -0.002 | -0.2664 | No | ||
| 113 | MYCT1 | 7572 | 10265 | -0.002 | -0.2711 | No | ||
| 114 | ATOH1 | 4419 | 10279 | -0.002 | -0.2717 | No | ||
| 115 | HNF4G | 15640 | 10349 | -0.002 | -0.2754 | No | ||
| 116 | VIP | 20096 | 10397 | -0.002 | -0.2779 | No | ||
| 117 | MMP3 | 19572 3148 | 10517 | -0.003 | -0.2842 | No | ||
| 118 | TAF7L | 24063 | 10559 | -0.003 | -0.2864 | No | ||
| 119 | A2BP1 | 11212 1652 1689 | 10571 | -0.003 | -0.2868 | No | ||
| 120 | BNIP3 | 17581 | 10612 | -0.003 | -0.2889 | No | ||
| 121 | EHF | 4657 | 10662 | -0.003 | -0.2915 | No | ||
| 122 | MMP20 | 19569 | 10828 | -0.003 | -0.3003 | No | ||
| 123 | DKK2 | 15418 | 10866 | -0.003 | -0.3022 | No | ||
| 124 | BMPR1B | 8661 15152 4451 | 10946 | -0.003 | -0.3064 | No | ||
| 125 | COL8A1 | 4548 | 11075 | -0.004 | -0.3132 | No | ||
| 126 | MYH2 | 20838 | 11220 | -0.004 | -0.3208 | No | ||
| 127 | ATF3 | 13720 | 11261 | -0.004 | -0.3229 | No | ||
| 128 | PITX2 | 15424 1878 | 11271 | -0.004 | -0.3232 | No | ||
| 129 | OLIG2 | 6953 | 11357 | -0.004 | -0.3276 | No | ||
| 130 | IL22 | 11948 | 11444 | -0.004 | -0.3321 | No | ||
| 131 | SLC12A2 | 23547 2026 | 11506 | -0.005 | -0.3353 | No | ||
| 132 | ITGB1BP2 | 24275 | 11539 | -0.005 | -0.3368 | No | ||
| 133 | NKX6-1 | 16463 | 11583 | -0.005 | -0.3390 | No | ||
| 134 | GPRC5D | 16962 | 11651 | -0.005 | -0.3424 | No | ||
| 135 | LMOD1 | 14119 | 11770 | -0.005 | -0.3487 | No | ||
| 136 | HOXB6 | 20688 | 11908 | -0.005 | -0.3559 | No | ||
| 137 | TRPC4 | 15593 | 12011 | -0.006 | -0.3612 | No | ||
| 138 | SIM1 | 20031 | 12041 | -0.006 | -0.3626 | No | ||
| 139 | RBP2 | 19347 | 12146 | -0.006 | -0.3680 | No | ||
| 140 | CD109 | 6174 10656 19369 | 12231 | -0.006 | -0.3723 | No | ||
| 141 | RHOH | 13226 | 12232 | -0.006 | -0.3721 | No | ||
| 142 | CD96 | 22580 | 12329 | -0.007 | -0.3770 | No | ||
| 143 | BARHL1 | 14634 | 12441 | -0.007 | -0.3828 | No | ||
| 144 | LDB2 | 16538 4989 | 12495 | -0.007 | -0.3854 | No | ||
| 145 | SLIT3 | 20925 | 12761 | -0.008 | -0.3995 | No | ||
| 146 | BCL11B | 7190 | 12808 | -0.008 | -0.4017 | No | ||
| 147 | KCNJ8 | 16940 | 12824 | -0.008 | -0.4022 | No | ||
| 148 | RAB15 | 21035 | 12903 | -0.008 | -0.4061 | No | ||
| 149 | VANGL1 | 6034 | 12925 | -0.008 | -0.4069 | No | ||
| 150 | AQP8 | 18087 | 12942 | -0.008 | -0.4075 | No | ||
| 151 | MYL1 | 4015 | 13087 | -0.009 | -0.4150 | No | ||
| 152 | TGFBR1 | 16213 10165 5745 | 13160 | -0.009 | -0.4185 | No | ||
| 153 | EVI1 | 4689 8926 | 13186 | -0.009 | -0.4195 | No | ||
| 154 | GJA5 | 1929 9017 4779 | 13233 | -0.009 | -0.4217 | No | ||
| 155 | RTN1 | 4177 2099 | 13237 | -0.009 | -0.4214 | No | ||
| 156 | TSHB | 15218 | 13287 | -0.010 | -0.4237 | No | ||
| 157 | EYA1 | 4695 4061 | 13430 | -0.010 | -0.4310 | No | ||
| 158 | ESR2 | 21038 | 13529 | -0.011 | -0.4360 | No | ||
| 159 | CITED1 | 24091 | 13580 | -0.011 | -0.4382 | No | ||
| 160 | LTBP1 | 1536 1604 6503 1559 1564 1518 | 13635 | -0.011 | -0.4407 | No | ||
| 161 | ABCA1 | 15881 | 13652 | -0.011 | -0.4412 | No | ||
| 162 | SETBP1 | 23401 | 13810 | -0.012 | -0.4492 | No | ||
| 163 | TGFB2 | 5743 | 13814 | -0.013 | -0.4489 | No | ||
| 164 | PURA | 9670 | 13829 | -0.013 | -0.4491 | No | ||
| 165 | OLIG3 | 20084 | 13839 | -0.013 | -0.4491 | No | ||
| 166 | KCNK3 | 4945 | 14010 | -0.014 | -0.4578 | No | ||
| 167 | ESRRG | 6447 | 14248 | -0.015 | -0.4701 | No | ||
| 168 | NEUROD1 | 14550 | 14259 | -0.015 | -0.4701 | No | ||
| 169 | MT3 | 9414 | 14376 | -0.016 | -0.4757 | No | ||
| 170 | HOXD9 | 14982 | 14509 | -0.018 | -0.4822 | No | ||
| 171 | PRKAR2B | 5288 2107 | 14517 | -0.018 | -0.4819 | No | ||
| 172 | ARHGEF2 | 4987 | 14526 | -0.018 | -0.4817 | No | ||
| 173 | TBX3 | 16723 | 14556 | -0.018 | -0.4825 | No | ||
| 174 | ELMO3 | 10629 18495 | 14614 | -0.018 | -0.4849 | No | ||
| 175 | HOXA9 | 17147 9109 1024 | 14728 | -0.019 | -0.4903 | No | ||
| 176 | PPAP2A | 21562 3258 3244 | 14750 | -0.020 | -0.4907 | No | ||
| 177 | PANK1 | 1687 23692 | 14811 | -0.020 | -0.4931 | No | ||
| 178 | TOPBP1 | 10659 | 14908 | -0.022 | -0.4975 | No | ||
| 179 | FGF23 | 17275 | 14925 | -0.022 | -0.4975 | No | ||
| 180 | GRK5 | 9036 23814 | 14977 | -0.023 | -0.4994 | No | ||
| 181 | CCL7 | 20743 | 15149 | -0.025 | -0.5077 | No | ||
| 182 | CDC26 | 7294 | 15167 | -0.025 | -0.5077 | No | ||
| 183 | PPARGC1B | 9318 | 15189 | -0.026 | -0.5078 | No | ||
| 184 | CD24 | 8711 | 15198 | -0.026 | -0.5072 | No | ||
| 185 | AKAP12 | 17600 | 15298 | -0.028 | -0.5115 | No | ||
| 186 | TWIST1 | 21291 | 15387 | -0.029 | -0.5152 | No | ||
| 187 | NKX2-5 | 23060 | 15512 | -0.032 | -0.5207 | No | ||
| 188 | ARRDC3 | 4191 | 15589 | -0.034 | -0.5235 | No | ||
| 189 | PACSIN3 | 14948 8149 2776 2665 | 15654 | -0.036 | -0.5255 | No | ||
| 190 | HOXB4 | 20686 | 15740 | -0.038 | -0.5287 | No | ||
| 191 | HNRPA0 | 13382 | 15807 | -0.040 | -0.5307 | No | ||
| 192 | PRPF4 | 7655 12845 | 15833 | -0.041 | -0.5304 | No | ||
| 193 | PA2G4 | 5265 | 15838 | -0.041 | -0.5291 | No | ||
| 194 | CYR61 | 9159 15145 9311 5024 | 15858 | -0.042 | -0.5285 | No | ||
| 195 | DIO2 | 21014 2159 | 15946 | -0.045 | -0.5314 | No | ||
| 196 | TGFB3 | 10161 | 16032 | -0.048 | -0.5342 | No | ||
| 197 | FIP1L1 | 888 16826 | 16224 | -0.056 | -0.5424 | No | ||
| 198 | PDYN | 14429 | 16257 | -0.058 | -0.5418 | No | ||
| 199 | HOXC4 | 4864 | 16394 | -0.064 | -0.5467 | No | ||
| 200 | SNRPF | 7645 | 16702 | -0.082 | -0.5602 | No | ||
| 201 | PPP2R5E | 6523 2136 | 17072 | -0.110 | -0.5760 | No | ||
| 202 | PRKAG1 | 22135 | 17334 | -0.137 | -0.5848 | Yes | ||
| 203 | SRPK2 | 5513 | 17366 | -0.141 | -0.5810 | Yes | ||
| 204 | MAPRE1 | 4652 | 17435 | -0.147 | -0.5789 | Yes | ||
| 205 | NDNL2 | 17804 | 17456 | -0.149 | -0.5742 | Yes | ||
| 206 | RAB5B | 359 19591 3430 | 17577 | -0.164 | -0.5743 | Yes | ||
| 207 | TCTA | 4167 | 17736 | -0.184 | -0.5757 | Yes | ||
| 208 | MRPS18B | 7365 | 17755 | -0.188 | -0.5694 | Yes | ||
| 209 | OTUB1 | 23795 | 17804 | -0.195 | -0.5644 | Yes | ||
| 210 | EEF2 | 8881 4654 8882 | 18189 | -0.311 | -0.5731 | Yes | ||
| 211 | PPP1R10 | 6972 11960 6971 | 18358 | -0.451 | -0.5646 | Yes | ||
| 212 | HERPUD1 | 18522 | 18438 | -0.621 | -0.5446 | Yes | ||
| 213 | EIF4A2 | 4660 1679 1645 | 18549 | -1.405 | -0.4958 | Yes | ||
| 214 | INVS | 2397 16211 | 18560 | -1.443 | -0.4399 | Yes | ||
| 215 | CRHBP | 8785 | 18573 | -1.695 | -0.3744 | Yes | ||
| 216 | SRF | 22961 1597 | 18585 | -2.048 | -0.2950 | Yes | ||
| 217 | EPC1 | 2039 8907 2033 | 18587 | -2.049 | -0.2150 | Yes | ||
| 218 | CA2 | 15631 1825 | 18589 | -2.059 | -0.1347 | Yes | ||
| 219 | ACTB | 8534 337 337 338 | 18610 | -3.484 | 0.0003 | Yes |

