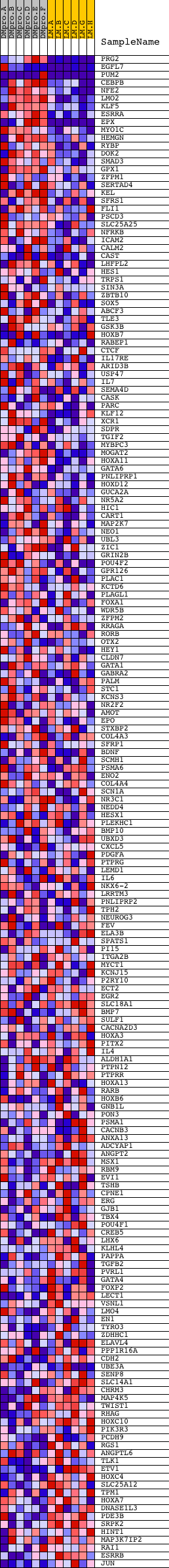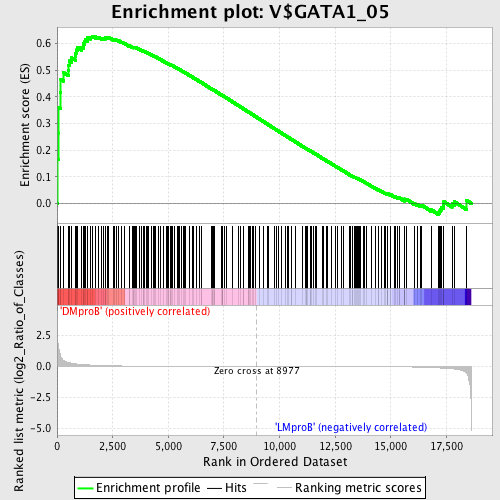
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_DMproB_versus_LMproB.phenotype_DMproB_versus_LMproB.cls #DMproB_versus_LMproB |
| Phenotype | phenotype_DMproB_versus_LMproB.cls#DMproB_versus_LMproB |
| Upregulated in class | DMproB |
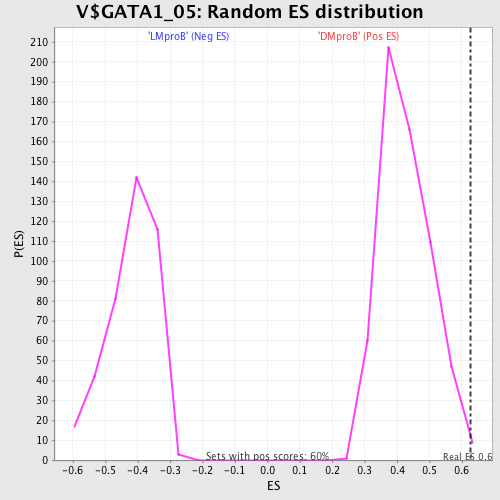
| GeneSet | V$GATA1_05 |
| Enrichment Score (ES) | 0.6273074 |
| Normalized Enrichment Score (NES) | 1.4711957 |
| Nominal p-value | 0.008347246 |
| FDR q-value | 0.70926994 |
| FWER p-Value | 0.998 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PRG2 | 14972 | 18 | 2.539 | 0.1652 | Yes | ||
| 2 | EGFL7 | 87 11687 2706 | 54 | 1.540 | 0.2641 | Yes | ||
| 3 | PUM2 | 2071 8161 | 58 | 1.485 | 0.3611 | Yes | ||
| 4 | CEBPB | 8733 | 151 | 0.910 | 0.4157 | Yes | ||
| 5 | NFE2 | 9456 | 170 | 0.762 | 0.4646 | Yes | ||
| 6 | LMO2 | 9281 | 280 | 0.490 | 0.4907 | Yes | ||
| 7 | KLF5 | 4456 | 502 | 0.312 | 0.4992 | Yes | ||
| 8 | ESRRA | 23802 | 527 | 0.299 | 0.5174 | Yes | ||
| 9 | EPX | 20302 | 559 | 0.284 | 0.5343 | Yes | ||
| 10 | MYO1C | 20777 | 651 | 0.254 | 0.5460 | Yes | ||
| 11 | HEMGN | 15888 | 818 | 0.208 | 0.5506 | Yes | ||
| 12 | RYBP | 17056 | 829 | 0.206 | 0.5636 | Yes | ||
| 13 | DOK2 | 354 | 854 | 0.201 | 0.5754 | Yes | ||
| 14 | SMAD3 | 19084 | 925 | 0.187 | 0.5839 | Yes | ||
| 15 | GPX1 | 19310 | 1094 | 0.159 | 0.5852 | Yes | ||
| 16 | ZFPM1 | 18439 | 1170 | 0.150 | 0.5909 | Yes | ||
| 17 | SERTAD4 | 13715 | 1185 | 0.147 | 0.5998 | Yes | ||
| 18 | KEL | 17170 | 1248 | 0.140 | 0.6056 | Yes | ||
| 19 | SFRS1 | 8492 | 1253 | 0.139 | 0.6144 | Yes | ||
| 20 | FLI1 | 4729 | 1367 | 0.124 | 0.6164 | Yes | ||
| 21 | PSCD3 | 16633 | 1379 | 0.123 | 0.6239 | Yes | ||
| 22 | SLC25A25 | 10431 | 1520 | 0.108 | 0.6233 | Yes | ||
| 23 | NFRKB | 10642 | 1573 | 0.104 | 0.6273 | Yes | ||
| 24 | ICAM2 | 20183 | 1713 | 0.092 | 0.6258 | No | ||
| 25 | CALM2 | 8681 | 1856 | 0.082 | 0.6235 | No | ||
| 26 | CAST | 3182 4478 3177 | 2016 | 0.071 | 0.6195 | No | ||
| 27 | LHFPL2 | 21584 | 2101 | 0.065 | 0.6192 | No | ||
| 28 | HES1 | 22798 | 2155 | 0.063 | 0.6204 | No | ||
| 29 | TRPS1 | 8195 | 2172 | 0.062 | 0.6236 | No | ||
| 30 | SIN3A | 5442 | 2277 | 0.057 | 0.6217 | No | ||
| 31 | ZBTB10 | 10468 | 2323 | 0.055 | 0.6228 | No | ||
| 32 | SOX5 | 9850 1044 5480 16937 | 2527 | 0.046 | 0.6148 | No | ||
| 33 | ABCF3 | 6578 | 2574 | 0.044 | 0.6152 | No | ||
| 34 | TLE3 | 5767 19416 | 2668 | 0.041 | 0.6129 | No | ||
| 35 | GSK3B | 22761 | 2764 | 0.038 | 0.6102 | No | ||
| 36 | HOXB7 | 666 9110 | 2905 | 0.034 | 0.6048 | No | ||
| 37 | RABEP1 | 12016 1344 | 3028 | 0.031 | 0.6002 | No | ||
| 38 | CTCF | 18490 | 3252 | 0.026 | 0.5898 | No | ||
| 39 | IL17RE | 434 17327 433 1049 | 3264 | 0.026 | 0.5909 | No | ||
| 40 | ARID3B | 12111 | 3406 | 0.024 | 0.5848 | No | ||
| 41 | USP47 | 6794 | 3445 | 0.023 | 0.5843 | No | ||
| 42 | IL7 | 4921 | 3448 | 0.023 | 0.5857 | No | ||
| 43 | SEMA4D | 9800 | 3495 | 0.022 | 0.5846 | No | ||
| 44 | CASK | 24184 2618 2604 | 3528 | 0.022 | 0.5843 | No | ||
| 45 | PARC | 22962 | 3561 | 0.021 | 0.5840 | No | ||
| 46 | KLF12 | 4960 2637 21732 9228 | 3713 | 0.019 | 0.5770 | No | ||
| 47 | XCR1 | 18953 | 3805 | 0.018 | 0.5733 | No | ||
| 48 | SDPR | 14252 | 3863 | 0.018 | 0.5714 | No | ||
| 49 | TGIF2 | 6013 6012 | 3881 | 0.017 | 0.5716 | No | ||
| 50 | MYBPC3 | 9434 | 3949 | 0.017 | 0.5690 | No | ||
| 51 | MOGAT2 | 17747 | 4005 | 0.016 | 0.5671 | No | ||
| 52 | HOXA11 | 17146 | 4084 | 0.015 | 0.5639 | No | ||
| 53 | GATA6 | 96 | 4127 | 0.015 | 0.5626 | No | ||
| 54 | PNLIPRP1 | 23835 | 4242 | 0.014 | 0.5573 | No | ||
| 55 | HOXD12 | 9113 | 4326 | 0.013 | 0.5537 | No | ||
| 56 | GUCA2A | 16104 | 4389 | 0.013 | 0.5512 | No | ||
| 57 | NR5A2 | 4105 13817 | 4432 | 0.013 | 0.5497 | No | ||
| 58 | HIC1 | 20350 | 4537 | 0.012 | 0.5449 | No | ||
| 59 | CART1 | 19634 | 4638 | 0.012 | 0.5402 | No | ||
| 60 | MAP2K7 | 6453 | 4764 | 0.011 | 0.5341 | No | ||
| 61 | NEO1 | 5161 | 4936 | 0.010 | 0.5255 | No | ||
| 62 | UBL3 | 16284 | 4939 | 0.010 | 0.5261 | No | ||
| 63 | ZIC1 | 19040 | 4985 | 0.010 | 0.5243 | No | ||
| 64 | GRIN2B | 16957 | 5022 | 0.010 | 0.5230 | No | ||
| 65 | POU4F2 | 9603 5276 | 5100 | 0.009 | 0.5194 | No | ||
| 66 | GPR126 | 19816 | 5116 | 0.009 | 0.5192 | No | ||
| 67 | PLAC1 | 12079 | 5133 | 0.009 | 0.5189 | No | ||
| 68 | KCTD6 | 22098 | 5171 | 0.009 | 0.5175 | No | ||
| 69 | PLAGL1 | 10372 3414 | 5260 | 0.009 | 0.5133 | No | ||
| 70 | FOXA1 | 21060 | 5421 | 0.008 | 0.5052 | No | ||
| 71 | WDR5B | 22775 | 5449 | 0.008 | 0.5042 | No | ||
| 72 | ZFPM2 | 22483 | 5490 | 0.008 | 0.5025 | No | ||
| 73 | RRAGA | 16180 | 5595 | 0.008 | 0.4974 | No | ||
| 74 | RORB | 10356 | 5677 | 0.007 | 0.4935 | No | ||
| 75 | OTX2 | 9519 | 5727 | 0.007 | 0.4913 | No | ||
| 76 | HEY1 | 15378 | 5781 | 0.007 | 0.4888 | No | ||
| 77 | CLDN7 | 20817 | 5950 | 0.006 | 0.4801 | No | ||
| 78 | GATA1 | 24196 | 6065 | 0.006 | 0.4744 | No | ||
| 79 | GABRA2 | 4746 | 6149 | 0.006 | 0.4702 | No | ||
| 80 | PALM | 19960 3350 | 6269 | 0.005 | 0.4641 | No | ||
| 81 | STC1 | 21974 | 6380 | 0.005 | 0.4585 | No | ||
| 82 | KCNS3 | 21109 | 6475 | 0.005 | 0.4537 | No | ||
| 83 | NR2F2 | 8619 409 | 6497 | 0.005 | 0.4529 | No | ||
| 84 | AMOT | 11341 24037 6587 | 6921 | 0.004 | 0.4302 | No | ||
| 85 | EPO | 8911 | 6982 | 0.004 | 0.4272 | No | ||
| 86 | STXBP2 | 18700 | 6986 | 0.004 | 0.4273 | No | ||
| 87 | COL4A3 | 8766 | 7015 | 0.004 | 0.4260 | No | ||
| 88 | SFRP1 | 9805 | 7061 | 0.004 | 0.4238 | No | ||
| 89 | BDNF | 14926 2797 | 7067 | 0.004 | 0.4237 | No | ||
| 90 | SCMH1 | 2351 6618 | 7378 | 0.003 | 0.4071 | No | ||
| 91 | PSMA6 | 21270 | 7419 | 0.003 | 0.4051 | No | ||
| 92 | ENO2 | 8904 | 7436 | 0.003 | 0.4044 | No | ||
| 93 | COL4A4 | 8767 | 7442 | 0.003 | 0.4043 | No | ||
| 94 | SCN1A | 9784 | 7541 | 0.003 | 0.3992 | No | ||
| 95 | NR3C1 | 9043 | 7598 | 0.003 | 0.3963 | No | ||
| 96 | NEDD4 | 19384 3101 | 7900 | 0.002 | 0.3801 | No | ||
| 97 | HESX1 | 22070 | 8145 | 0.001 | 0.3670 | No | ||
| 98 | PLEKHC1 | 21864 | 8226 | 0.001 | 0.3627 | No | ||
| 99 | BMP10 | 17376 | 8365 | 0.001 | 0.3553 | No | ||
| 100 | UBXD3 | 15701 | 8382 | 0.001 | 0.3545 | No | ||
| 101 | CXCL5 | 16802 | 8583 | 0.001 | 0.3437 | No | ||
| 102 | PDGFA | 5237 | 8589 | 0.001 | 0.3434 | No | ||
| 103 | PTPRG | 22097 3151 | 8637 | 0.001 | 0.3409 | No | ||
| 104 | LEMD1 | 10023 537 | 8702 | 0.000 | 0.3375 | No | ||
| 105 | IL6 | 16895 | 8787 | 0.000 | 0.3329 | No | ||
| 106 | NKX6-2 | 17579 4815 | 8805 | 0.000 | 0.3320 | No | ||
| 107 | LRRTM3 | 19744 | 8922 | 0.000 | 0.3258 | No | ||
| 108 | PNLIPRP2 | 23836 | 9085 | -0.000 | 0.3170 | No | ||
| 109 | TPH2 | 19626 | 9290 | -0.001 | 0.3059 | No | ||
| 110 | NEUROG3 | 20005 | 9459 | -0.001 | 0.2969 | No | ||
| 111 | FEV | 11153 6434 | 9520 | -0.001 | 0.2937 | No | ||
| 112 | ELA3B | 12592 7494 | 9772 | -0.001 | 0.2801 | No | ||
| 113 | SPATS1 | 22973 | 9847 | -0.001 | 0.2762 | No | ||
| 114 | PI15 | 13586 | 9948 | -0.002 | 0.2709 | No | ||
| 115 | ITGA2B | 9186 | 10070 | -0.002 | 0.2644 | No | ||
| 116 | MYCT1 | 7572 | 10265 | -0.002 | 0.2540 | No | ||
| 117 | KCNJ15 | 4943 9208 | 10358 | -0.002 | 0.2492 | No | ||
| 118 | P2RY10 | 2613 24269 | 10394 | -0.002 | 0.2475 | No | ||
| 119 | ECT2 | 15367 | 10521 | -0.003 | 0.2408 | No | ||
| 120 | EGR2 | 8886 | 10551 | -0.003 | 0.2394 | No | ||
| 121 | SLC18A1 | 18862 | 10695 | -0.003 | 0.2318 | No | ||
| 122 | BMP7 | 14324 | 11007 | -0.004 | 0.2152 | No | ||
| 123 | SULF1 | 6285 | 11182 | -0.004 | 0.2060 | No | ||
| 124 | CACNA2D3 | 8677 | 11190 | -0.004 | 0.2058 | No | ||
| 125 | HOXA3 | 9108 1075 | 11197 | -0.004 | 0.2058 | No | ||
| 126 | PITX2 | 15424 1878 | 11271 | -0.004 | 0.2021 | No | ||
| 127 | IL4 | 9174 | 11369 | -0.004 | 0.1971 | No | ||
| 128 | ALDH1A1 | 8569 | 11399 | -0.004 | 0.1958 | No | ||
| 129 | PTPN12 | 16597 3502 3586 3665 | 11440 | -0.004 | 0.1939 | No | ||
| 130 | PTPRR | 3436 19876 | 11540 | -0.005 | 0.1889 | No | ||
| 131 | HOXA13 | 9107 | 11614 | -0.005 | 0.1852 | No | ||
| 132 | RARB | 3011 21915 | 11666 | -0.005 | 0.1828 | No | ||
| 133 | HOXB6 | 20688 | 11908 | -0.005 | 0.1700 | No | ||
| 134 | GNB1L | 8921 22827 | 11980 | -0.006 | 0.1666 | No | ||
| 135 | PON3 | 1141 17223 | 12129 | -0.006 | 0.1589 | No | ||
| 136 | PSMA1 | 1627 17669 | 12137 | -0.006 | 0.1589 | No | ||
| 137 | CACNB3 | 22373 8679 | 12327 | -0.007 | 0.1491 | No | ||
| 138 | ANXA13 | 22287 | 12503 | -0.007 | 0.1401 | No | ||
| 139 | ADCYAP1 | 4348 | 12612 | -0.007 | 0.1347 | No | ||
| 140 | ANGPT2 | 14885 3783 18923 | 12795 | -0.008 | 0.1253 | No | ||
| 141 | MSX1 | 16547 | 12856 | -0.008 | 0.1226 | No | ||
| 142 | RBM9 | 2276 13528 | 13163 | -0.009 | 0.1066 | No | ||
| 143 | EVI1 | 4689 8926 | 13186 | -0.009 | 0.1060 | No | ||
| 144 | TSHB | 15218 | 13287 | -0.010 | 0.1012 | No | ||
| 145 | CPNE1 | 11186 | 13297 | -0.010 | 0.1013 | No | ||
| 146 | ERG | 1686 8915 | 13370 | -0.010 | 0.0981 | No | ||
| 147 | GJB1 | 24276 | 13398 | -0.010 | 0.0973 | No | ||
| 148 | TBX4 | 5633 | 13460 | -0.010 | 0.0946 | No | ||
| 149 | POU4F1 | 21727 | 13517 | -0.011 | 0.0923 | No | ||
| 150 | CREB5 | 10551 | 13526 | -0.011 | 0.0926 | No | ||
| 151 | LHX6 | 9275 2914 | 13604 | -0.011 | 0.0891 | No | ||
| 152 | KLHL4 | 24076 | 13639 | -0.011 | 0.0880 | No | ||
| 153 | PAPPA | 16190 | 13762 | -0.012 | 0.0822 | No | ||
| 154 | TGFB2 | 5743 | 13814 | -0.013 | 0.0803 | No | ||
| 155 | PVRL1 | 3006 19482 12211 7192 | 13902 | -0.013 | 0.0764 | No | ||
| 156 | GATA4 | 21792 4755 | 14125 | -0.014 | 0.0653 | No | ||
| 157 | FOXP2 | 4312 17523 8523 | 14306 | -0.016 | 0.0565 | No | ||
| 158 | LECT1 | 21740 | 14432 | -0.017 | 0.0509 | No | ||
| 159 | VSNL1 | 6527 | 14440 | -0.017 | 0.0516 | No | ||
| 160 | LMO4 | 15151 | 14582 | -0.018 | 0.0451 | No | ||
| 161 | EN1 | 387 14155 | 14727 | -0.019 | 0.0386 | No | ||
| 162 | TYRO3 | 5811 | 14755 | -0.020 | 0.0384 | No | ||
| 163 | ZDHHC1 | 18770 | 14832 | -0.021 | 0.0356 | No | ||
| 164 | ELAVL4 | 15805 4889 9137 | 14859 | -0.021 | 0.0356 | No | ||
| 165 | PPP1R16A | 22441 | 14869 | -0.021 | 0.0365 | No | ||
| 166 | CDH2 | 1963 8727 8726 4508 | 14992 | -0.023 | 0.0313 | No | ||
| 167 | UBE3A | 1209 1084 1830 | 14993 | -0.023 | 0.0328 | No | ||
| 168 | SENP8 | 7740 | 15160 | -0.025 | 0.0255 | No | ||
| 169 | SLC14A1 | 4231 23403 8425 | 15204 | -0.026 | 0.0248 | No | ||
| 170 | CHRM3 | 21547 | 15318 | -0.028 | 0.0205 | No | ||
| 171 | MAP4K5 | 11853 21048 2101 2142 | 15377 | -0.029 | 0.0193 | No | ||
| 172 | TWIST1 | 21291 | 15387 | -0.029 | 0.0207 | No | ||
| 173 | RHAG | 1545 1591 23235 | 15609 | -0.035 | 0.0110 | No | ||
| 174 | HOXC10 | 22342 | 15612 | -0.035 | 0.0132 | No | ||
| 175 | PIK3R3 | 5248 | 15620 | -0.035 | 0.0151 | No | ||
| 176 | PCDH9 | 21736 | 15635 | -0.035 | 0.0166 | No | ||
| 177 | RGS1 | 3970 6936 | 15699 | -0.037 | 0.0156 | No | ||
| 178 | ANGPTL6 | 19218 | 16067 | -0.050 | -0.0011 | No | ||
| 179 | TLK1 | 14563 | 16181 | -0.054 | -0.0036 | No | ||
| 180 | ETV1 | 4688 8925 8924 | 16353 | -0.062 | -0.0088 | No | ||
| 181 | HOXC4 | 4864 | 16394 | -0.064 | -0.0068 | No | ||
| 182 | SLC25A12 | 14562 | 16829 | -0.092 | -0.0244 | No | ||
| 183 | TPM1 | 19070 | 17146 | -0.117 | -0.0339 | No | ||
| 184 | HOXA7 | 4862 | 17192 | -0.121 | -0.0284 | No | ||
| 185 | DNASE1L3 | 21925 | 17234 | -0.126 | -0.0223 | No | ||
| 186 | PDE3B | 5236 | 17283 | -0.132 | -0.0163 | No | ||
| 187 | SRPK2 | 5513 | 17366 | -0.141 | -0.0115 | No | ||
| 188 | HINT1 | 20881 | 17377 | -0.142 | -0.0028 | No | ||
| 189 | MAP3K7IP2 | 19827 | 17385 | -0.143 | 0.0062 | No | ||
| 190 | RAI1 | 5355 20861 | 17751 | -0.187 | -0.0014 | No | ||
| 191 | ESRRB | 21196 | 17859 | -0.205 | 0.0063 | No | ||
| 192 | JUN | 15832 | 18408 | -0.531 | 0.0113 | No |