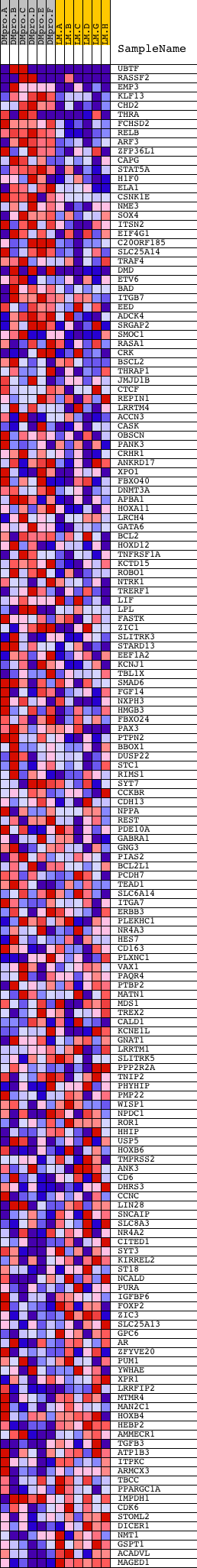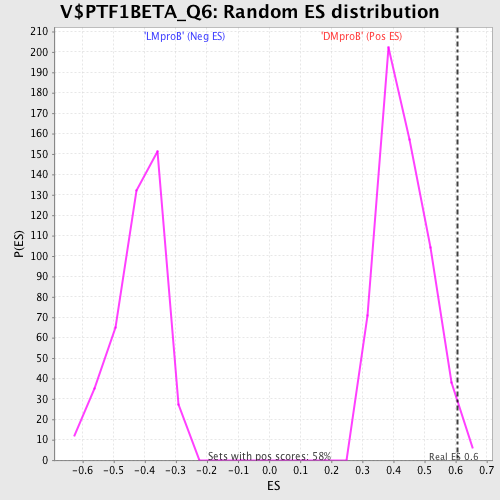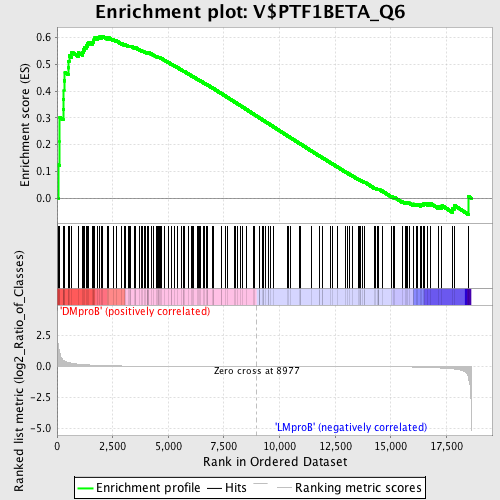
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_DMproB_versus_LMproB.phenotype_DMproB_versus_LMproB.cls #DMproB_versus_LMproB |
| Phenotype | phenotype_DMproB_versus_LMproB.cls#DMproB_versus_LMproB |
| Upregulated in class | DMproB |
| GeneSet | V$PTF1BETA_Q6 |
| Enrichment Score (ES) | 0.6052212 |
| Normalized Enrichment Score (NES) | 1.3964775 |
| Nominal p-value | 0.013840831 |
| FDR q-value | 0.8128674 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | UBTF | 1313 10052 | 49 | 1.608 | 0.1243 | Yes | ||
| 2 | RASSF2 | 2802 14422 | 120 | 1.156 | 0.2117 | Yes | ||
| 3 | EMP3 | 17824 968 | 124 | 1.130 | 0.3007 | Yes | ||
| 4 | KLF13 | 17807 | 272 | 0.499 | 0.3321 | Yes | ||
| 5 | CHD2 | 10847 | 290 | 0.478 | 0.3689 | Yes | ||
| 6 | THRA | 1447 10171 1406 | 308 | 0.456 | 0.4040 | Yes | ||
| 7 | FCHSD2 | 18172 | 311 | 0.453 | 0.4397 | Yes | ||
| 8 | RELB | 17942 | 353 | 0.406 | 0.4695 | Yes | ||
| 9 | ARF3 | 22137 | 497 | 0.314 | 0.4866 | Yes | ||
| 10 | ZFP36L1 | 4453 4454 | 505 | 0.312 | 0.5108 | Yes | ||
| 11 | CAPG | 17411 | 541 | 0.292 | 0.5319 | Yes | ||
| 12 | STAT5A | 20664 | 658 | 0.251 | 0.5455 | Yes | ||
| 13 | H1F0 | 22428 | 960 | 0.182 | 0.5435 | Yes | ||
| 14 | ELA1 | 22127 | 1141 | 0.154 | 0.5459 | Yes | ||
| 15 | CSNK1E | 6570 2211 11332 | 1179 | 0.148 | 0.5556 | Yes | ||
| 16 | NME3 | 23349 | 1230 | 0.142 | 0.5641 | Yes | ||
| 17 | SOX4 | 5479 | 1304 | 0.132 | 0.5705 | Yes | ||
| 18 | ITSN2 | 21327 | 1376 | 0.123 | 0.5764 | Yes | ||
| 19 | EIF4G1 | 22818 | 1424 | 0.118 | 0.5831 | Yes | ||
| 20 | C20ORF185 | 11713 | 1578 | 0.103 | 0.5830 | Yes | ||
| 21 | SLC25A14 | 24344 | 1638 | 0.098 | 0.5875 | Yes | ||
| 22 | TRAF4 | 10217 5796 1400 | 1656 | 0.097 | 0.5942 | Yes | ||
| 23 | DMD | 24295 2647 | 1698 | 0.093 | 0.5994 | Yes | ||
| 24 | ETV6 | 17264 | 1825 | 0.084 | 0.5992 | Yes | ||
| 25 | BAD | 24000 | 1887 | 0.080 | 0.6022 | Yes | ||
| 26 | ITGB7 | 22112 | 1981 | 0.073 | 0.6030 | Yes | ||
| 27 | EED | 17759 | 2041 | 0.069 | 0.6052 | Yes | ||
| 28 | ADCK4 | 13364 | 2275 | 0.057 | 0.5971 | No | ||
| 29 | SRGAP2 | 13836 17045 6433 17046 | 2300 | 0.056 | 0.6002 | No | ||
| 30 | SMOC1 | 21221 | 2516 | 0.047 | 0.5923 | No | ||
| 31 | RASA1 | 10174 | 2654 | 0.041 | 0.5881 | No | ||
| 32 | CRK | 4559 1249 | 2908 | 0.034 | 0.5771 | No | ||
| 33 | BSCL2 | 23940 | 3040 | 0.031 | 0.5724 | No | ||
| 34 | THRAP1 | 20309 | 3080 | 0.030 | 0.5726 | No | ||
| 35 | JMJD1B | 23600 | 3216 | 0.027 | 0.5675 | No | ||
| 36 | CTCF | 18490 | 3252 | 0.026 | 0.5676 | No | ||
| 37 | REPIN1 | 12220 | 3307 | 0.025 | 0.5667 | No | ||
| 38 | LRRTM4 | 17405 | 3316 | 0.025 | 0.5683 | No | ||
| 39 | ACCN3 | 3532 3647 | 3487 | 0.023 | 0.5609 | No | ||
| 40 | CASK | 24184 2618 2604 | 3528 | 0.022 | 0.5604 | No | ||
| 41 | OBSCN | 20434 | 3536 | 0.022 | 0.5617 | No | ||
| 42 | PANK3 | 20924 | 3712 | 0.019 | 0.5538 | No | ||
| 43 | CRHR1 | 1464 1468 20633 | 3809 | 0.018 | 0.5500 | No | ||
| 44 | ANKRD17 | 13496 3474 8170 | 3856 | 0.018 | 0.5489 | No | ||
| 45 | XPO1 | 4172 | 3943 | 0.017 | 0.5456 | No | ||
| 46 | FBXO40 | 22599 444 | 3969 | 0.016 | 0.5455 | No | ||
| 47 | DNMT3A | 2167 21330 | 4045 | 0.016 | 0.5427 | No | ||
| 48 | APBA1 | 11483 | 4076 | 0.015 | 0.5423 | No | ||
| 49 | HOXA11 | 17146 | 4084 | 0.015 | 0.5431 | No | ||
| 50 | LRCH4 | 10544 6090 | 4087 | 0.015 | 0.5442 | No | ||
| 51 | GATA6 | 96 | 4127 | 0.015 | 0.5433 | No | ||
| 52 | BCL2 | 8651 3928 13864 4435 981 4062 13863 4027 | 4227 | 0.014 | 0.5391 | No | ||
| 53 | HOXD12 | 9113 | 4326 | 0.013 | 0.5348 | No | ||
| 54 | TNFRSF1A | 1181 10206 | 4458 | 0.013 | 0.5287 | No | ||
| 55 | KCTD15 | 17864 | 4498 | 0.012 | 0.5276 | No | ||
| 56 | ROBO1 | 9731 | 4561 | 0.012 | 0.5252 | No | ||
| 57 | NTRK1 | 15299 | 4564 | 0.012 | 0.5260 | No | ||
| 58 | TRERF1 | 1546 23213 | 4585 | 0.012 | 0.5259 | No | ||
| 59 | LIF | 956 667 20960 | 4643 | 0.012 | 0.5237 | No | ||
| 60 | LPL | 9283 5003 | 4683 | 0.011 | 0.5225 | No | ||
| 61 | FASTK | 16589 3661 3478 3664 | 4841 | 0.010 | 0.5148 | No | ||
| 62 | ZIC1 | 19040 | 4985 | 0.010 | 0.5078 | No | ||
| 63 | SLITRK3 | 11816 | 5144 | 0.009 | 0.5000 | No | ||
| 64 | STARD13 | 10833 | 5163 | 0.009 | 0.4997 | No | ||
| 65 | EEF1A2 | 8880 14309 | 5297 | 0.009 | 0.4932 | No | ||
| 66 | KCNJ1 | 3121 12110 | 5417 | 0.008 | 0.4874 | No | ||
| 67 | TBL1X | 5628 | 5604 | 0.007 | 0.4779 | No | ||
| 68 | SMAD6 | 19083 | 5691 | 0.007 | 0.4738 | No | ||
| 69 | FGF14 | 21716 3005 3463 | 5695 | 0.007 | 0.4742 | No | ||
| 70 | NXPH3 | 20281 | 5716 | 0.007 | 0.4737 | No | ||
| 71 | HMGB3 | 9096 | 5916 | 0.006 | 0.4634 | No | ||
| 72 | FBXO24 | 12918 3649 | 6055 | 0.006 | 0.4564 | No | ||
| 73 | PAX3 | 13910 | 6088 | 0.006 | 0.4551 | No | ||
| 74 | PTPN2 | 23414 | 6127 | 0.006 | 0.4535 | No | ||
| 75 | BBOX1 | 5010 9290 | 6321 | 0.005 | 0.4435 | No | ||
| 76 | DUSP22 | 466 21680 | 6359 | 0.005 | 0.4419 | No | ||
| 77 | STC1 | 21974 | 6380 | 0.005 | 0.4412 | No | ||
| 78 | RIMS1 | 8575 | 6443 | 0.005 | 0.4383 | No | ||
| 79 | SYT7 | 3670 3698 23932 | 6597 | 0.005 | 0.4303 | No | ||
| 80 | CCKBR | 8706 | 6640 | 0.004 | 0.4284 | No | ||
| 81 | CDH13 | 4506 3826 | 6712 | 0.004 | 0.4249 | No | ||
| 82 | NPPA | 298 | 6741 | 0.004 | 0.4237 | No | ||
| 83 | REST | 16818 | 6743 | 0.004 | 0.4240 | No | ||
| 84 | PDE10A | 23387 | 6970 | 0.004 | 0.4121 | No | ||
| 85 | GABRA1 | 20492 | 7028 | 0.004 | 0.4092 | No | ||
| 86 | GNG3 | 23752 | 7029 | 0.004 | 0.4095 | No | ||
| 87 | PIAS2 | 1948 5102 | 7371 | 0.003 | 0.3913 | No | ||
| 88 | BCL2L1 | 4440 2930 8652 | 7409 | 0.003 | 0.3895 | No | ||
| 89 | PCDH7 | 7029 3460 | 7582 | 0.003 | 0.3804 | No | ||
| 90 | TEAD1 | 18121 | 7671 | 0.002 | 0.3758 | No | ||
| 91 | SLC6A14 | 12168 | 7954 | 0.002 | 0.3607 | No | ||
| 92 | ITGA7 | 19835 | 8031 | 0.002 | 0.3567 | No | ||
| 93 | ERBB3 | 19593 | 8085 | 0.002 | 0.3539 | No | ||
| 94 | PLEKHC1 | 21864 | 8226 | 0.001 | 0.3465 | No | ||
| 95 | NR4A3 | 9473 16212 5183 | 8351 | 0.001 | 0.3398 | No | ||
| 96 | HES7 | 20828 | 8525 | 0.001 | 0.3305 | No | ||
| 97 | CD163 | 17289 | 8808 | 0.000 | 0.3152 | No | ||
| 98 | PLXNC1 | 7056 | 8873 | 0.000 | 0.3118 | No | ||
| 99 | VAX1 | 23636 | 9088 | -0.000 | 0.3002 | No | ||
| 100 | PAQR4 | 23105 | 9093 | -0.000 | 0.3000 | No | ||
| 101 | PTBP2 | 7074 | 9116 | -0.000 | 0.2988 | No | ||
| 102 | MATN1 | 16060 | 9251 | -0.001 | 0.2916 | No | ||
| 103 | MDS1 | 9375 | 9267 | -0.001 | 0.2908 | No | ||
| 104 | TREX2 | 24137 | 9295 | -0.001 | 0.2894 | No | ||
| 105 | CALD1 | 4273 8463 | 9344 | -0.001 | 0.2869 | No | ||
| 106 | KCNE1L | 24044 | 9501 | -0.001 | 0.2785 | No | ||
| 107 | GNAT1 | 3027 19000 | 9511 | -0.001 | 0.2781 | No | ||
| 108 | LRRTM1 | 17408 | 9601 | -0.001 | 0.2733 | No | ||
| 109 | SLITRK5 | 21939 | 9710 | -0.001 | 0.2676 | No | ||
| 110 | PPP2R2A | 3222 21774 | 10346 | -0.002 | 0.2333 | No | ||
| 111 | TNIP2 | 16561 | 10395 | -0.002 | 0.2309 | No | ||
| 112 | PHYHIP | 21968 | 10484 | -0.002 | 0.2264 | No | ||
| 113 | PMP22 | 9594 1312 | 10914 | -0.003 | 0.2034 | No | ||
| 114 | WISP1 | 22463 2289 | 10921 | -0.003 | 0.2033 | No | ||
| 115 | NPDC1 | 9479 | 10955 | -0.003 | 0.2018 | No | ||
| 116 | ROR1 | 6472 | 11431 | -0.004 | 0.1764 | No | ||
| 117 | HHIP | 4849 | 11771 | -0.005 | 0.1584 | No | ||
| 118 | USP5 | 17004 | 11803 | -0.005 | 0.1572 | No | ||
| 119 | HOXB6 | 20688 | 11908 | -0.005 | 0.1520 | No | ||
| 120 | TMPRSS2 | 22527 | 11948 | -0.006 | 0.1503 | No | ||
| 121 | ANK3 | 8591 3445 3304 3381 | 12275 | -0.006 | 0.1331 | No | ||
| 122 | CD6 | 3669 23740 | 12359 | -0.007 | 0.1291 | No | ||
| 123 | DHRS3 | 15997 | 12584 | -0.007 | 0.1176 | No | ||
| 124 | CCNC | 16263 | 12970 | -0.008 | 0.0974 | No | ||
| 125 | LIN28 | 15723 | 13032 | -0.009 | 0.0947 | No | ||
| 126 | SNCAIP | 12589 | 13123 | -0.009 | 0.0906 | No | ||
| 127 | SLC8A3 | 2065 4293 | 13292 | -0.010 | 0.0822 | No | ||
| 128 | NR4A2 | 5203 | 13528 | -0.011 | 0.0704 | No | ||
| 129 | CITED1 | 24091 | 13580 | -0.011 | 0.0685 | No | ||
| 130 | SYT3 | 18264 | 13615 | -0.011 | 0.0675 | No | ||
| 131 | KIRREL2 | 17889 3922 | 13623 | -0.011 | 0.0680 | No | ||
| 132 | ST18 | 14298 | 13719 | -0.012 | 0.0638 | No | ||
| 133 | NCALD | 22310 | 13816 | -0.013 | 0.0596 | No | ||
| 134 | PURA | 9670 | 13829 | -0.013 | 0.0599 | No | ||
| 135 | IGFBP6 | 9161 | 14287 | -0.016 | 0.0364 | No | ||
| 136 | FOXP2 | 4312 17523 8523 | 14306 | -0.016 | 0.0367 | No | ||
| 137 | ZIC3 | 10430 | 14378 | -0.016 | 0.0341 | No | ||
| 138 | SLC25A13 | 17221 1159 | 14428 | -0.017 | 0.0328 | No | ||
| 139 | GPC6 | 21937 | 14445 | -0.017 | 0.0332 | No | ||
| 140 | AR | 24286 | 14451 | -0.017 | 0.0343 | No | ||
| 141 | ZFYVE20 | 17065 | 14610 | -0.018 | 0.0272 | No | ||
| 142 | PUM1 | 8160 | 15033 | -0.023 | 0.0062 | No | ||
| 143 | YWHAE | 20776 | 15136 | -0.025 | 0.0026 | No | ||
| 144 | XPR1 | 5383 | 15180 | -0.026 | 0.0023 | No | ||
| 145 | LRRFIP2 | 3024 3071 19285 | 15515 | -0.032 | -0.0132 | No | ||
| 146 | MTMR4 | 20712 | 15646 | -0.036 | -0.0175 | No | ||
| 147 | MAN2C1 | 19433 | 15696 | -0.037 | -0.0172 | No | ||
| 148 | HOXB4 | 20686 | 15740 | -0.038 | -0.0166 | No | ||
| 149 | HEBP2 | 19812 | 15821 | -0.041 | -0.0177 | No | ||
| 150 | AMMECR1 | 7067 | 15996 | -0.047 | -0.0234 | No | ||
| 151 | TGFB3 | 10161 | 16032 | -0.048 | -0.0215 | No | ||
| 152 | ATP1B3 | 19037 | 16138 | -0.053 | -0.0231 | No | ||
| 153 | ITPKC | 17919 | 16207 | -0.056 | -0.0224 | No | ||
| 154 | ARMCX3 | 12951 | 16345 | -0.062 | -0.0249 | No | ||
| 155 | TBCC | 23215 | 16396 | -0.064 | -0.0225 | No | ||
| 156 | PPARGC1A | 16533 | 16471 | -0.068 | -0.0212 | No | ||
| 157 | IMPDH1 | 17197 1131 | 16527 | -0.070 | -0.0186 | No | ||
| 158 | CDK6 | 16600 | 16668 | -0.079 | -0.0200 | No | ||
| 159 | STOML2 | 15902 | 16775 | -0.088 | -0.0188 | No | ||
| 160 | DICER1 | 20989 | 17140 | -0.116 | -0.0293 | No | ||
| 161 | NMT1 | 20638 | 17299 | -0.134 | -0.0273 | No | ||
| 162 | GSPT1 | 9046 | 17765 | -0.189 | -0.0376 | No | ||
| 163 | ACADVL | 20375 | 17844 | -0.201 | -0.0260 | No | ||
| 164 | MAGED1 | 24108 | 18486 | -0.858 | 0.0070 | No |