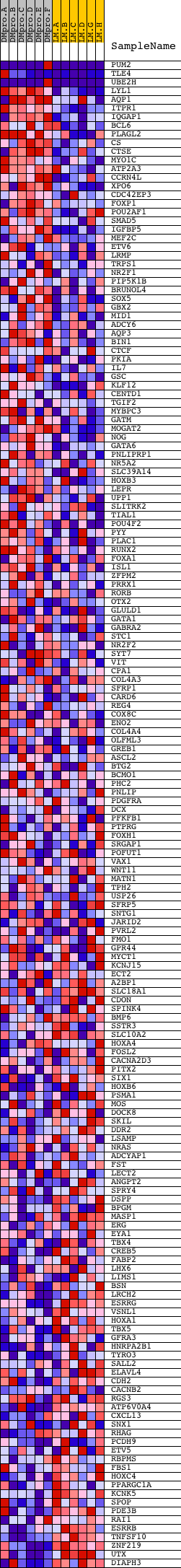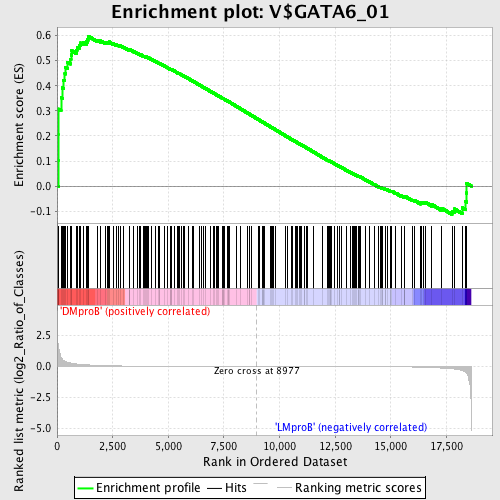
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_DMproB_versus_LMproB.phenotype_DMproB_versus_LMproB.cls #DMproB_versus_LMproB |
| Phenotype | phenotype_DMproB_versus_LMproB.cls#DMproB_versus_LMproB |
| Upregulated in class | DMproB |
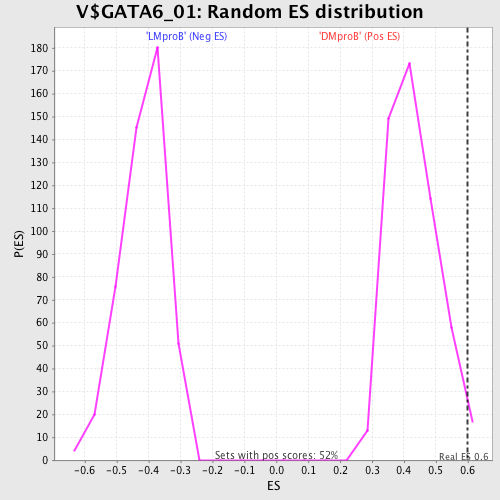
| GeneSet | V$GATA6_01 |
| Enrichment Score (ES) | 0.59772533 |
| Normalized Enrichment Score (NES) | 1.3923244 |
| Nominal p-value | 0.026717557 |
| FDR q-value | 0.7649079 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PUM2 | 2071 8161 | 58 | 1.485 | 0.1028 | Yes | ||
| 2 | TLE4 | 23720 3697 | 64 | 1.444 | 0.2054 | Yes | ||
| 3 | UBE2H | 5823 5822 | 68 | 1.432 | 0.3074 | Yes | ||
| 4 | LYL1 | 18546 | 177 | 0.727 | 0.3533 | Yes | ||
| 5 | AQP1 | 1057 17437 | 231 | 0.576 | 0.3915 | Yes | ||
| 6 | ITPR1 | 17341 | 293 | 0.475 | 0.4221 | Yes | ||
| 7 | IQGAP1 | 6619 | 350 | 0.409 | 0.4482 | Yes | ||
| 8 | BCL6 | 22624 | 380 | 0.381 | 0.4738 | Yes | ||
| 9 | PLAGL2 | 7055 | 461 | 0.334 | 0.4933 | Yes | ||
| 10 | CS | 19839 | 588 | 0.276 | 0.5061 | Yes | ||
| 11 | CTSE | 8816 | 626 | 0.263 | 0.5229 | Yes | ||
| 12 | MYO1C | 20777 | 651 | 0.254 | 0.5397 | Yes | ||
| 13 | ATP2A3 | 20795 1314 | 875 | 0.197 | 0.5416 | Yes | ||
| 14 | CCRN4L | 15599 8708 | 896 | 0.193 | 0.5543 | Yes | ||
| 15 | XPO6 | 17644 | 994 | 0.175 | 0.5615 | Yes | ||
| 16 | CDC42EP3 | 6436 | 1048 | 0.167 | 0.5705 | Yes | ||
| 17 | FOXP1 | 4242 | 1181 | 0.148 | 0.5739 | Yes | ||
| 18 | POU2AF1 | 19455 | 1332 | 0.129 | 0.5749 | Yes | ||
| 19 | SMAD5 | 21621 | 1352 | 0.125 | 0.5828 | Yes | ||
| 20 | IGFBP5 | 4899 | 1391 | 0.121 | 0.5894 | Yes | ||
| 21 | MEF2C | 3204 9378 | 1398 | 0.121 | 0.5977 | Yes | ||
| 22 | ETV6 | 17264 | 1825 | 0.084 | 0.5806 | No | ||
| 23 | LRMP | 17244 | 1960 | 0.075 | 0.5787 | No | ||
| 24 | TRPS1 | 8195 | 2172 | 0.062 | 0.5717 | No | ||
| 25 | NR2F1 | 7015 21401 | 2269 | 0.057 | 0.5706 | No | ||
| 26 | PIP5K1B | 5251 1841 | 2307 | 0.056 | 0.5725 | No | ||
| 27 | BRUNOL4 | 2010 4229 | 2337 | 0.054 | 0.5748 | No | ||
| 28 | SOX5 | 9850 1044 5480 16937 | 2527 | 0.046 | 0.5678 | No | ||
| 29 | GBX2 | 13887 | 2661 | 0.041 | 0.5636 | No | ||
| 30 | MID1 | 5097 5098 | 2759 | 0.038 | 0.5610 | No | ||
| 31 | ADCY6 | 22139 2283 8551 | 2830 | 0.035 | 0.5597 | No | ||
| 32 | AQP3 | 15915 | 2990 | 0.032 | 0.5534 | No | ||
| 33 | BIN1 | 6634 2017 | 3240 | 0.027 | 0.5418 | No | ||
| 34 | CTCF | 18490 | 3252 | 0.026 | 0.5431 | No | ||
| 35 | PKIA | 15639 | 3272 | 0.026 | 0.5439 | No | ||
| 36 | IL7 | 4921 | 3448 | 0.023 | 0.5360 | No | ||
| 37 | GSC | 20990 | 3596 | 0.021 | 0.5295 | No | ||
| 38 | KLF12 | 4960 2637 21732 9228 | 3713 | 0.019 | 0.5246 | No | ||
| 39 | CENTD1 | 16529 | 3769 | 0.019 | 0.5230 | No | ||
| 40 | TGIF2 | 6013 6012 | 3881 | 0.017 | 0.5182 | No | ||
| 41 | MYBPC3 | 9434 | 3949 | 0.017 | 0.5158 | No | ||
| 42 | GATM | 14451 | 3989 | 0.016 | 0.5148 | No | ||
| 43 | MOGAT2 | 17747 | 4005 | 0.016 | 0.5151 | No | ||
| 44 | NOG | 13663 | 4054 | 0.016 | 0.5136 | No | ||
| 45 | GATA6 | 96 | 4127 | 0.015 | 0.5108 | No | ||
| 46 | PNLIPRP1 | 23835 | 4242 | 0.014 | 0.5056 | No | ||
| 47 | NR5A2 | 4105 13817 | 4432 | 0.013 | 0.4963 | No | ||
| 48 | SLC39A14 | 3517 5611 21762 | 4569 | 0.012 | 0.4898 | No | ||
| 49 | HOXB3 | 20685 | 4603 | 0.012 | 0.4888 | No | ||
| 50 | LEPR | 2531 2349 | 4615 | 0.012 | 0.4891 | No | ||
| 51 | UPP1 | 20947 1385 | 4812 | 0.011 | 0.4792 | No | ||
| 52 | SLITRK2 | 24322 | 4950 | 0.010 | 0.4725 | No | ||
| 53 | TIAL1 | 996 5753 | 5079 | 0.009 | 0.4662 | No | ||
| 54 | POU4F2 | 9603 5276 | 5100 | 0.009 | 0.4658 | No | ||
| 55 | PYY | 20207 | 5130 | 0.009 | 0.4649 | No | ||
| 56 | PLAC1 | 12079 | 5133 | 0.009 | 0.4654 | No | ||
| 57 | RUNX2 | 4480 8700 | 5256 | 0.009 | 0.4594 | No | ||
| 58 | FOXA1 | 21060 | 5421 | 0.008 | 0.4511 | No | ||
| 59 | ISL1 | 21338 | 5462 | 0.008 | 0.4495 | No | ||
| 60 | ZFPM2 | 22483 | 5490 | 0.008 | 0.4486 | No | ||
| 61 | PRRX1 | 9595 5271 | 5585 | 0.008 | 0.4441 | No | ||
| 62 | RORB | 10356 | 5677 | 0.007 | 0.4396 | No | ||
| 63 | OTX2 | 9519 | 5727 | 0.007 | 0.4375 | No | ||
| 64 | GLULD1 | 14285 | 5909 | 0.006 | 0.4281 | No | ||
| 65 | GATA1 | 24196 | 6065 | 0.006 | 0.4202 | No | ||
| 66 | GABRA2 | 4746 | 6149 | 0.006 | 0.4161 | No | ||
| 67 | STC1 | 21974 | 6380 | 0.005 | 0.4040 | No | ||
| 68 | NR2F2 | 8619 409 | 6497 | 0.005 | 0.3980 | No | ||
| 69 | SYT7 | 3670 3698 23932 | 6597 | 0.005 | 0.3930 | No | ||
| 70 | VIT | 23155 | 6673 | 0.004 | 0.3892 | No | ||
| 71 | CPA1 | 17496 | 6915 | 0.004 | 0.3764 | No | ||
| 72 | COL4A3 | 8766 | 7015 | 0.004 | 0.3713 | No | ||
| 73 | SFRP1 | 9805 | 7061 | 0.004 | 0.3691 | No | ||
| 74 | CARD6 | 22334 | 7161 | 0.003 | 0.3640 | No | ||
| 75 | REG4 | 15484 | 7186 | 0.003 | 0.3630 | No | ||
| 76 | COX8C | 21175 | 7264 | 0.003 | 0.3590 | No | ||
| 77 | ENO2 | 8904 | 7436 | 0.003 | 0.3499 | No | ||
| 78 | COL4A4 | 8767 | 7442 | 0.003 | 0.3499 | No | ||
| 79 | OLFML3 | 15216 | 7477 | 0.003 | 0.3482 | No | ||
| 80 | GREB1 | 6490 2152 | 7486 | 0.003 | 0.3480 | No | ||
| 81 | ASCL2 | 5079 | 7533 | 0.003 | 0.3457 | No | ||
| 82 | BTG2 | 13832 | 7656 | 0.002 | 0.3392 | No | ||
| 83 | BCMO1 | 18454 | 7686 | 0.002 | 0.3378 | No | ||
| 84 | PHC2 | 16075 | 7711 | 0.002 | 0.3367 | No | ||
| 85 | PNLIP | 23830 | 7732 | 0.002 | 0.3358 | No | ||
| 86 | PDGFRA | 16824 | 8042 | 0.002 | 0.3191 | No | ||
| 87 | DCX | 24040 8841 4604 | 8248 | 0.001 | 0.3081 | No | ||
| 88 | PFKFB1 | 9552 | 8574 | 0.001 | 0.2905 | No | ||
| 89 | PTPRG | 22097 3151 | 8637 | 0.001 | 0.2872 | No | ||
| 90 | FOXH1 | 22238 | 8648 | 0.001 | 0.2867 | No | ||
| 91 | SRGAP1 | 19615 | 8733 | 0.000 | 0.2822 | No | ||
| 92 | POFUT1 | 4697 | 9033 | -0.000 | 0.2660 | No | ||
| 93 | VAX1 | 23636 | 9088 | -0.000 | 0.2631 | No | ||
| 94 | WNT11 | 5876 | 9243 | -0.000 | 0.2548 | No | ||
| 95 | MATN1 | 16060 | 9251 | -0.001 | 0.2544 | No | ||
| 96 | TPH2 | 19626 | 9290 | -0.001 | 0.2524 | No | ||
| 97 | USP26 | 24155 | 9297 | -0.001 | 0.2521 | No | ||
| 98 | SFRP5 | 23674 | 9303 | -0.001 | 0.2519 | No | ||
| 99 | SNTG1 | 7717 | 9602 | -0.001 | 0.2358 | No | ||
| 100 | JARID2 | 9200 | 9651 | -0.001 | 0.2333 | No | ||
| 101 | PVRL2 | 9671 5336 | 9696 | -0.001 | 0.2310 | No | ||
| 102 | FMO1 | 8976 | 9725 | -0.001 | 0.2295 | No | ||
| 103 | GPR44 | 23926 | 9833 | -0.001 | 0.2238 | No | ||
| 104 | MYCT1 | 7572 | 10265 | -0.002 | 0.2006 | No | ||
| 105 | KCNJ15 | 4943 9208 | 10358 | -0.002 | 0.1958 | No | ||
| 106 | ECT2 | 15367 | 10521 | -0.003 | 0.1872 | No | ||
| 107 | A2BP1 | 11212 1652 1689 | 10571 | -0.003 | 0.1847 | No | ||
| 108 | SLC18A1 | 18862 | 10695 | -0.003 | 0.1783 | No | ||
| 109 | CDON | 12195 | 10738 | -0.003 | 0.1762 | No | ||
| 110 | SPINK4 | 16241 | 10809 | -0.003 | 0.1726 | No | ||
| 111 | BMP6 | 8659 | 10887 | -0.003 | 0.1687 | No | ||
| 112 | SSTR3 | 22218 | 10934 | -0.003 | 0.1664 | No | ||
| 113 | SLC10A2 | 18939 | 10981 | -0.003 | 0.1642 | No | ||
| 114 | HOXA4 | 17148 | 11123 | -0.004 | 0.1568 | No | ||
| 115 | FOSL2 | 4733 8978 16878 | 11130 | -0.004 | 0.1567 | No | ||
| 116 | CACNA2D3 | 8677 | 11190 | -0.004 | 0.1538 | No | ||
| 117 | PITX2 | 15424 1878 | 11271 | -0.004 | 0.1498 | No | ||
| 118 | SIX1 | 9821 | 11513 | -0.005 | 0.1370 | No | ||
| 119 | HOXB6 | 20688 | 11908 | -0.005 | 0.1161 | No | ||
| 120 | PSMA1 | 1627 17669 | 12137 | -0.006 | 0.1041 | No | ||
| 121 | MOS | 16254 | 12210 | -0.006 | 0.1007 | No | ||
| 122 | DOCK8 | 13170 | 12223 | -0.006 | 0.1005 | No | ||
| 123 | SKIL | 15617 | 12273 | -0.006 | 0.0982 | No | ||
| 124 | DDR2 | 13767 | 12274 | -0.006 | 0.0987 | No | ||
| 125 | LSAMP | 1690 6500 | 12346 | -0.007 | 0.0953 | No | ||
| 126 | NRAS | 5191 | 12478 | -0.007 | 0.0887 | No | ||
| 127 | ADCYAP1 | 4348 | 12612 | -0.007 | 0.0820 | No | ||
| 128 | FST | 8985 | 12685 | -0.008 | 0.0787 | No | ||
| 129 | LECT2 | 21443 | 12707 | -0.008 | 0.0781 | No | ||
| 130 | ANGPT2 | 14885 3783 18923 | 12795 | -0.008 | 0.0739 | No | ||
| 131 | SPRY4 | 23448 | 13025 | -0.009 | 0.0621 | No | ||
| 132 | DSPP | 8867 | 13172 | -0.009 | 0.0548 | No | ||
| 133 | BPGM | 17489 | 13279 | -0.010 | 0.0498 | No | ||
| 134 | MASP1 | 1632 22626 | 13328 | -0.010 | 0.0479 | No | ||
| 135 | ERG | 1686 8915 | 13370 | -0.010 | 0.0464 | No | ||
| 136 | EYA1 | 4695 4061 | 13430 | -0.010 | 0.0439 | No | ||
| 137 | TBX4 | 5633 | 13460 | -0.010 | 0.0431 | No | ||
| 138 | CREB5 | 10551 | 13526 | -0.011 | 0.0403 | No | ||
| 139 | FABP2 | 15430 | 13550 | -0.011 | 0.0399 | No | ||
| 140 | LHX6 | 9275 2914 | 13604 | -0.011 | 0.0378 | No | ||
| 141 | LIMS1 | 8493 4290 | 13625 | -0.011 | 0.0375 | No | ||
| 142 | BSN | 18997 | 13882 | -0.013 | 0.0245 | No | ||
| 143 | LRCH2 | 24035 | 14019 | -0.014 | 0.0181 | No | ||
| 144 | ESRRG | 6447 | 14248 | -0.015 | 0.0069 | No | ||
| 145 | VSNL1 | 6527 | 14440 | -0.017 | -0.0023 | No | ||
| 146 | HOXA1 | 1005 17152 | 14529 | -0.018 | -0.0058 | No | ||
| 147 | TBX5 | 3531 5634 | 14576 | -0.018 | -0.0070 | No | ||
| 148 | GFRA3 | 23469 | 14612 | -0.018 | -0.0076 | No | ||
| 149 | HNRPA2B1 | 1187 7001 7000 1040 | 14647 | -0.019 | -0.0081 | No | ||
| 150 | TYRO3 | 5811 | 14755 | -0.020 | -0.0125 | No | ||
| 151 | SALL2 | 21840 | 14765 | -0.020 | -0.0116 | No | ||
| 152 | ELAVL4 | 15805 4889 9137 | 14859 | -0.021 | -0.0151 | No | ||
| 153 | CDH2 | 1963 8727 8726 4508 | 14992 | -0.023 | -0.0207 | No | ||
| 154 | CACNB2 | 4466 8678 | 15004 | -0.023 | -0.0196 | No | ||
| 155 | RGS3 | 2434 2474 6937 11935 2374 2332 2445 2390 | 15048 | -0.024 | -0.0203 | No | ||
| 156 | ATP6V0A4 | 8934 | 15220 | -0.026 | -0.0277 | No | ||
| 157 | CXCL13 | 16789 | 15469 | -0.031 | -0.0389 | No | ||
| 158 | SNX1 | 19076 | 15597 | -0.034 | -0.0433 | No | ||
| 159 | RHAG | 1545 1591 23235 | 15609 | -0.035 | -0.0415 | No | ||
| 160 | PCDH9 | 21736 | 15635 | -0.035 | -0.0403 | No | ||
| 161 | ETV5 | 22630 | 15983 | -0.046 | -0.0558 | No | ||
| 162 | RBPMS | 5369 | 16056 | -0.049 | -0.0562 | No | ||
| 163 | FBS1 | 18067 | 16336 | -0.062 | -0.0670 | No | ||
| 164 | HOXC4 | 4864 | 16394 | -0.064 | -0.0655 | No | ||
| 165 | PPARGC1A | 16533 | 16471 | -0.068 | -0.0648 | No | ||
| 166 | KCNK5 | 4946 9212 | 16537 | -0.071 | -0.0633 | No | ||
| 167 | SPOP | 5498 | 16848 | -0.093 | -0.0735 | No | ||
| 168 | PDE3B | 5236 | 17283 | -0.132 | -0.0876 | No | ||
| 169 | RAI1 | 5355 20861 | 17751 | -0.187 | -0.0996 | No | ||
| 170 | ESRRB | 21196 | 17859 | -0.205 | -0.0908 | No | ||
| 171 | TNFSF10 | 15626 | 18208 | -0.325 | -0.0865 | No | ||
| 172 | ZNF219 | 12835 | 18361 | -0.461 | -0.0619 | No | ||
| 173 | UTX | 10266 2574 | 18399 | -0.516 | -0.0271 | No | ||
| 174 | DIAPH3 | 7117 | 18411 | -0.544 | 0.0111 | No |