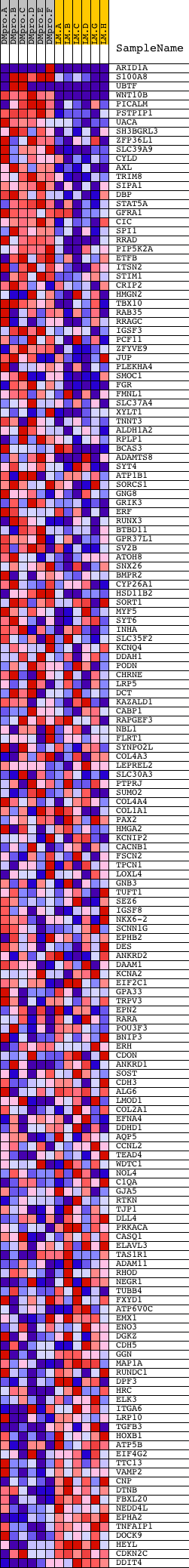
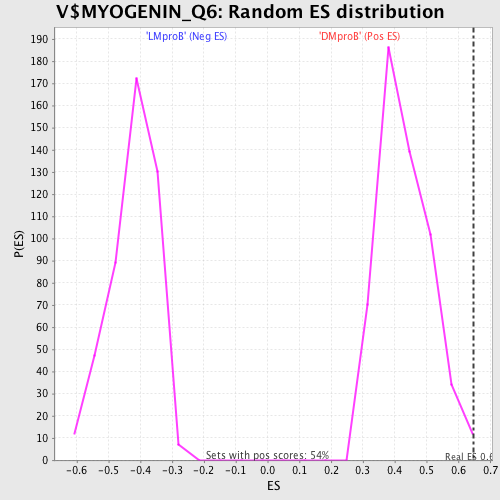
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_DMproB_versus_LMproB.phenotype_DMproB_versus_LMproB.cls #DMproB_versus_LMproB |
| Phenotype | phenotype_DMproB_versus_LMproB.cls#DMproB_versus_LMproB |
| Upregulated in class | DMproB |
| GeneSet | V$MYOGENIN_Q6 |
| Enrichment Score (ES) | 0.64733875 |
| Normalized Enrichment Score (NES) | 1.504095 |
| Nominal p-value | 0.005524862 |
| FDR q-value | 1.0 |
| FWER p-Value | 0.982 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ARID1A | 8214 2501 13554 | 3 | 4.234 | 0.2145 | Yes | ||
| 2 | S100A8 | 15527 | 40 | 1.871 | 0.3074 | Yes | ||
| 3 | UBTF | 1313 10052 | 49 | 1.608 | 0.3885 | Yes | ||
| 4 | WNT10B | 5874 | 78 | 1.365 | 0.4562 | Yes | ||
| 5 | PICALM | 18191 | 270 | 0.504 | 0.4714 | Yes | ||
| 6 | PSTPIP1 | 19437 | 288 | 0.478 | 0.4947 | Yes | ||
| 7 | UACA | 19417 | 429 | 0.353 | 0.5051 | Yes | ||
| 8 | SH3BGRL3 | 15722 | 481 | 0.322 | 0.5186 | Yes | ||
| 9 | ZFP36L1 | 4453 4454 | 505 | 0.312 | 0.5332 | Yes | ||
| 10 | SLC39A9 | 11597 | 536 | 0.294 | 0.5465 | Yes | ||
| 11 | CYLD | 18532 | 576 | 0.278 | 0.5585 | Yes | ||
| 12 | AXL | 17922 | 579 | 0.278 | 0.5724 | Yes | ||
| 13 | TRIM8 | 23824 | 603 | 0.271 | 0.5849 | Yes | ||
| 14 | SIPA1 | 9820 | 604 | 0.270 | 0.5986 | Yes | ||
| 15 | DBP | 18242 | 608 | 0.269 | 0.6121 | Yes | ||
| 16 | STAT5A | 20664 | 658 | 0.251 | 0.6222 | Yes | ||
| 17 | GFRA1 | 9015 | 850 | 0.202 | 0.6221 | Yes | ||
| 18 | CIC | 18338 | 888 | 0.194 | 0.6299 | Yes | ||
| 19 | SPI1 | 14949 | 999 | 0.174 | 0.6328 | Yes | ||
| 20 | RRAD | 18775 | 1113 | 0.157 | 0.6346 | Yes | ||
| 21 | PIP5K2A | 14676 | 1216 | 0.143 | 0.6363 | Yes | ||
| 22 | ETFB | 18279 | 1325 | 0.129 | 0.6370 | Yes | ||
| 23 | ITSN2 | 21327 | 1376 | 0.123 | 0.6406 | Yes | ||
| 24 | STIM1 | 18164 | 1434 | 0.118 | 0.6434 | Yes | ||
| 25 | CRIP2 | 12681 | 1544 | 0.106 | 0.6429 | Yes | ||
| 26 | HMGN2 | 9095 | 1595 | 0.102 | 0.6454 | Yes | ||
| 27 | TBX10 | 4272 5631 | 1706 | 0.092 | 0.6441 | Yes | ||
| 28 | RAB35 | 13386 | 1732 | 0.090 | 0.6473 | Yes | ||
| 29 | RRAGC | 12015 | 1883 | 0.080 | 0.6433 | No | ||
| 30 | IGSF3 | 8128 | 1964 | 0.075 | 0.6428 | No | ||
| 31 | PCF11 | 121 | 2122 | 0.064 | 0.6375 | No | ||
| 32 | ZFYVE9 | 10501 2449 | 2151 | 0.063 | 0.6392 | No | ||
| 33 | JUP | 4937 | 2467 | 0.048 | 0.6246 | No | ||
| 34 | PLEKHA4 | 18246 | 2514 | 0.047 | 0.6244 | No | ||
| 35 | SMOC1 | 21221 | 2516 | 0.047 | 0.6267 | No | ||
| 36 | FGR | 4723 | 2604 | 0.043 | 0.6242 | No | ||
| 37 | FMNL1 | 12191 | 2637 | 0.042 | 0.6246 | No | ||
| 38 | SLC37A4 | 8994 | 2718 | 0.039 | 0.6223 | No | ||
| 39 | XYLT1 | 18113 | 2740 | 0.038 | 0.6230 | No | ||
| 40 | TNNT3 | 18004 | 2804 | 0.036 | 0.6215 | No | ||
| 41 | ALDH1A2 | 19386 | 2986 | 0.032 | 0.6133 | No | ||
| 42 | RPLP1 | 3010 | 3012 | 0.031 | 0.6135 | No | ||
| 43 | BCAS3 | 5314 | 3083 | 0.030 | 0.6112 | No | ||
| 44 | ADAMTS8 | 6626 | 3181 | 0.028 | 0.6074 | No | ||
| 45 | SYT4 | 5566 | 3186 | 0.028 | 0.6086 | No | ||
| 46 | ATP1B1 | 4420 | 3325 | 0.025 | 0.6024 | No | ||
| 47 | SORCS1 | 7184 12203 | 3365 | 0.024 | 0.6015 | No | ||
| 48 | GNG8 | 18378 | 3371 | 0.024 | 0.6024 | No | ||
| 49 | GRIK3 | 16083 | 3390 | 0.024 | 0.6027 | No | ||
| 50 | ERF | 4680 8914 | 3400 | 0.024 | 0.6034 | No | ||
| 51 | RUNX3 | 8702 | 4310 | 0.014 | 0.5548 | No | ||
| 52 | BTBD11 | 13147 | 4324 | 0.013 | 0.5548 | No | ||
| 53 | GPR37L1 | 13825 | 4443 | 0.013 | 0.5491 | No | ||
| 54 | SV2B | 17795 | 4488 | 0.012 | 0.5473 | No | ||
| 55 | ATOH8 | 17120 | 4659 | 0.011 | 0.5387 | No | ||
| 56 | SNX26 | 10582 | 4759 | 0.011 | 0.5339 | No | ||
| 57 | BMPR2 | 14241 4081 | 4816 | 0.011 | 0.5314 | No | ||
| 58 | CYP26A1 | 23871 3695 | 4866 | 0.010 | 0.5292 | No | ||
| 59 | HSD11B2 | 4872 | 4998 | 0.010 | 0.5226 | No | ||
| 60 | SORT1 | 15452 5475 9847 | 5071 | 0.010 | 0.5192 | No | ||
| 61 | MYF5 | 19632 | 5625 | 0.007 | 0.4896 | No | ||
| 62 | SYT6 | 13437 7042 12040 | 5859 | 0.007 | 0.4773 | No | ||
| 63 | INHA | 14214 | 5901 | 0.007 | 0.4754 | No | ||
| 64 | SLC35F2 | 19451 | 6084 | 0.006 | 0.4659 | No | ||
| 65 | KCNQ4 | 12247 2533 | 6257 | 0.005 | 0.4568 | No | ||
| 66 | DDAH1 | 7610 | 6302 | 0.005 | 0.4547 | No | ||
| 67 | PODN | 15811 | 6445 | 0.005 | 0.4473 | No | ||
| 68 | CHRNE | 20368 | 6487 | 0.005 | 0.4453 | No | ||
| 69 | LRP5 | 23948 9285 | 6508 | 0.005 | 0.4444 | No | ||
| 70 | DCT | 4603 21723 | 6567 | 0.005 | 0.4415 | No | ||
| 71 | KAZALD1 | 4206 | 6599 | 0.005 | 0.4401 | No | ||
| 72 | CABP1 | 16412 | 6896 | 0.004 | 0.4242 | No | ||
| 73 | RAPGEF3 | 22146 | 6917 | 0.004 | 0.4234 | No | ||
| 74 | NBL1 | 15699 | 6939 | 0.004 | 0.4224 | No | ||
| 75 | FLRT1 | 11852 | 6987 | 0.004 | 0.4201 | No | ||
| 76 | SYNPO2L | 7582 | 7011 | 0.004 | 0.4190 | No | ||
| 77 | COL4A3 | 8766 | 7015 | 0.004 | 0.4190 | No | ||
| 78 | LEPREL2 | 17001 | 7017 | 0.004 | 0.4192 | No | ||
| 79 | SLC30A3 | 5995 | 7215 | 0.003 | 0.4086 | No | ||
| 80 | PTPRJ | 9664 | 7224 | 0.003 | 0.4084 | No | ||
| 81 | SUMO2 | 1293 9319 | 7414 | 0.003 | 0.3983 | No | ||
| 82 | COL4A4 | 8767 | 7442 | 0.003 | 0.3970 | No | ||
| 83 | COL1A1 | 8771 | 7716 | 0.002 | 0.3823 | No | ||
| 84 | PAX2 | 9530 24416 | 7939 | 0.002 | 0.3703 | No | ||
| 85 | HMGA2 | 4857 9098 3341 3444 | 8119 | 0.002 | 0.3607 | No | ||
| 86 | KCNIP2 | 23655 3725 | 8206 | 0.001 | 0.3561 | No | ||
| 87 | CACNB1 | 1443 1304 | 8222 | 0.001 | 0.3554 | No | ||
| 88 | FSCN2 | 20569 | 8232 | 0.001 | 0.3550 | No | ||
| 89 | TPCN1 | 16398 6383 10915 | 8277 | 0.001 | 0.3526 | No | ||
| 90 | LOXL4 | 23672 | 8394 | 0.001 | 0.3464 | No | ||
| 91 | GNB3 | 9027 | 8397 | 0.001 | 0.3463 | No | ||
| 92 | TUFT1 | 1847 15253 | 8578 | 0.001 | 0.3366 | No | ||
| 93 | SEZ6 | 20764 | 8728 | 0.000 | 0.3286 | No | ||
| 94 | IGSF8 | 14048 | 8756 | 0.000 | 0.3271 | No | ||
| 95 | NKX6-2 | 17579 4815 | 8805 | 0.000 | 0.3245 | No | ||
| 96 | SCNN1G | 18095 | 8911 | 0.000 | 0.3188 | No | ||
| 97 | EPHB2 | 4675 2440 8910 | 8933 | 0.000 | 0.3177 | No | ||
| 98 | DES | 14215 | 8940 | 0.000 | 0.3174 | No | ||
| 99 | ANKRD2 | 23853 | 8947 | 0.000 | 0.3171 | No | ||
| 100 | DAAM1 | 5536 | 9000 | -0.000 | 0.3142 | No | ||
| 101 | KCNA2 | 15458 | 9001 | -0.000 | 0.3142 | No | ||
| 102 | EIF2C1 | 10672 | 9174 | -0.000 | 0.3049 | No | ||
| 103 | GPA33 | 14066 | 9278 | -0.001 | 0.2994 | No | ||
| 104 | TRPV3 | 20788 | 9360 | -0.001 | 0.2950 | No | ||
| 105 | EPN2 | 20413 945 946 955 | 9806 | -0.001 | 0.2710 | No | ||
| 106 | RARA | 5358 | 10355 | -0.002 | 0.2414 | No | ||
| 107 | POU3F3 | 14261 | 10553 | -0.003 | 0.2309 | No | ||
| 108 | BNIP3 | 17581 | 10612 | -0.003 | 0.2279 | No | ||
| 109 | ERH | 9614 4681 | 10680 | -0.003 | 0.2244 | No | ||
| 110 | CDON | 12195 | 10738 | -0.003 | 0.2214 | No | ||
| 111 | ANKRD1 | 23689 | 11035 | -0.004 | 0.2056 | No | ||
| 112 | SOST | 20210 | 11507 | -0.005 | 0.1803 | No | ||
| 113 | CDH3 | 3919 4509 | 11519 | -0.005 | 0.1799 | No | ||
| 114 | ALG6 | 6757 | 11558 | -0.005 | 0.1781 | No | ||
| 115 | LMOD1 | 14119 | 11770 | -0.005 | 0.1669 | No | ||
| 116 | COL2A1 | 22145 4542 | 11992 | -0.006 | 0.1552 | No | ||
| 117 | EFNA4 | 1866 15277 | 12001 | -0.006 | 0.1551 | No | ||
| 118 | DDHD1 | 497 496 4339 | 12201 | -0.006 | 0.1446 | No | ||
| 119 | AQP5 | 22365 | 12228 | -0.006 | 0.1435 | No | ||
| 120 | CCNL2 | 15957 | 12235 | -0.006 | 0.1435 | No | ||
| 121 | TEAD4 | 997 5708 | 12677 | -0.007 | 0.1200 | No | ||
| 122 | WDTC1 | 10512 | 12681 | -0.008 | 0.1202 | No | ||
| 123 | NOL4 | 2028 11432 | 12895 | -0.008 | 0.1091 | No | ||
| 124 | C1QA | 8666 | 13039 | -0.009 | 0.1018 | No | ||
| 125 | GJA5 | 1929 9017 4779 | 13233 | -0.009 | 0.0918 | No | ||
| 126 | RTKN | 17394 1156 | 13317 | -0.010 | 0.0878 | No | ||
| 127 | TJP1 | 17803 | 13375 | -0.010 | 0.0852 | No | ||
| 128 | DLL4 | 7040 | 13405 | -0.010 | 0.0841 | No | ||
| 129 | PRKACA | 18549 3844 | 13408 | -0.010 | 0.0845 | No | ||
| 130 | CASQ1 | 13752 8695 | 13418 | -0.010 | 0.0846 | No | ||
| 131 | ELAVL3 | 9136 | 13534 | -0.011 | 0.0789 | No | ||
| 132 | TAS1R1 | 15657 | 13654 | -0.011 | 0.0730 | No | ||
| 133 | ADAM11 | 4340 1432 | 13662 | -0.012 | 0.0732 | No | ||
| 134 | RHOD | 23961 | 13939 | -0.013 | 0.0589 | No | ||
| 135 | NEGR1 | 6806 11572 | 13952 | -0.013 | 0.0590 | No | ||
| 136 | TUBB4 | 22917 | 13966 | -0.013 | 0.0590 | No | ||
| 137 | FXYD1 | 17874 85 | 14006 | -0.014 | 0.0575 | No | ||
| 138 | ATP6V0C | 8643 | 14037 | -0.014 | 0.0566 | No | ||
| 139 | EMX1 | 8900 | 14192 | -0.015 | 0.0490 | No | ||
| 140 | ENO3 | 8905 | 14203 | -0.015 | 0.0492 | No | ||
| 141 | DGKZ | 2836 14522 | 14263 | -0.015 | 0.0468 | No | ||
| 142 | CDH5 | 18512 | 14407 | -0.017 | 0.0399 | No | ||
| 143 | GGN | 2472 1171 18312 | 14918 | -0.022 | 0.0134 | No | ||
| 144 | MAP1A | 5124 | 14943 | -0.022 | 0.0132 | No | ||
| 145 | RUNDC1 | 20655 | 15238 | -0.027 | -0.0014 | No | ||
| 146 | DPF3 | 7659 12853 | 15325 | -0.028 | -0.0046 | No | ||
| 147 | HRC | 18251 | 15498 | -0.032 | -0.0123 | No | ||
| 148 | ELK3 | 4663 94 3388 | 15500 | -0.032 | -0.0108 | No | ||
| 149 | ITGA6 | 4930 | 15653 | -0.036 | -0.0172 | No | ||
| 150 | LRP10 | 22018 | 16016 | -0.048 | -0.0344 | No | ||
| 151 | TGFB3 | 10161 | 16032 | -0.048 | -0.0328 | No | ||
| 152 | HOXB1 | 20684 | 16131 | -0.052 | -0.0354 | No | ||
| 153 | ATP5B | 19846 | 16547 | -0.071 | -0.0543 | No | ||
| 154 | EIF4G2 | 1908 8892 | 16656 | -0.078 | -0.0562 | No | ||
| 155 | TTC13 | 10636 | 16738 | -0.085 | -0.0562 | No | ||
| 156 | VAMP2 | 20826 | 17025 | -0.106 | -0.0663 | No | ||
| 157 | CNP | 8760 | 17479 | -0.151 | -0.0832 | No | ||
| 158 | DTNB | 21333 | 17509 | -0.155 | -0.0770 | No | ||
| 159 | FBXL20 | 13010 | 17887 | -0.212 | -0.0866 | No | ||
| 160 | NEDD4L | 8192 | 17903 | -0.215 | -0.0766 | No | ||
| 161 | EPHA2 | 16006 | 18060 | -0.262 | -0.0717 | No | ||
| 162 | TNFAIP1 | 20333 | 18066 | -0.263 | -0.0586 | No | ||
| 163 | DOCK9 | 8369 | 18170 | -0.299 | -0.0491 | No | ||
| 164 | HEYL | 16095 | 18276 | -0.373 | -0.0358 | No | ||
| 165 | CDKN2C | 15806 2321 | 18342 | -0.432 | -0.0175 | No | ||
| 166 | DDIT4 | 19762 | 18442 | -0.636 | 0.0094 | No |