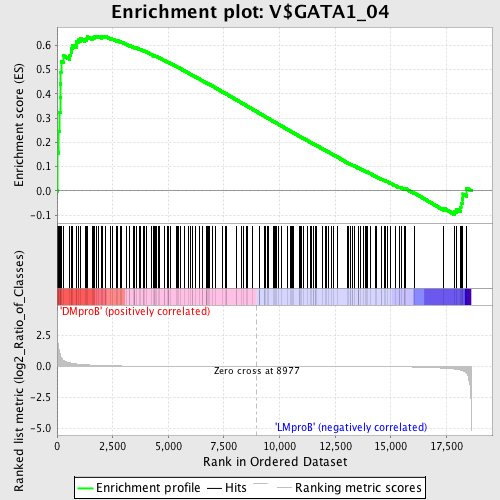
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_DMproB_versus_LMproB.phenotype_DMproB_versus_LMproB.cls #DMproB_versus_LMproB |
| Phenotype | phenotype_DMproB_versus_LMproB.cls#DMproB_versus_LMproB |
| Upregulated in class | DMproB |
| GeneSet | V$GATA1_04 |
| Enrichment Score (ES) | 0.6385554 |
| Normalized Enrichment Score (NES) | 1.4938216 |
| Nominal p-value | 0.0076923077 |
| FDR q-value | 0.6654779 |
| FWER p-Value | 0.989 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PRG2 | 14972 | 18 | 2.539 | 0.1578 | Yes | ||
| 2 | UBE2H | 5823 5822 | 68 | 1.432 | 0.2447 | Yes | ||
| 3 | TNFRSF19 | 21799 3033 | 98 | 1.294 | 0.3241 | Yes | ||
| 4 | SLC4A1 | 20204 | 136 | 1.010 | 0.3852 | Yes | ||
| 5 | CEBPB | 8733 | 151 | 0.910 | 0.4413 | Yes | ||
| 6 | NFE2 | 9456 | 170 | 0.762 | 0.4880 | Yes | ||
| 7 | LYL1 | 18546 | 177 | 0.727 | 0.5332 | Yes | ||
| 8 | LMO2 | 9281 | 280 | 0.490 | 0.5583 | Yes | ||
| 9 | GADD45G | 21637 | 573 | 0.279 | 0.5599 | Yes | ||
| 10 | KLF1 | 18544 | 624 | 0.263 | 0.5737 | Yes | ||
| 11 | MYO1C | 20777 | 651 | 0.254 | 0.5881 | Yes | ||
| 12 | RAB24 | 12584 5344 | 700 | 0.238 | 0.6004 | Yes | ||
| 13 | MBNL2 | 21932 | 859 | 0.199 | 0.6043 | Yes | ||
| 14 | ARPC2 | 14225 | 867 | 0.198 | 0.6164 | Yes | ||
| 15 | ADM | 18125 | 961 | 0.181 | 0.6227 | Yes | ||
| 16 | CDC42EP3 | 6436 | 1048 | 0.167 | 0.6284 | Yes | ||
| 17 | SFRS1 | 8492 | 1253 | 0.139 | 0.6260 | Yes | ||
| 18 | CHCHD7 | 16280 | 1337 | 0.127 | 0.6295 | Yes | ||
| 19 | FLI1 | 4729 | 1367 | 0.124 | 0.6357 | Yes | ||
| 20 | NFRKB | 10642 | 1573 | 0.104 | 0.6311 | Yes | ||
| 21 | USP52 | 3397 19841 | 1642 | 0.098 | 0.6335 | Yes | ||
| 22 | DMD | 24295 2647 | 1698 | 0.093 | 0.6363 | Yes | ||
| 23 | CCL27 | 5419 | 1761 | 0.089 | 0.6385 | Yes | ||
| 24 | CALM2 | 8681 | 1856 | 0.082 | 0.6386 | Yes | ||
| 25 | CAST | 3182 4478 3177 | 2016 | 0.071 | 0.6344 | No | ||
| 26 | ICAM4 | 19544 | 2045 | 0.069 | 0.6371 | No | ||
| 27 | HES1 | 22798 | 2155 | 0.063 | 0.6352 | No | ||
| 28 | TRPS1 | 8195 | 2172 | 0.062 | 0.6381 | No | ||
| 29 | SLC35A2 | 24389 | 2405 | 0.051 | 0.6288 | No | ||
| 30 | IFRD2 | 19320 | 2507 | 0.047 | 0.6262 | No | ||
| 31 | GBX2 | 13887 | 2661 | 0.041 | 0.6205 | No | ||
| 32 | AQP2 | 22366 | 2707 | 0.040 | 0.6205 | No | ||
| 33 | IGF1 | 3352 9156 3409 | 2839 | 0.035 | 0.6156 | No | ||
| 34 | HOXB7 | 666 9110 | 2905 | 0.034 | 0.6142 | No | ||
| 35 | PRDX2 | 3871 18542 5705 | 3101 | 0.029 | 0.6055 | No | ||
| 36 | CTCF | 18490 | 3252 | 0.026 | 0.5990 | No | ||
| 37 | PKIA | 15639 | 3272 | 0.026 | 0.5996 | No | ||
| 38 | HIPK1 | 4851 | 3439 | 0.023 | 0.5920 | No | ||
| 39 | CNTF | 3741 8761 3680 3736 | 3489 | 0.022 | 0.5908 | No | ||
| 40 | ANK2 | 4275 | 3492 | 0.022 | 0.5921 | No | ||
| 41 | SEMA4D | 9800 | 3495 | 0.022 | 0.5934 | No | ||
| 42 | STAG2 | 5521 | 3577 | 0.021 | 0.5903 | No | ||
| 43 | KLF12 | 4960 2637 21732 9228 | 3713 | 0.019 | 0.5842 | No | ||
| 44 | TNNC1 | 22058 | 3746 | 0.019 | 0.5836 | No | ||
| 45 | PGGT1B | 10344 | 3873 | 0.017 | 0.5779 | No | ||
| 46 | HOXB9 | 20689 | 3875 | 0.017 | 0.5789 | No | ||
| 47 | MYBPC3 | 9434 | 3949 | 0.017 | 0.5760 | No | ||
| 48 | MOGAT2 | 17747 | 4005 | 0.016 | 0.5740 | No | ||
| 49 | PNLIPRP1 | 23835 | 4242 | 0.014 | 0.5621 | No | ||
| 50 | NHLH2 | 9462 | 4353 | 0.013 | 0.5569 | No | ||
| 51 | CMYA1 | 18966 | 4379 | 0.013 | 0.5564 | No | ||
| 52 | GUCA2A | 16104 | 4389 | 0.013 | 0.5567 | No | ||
| 53 | NR5A2 | 4105 13817 | 4432 | 0.013 | 0.5553 | No | ||
| 54 | CAPN1 | 23989 | 4459 | 0.013 | 0.5546 | No | ||
| 55 | HIC1 | 20350 | 4537 | 0.012 | 0.5512 | No | ||
| 56 | BNC2 | 15850 | 4580 | 0.012 | 0.5497 | No | ||
| 57 | EYA4 | 8937 | 4831 | 0.011 | 0.5368 | No | ||
| 58 | LEPREL1 | 5575 | 4842 | 0.010 | 0.5369 | No | ||
| 59 | SLITRK2 | 24322 | 4950 | 0.010 | 0.5317 | No | ||
| 60 | ZIC1 | 19040 | 4985 | 0.010 | 0.5305 | No | ||
| 61 | AMHR2 | 8487 22345 | 4990 | 0.010 | 0.5309 | No | ||
| 62 | GRIN2B | 16957 | 5022 | 0.010 | 0.5298 | No | ||
| 63 | POU4F2 | 9603 5276 | 5100 | 0.009 | 0.5263 | No | ||
| 64 | IMPG1 | 19052 | 5368 | 0.008 | 0.5123 | No | ||
| 65 | HOXA10 | 17145 989 | 5413 | 0.008 | 0.5104 | No | ||
| 66 | ISL1 | 21338 | 5462 | 0.008 | 0.5083 | No | ||
| 67 | PRKAA1 | 22520 | 5526 | 0.008 | 0.5054 | No | ||
| 68 | CD34 | 14006 | 5733 | 0.007 | 0.4946 | No | ||
| 69 | INHA | 14214 | 5901 | 0.007 | 0.4860 | No | ||
| 70 | ABCA12 | 13931 | 6005 | 0.006 | 0.4808 | No | ||
| 71 | GATA1 | 24196 | 6065 | 0.006 | 0.4780 | No | ||
| 72 | SLC26A9 | 11560 980 4067 | 6087 | 0.006 | 0.4772 | No | ||
| 73 | ABCG5 | 22884 | 6231 | 0.006 | 0.4698 | No | ||
| 74 | NEUROD6 | 17138 | 6389 | 0.005 | 0.4616 | No | ||
| 75 | SLC41A1 | 8269 | 6530 | 0.005 | 0.4543 | No | ||
| 76 | SLC26A7 | 5538 2372 | 6547 | 0.005 | 0.4537 | No | ||
| 77 | PPIL5 | 21255 | 6696 | 0.004 | 0.4460 | No | ||
| 78 | SYN1 | 24175 5558 | 6708 | 0.004 | 0.4457 | No | ||
| 79 | GPX5 | 21532 3242 | 6715 | 0.004 | 0.4456 | No | ||
| 80 | NRP2 | 5194 9486 5195 | 6758 | 0.004 | 0.4436 | No | ||
| 81 | MITF | 17349 | 6811 | 0.004 | 0.4410 | No | ||
| 82 | MAP4K3 | 22887 | 6849 | 0.004 | 0.4393 | No | ||
| 83 | EPO | 8911 | 6982 | 0.004 | 0.4324 | No | ||
| 84 | CYP11A1 | 4580 | 7102 | 0.003 | 0.4261 | No | ||
| 85 | ENO2 | 8904 | 7436 | 0.003 | 0.4082 | No | ||
| 86 | PCDH7 | 7029 3460 | 7582 | 0.003 | 0.4005 | No | ||
| 87 | NR3C1 | 9043 | 7598 | 0.003 | 0.3999 | No | ||
| 88 | RAB3C | 21351 | 7602 | 0.003 | 0.3999 | No | ||
| 89 | PDGFRA | 16824 | 8042 | 0.002 | 0.3762 | No | ||
| 90 | MST1 | 9089 | 8073 | 0.002 | 0.3747 | No | ||
| 91 | C3AR1 | 8668 | 8275 | 0.001 | 0.3638 | No | ||
| 92 | TCF21 | 3437 | 8376 | 0.001 | 0.3585 | No | ||
| 93 | APOBEC2 | 22946 | 8495 | 0.001 | 0.3521 | No | ||
| 94 | DRD3 | 22750 | 8535 | 0.001 | 0.3501 | No | ||
| 95 | ROS1 | 19771 | 8789 | 0.000 | 0.3364 | No | ||
| 96 | PNLIPRP2 | 23836 | 9085 | -0.000 | 0.3204 | No | ||
| 97 | HAMP | 17878 | 9092 | -0.000 | 0.3201 | No | ||
| 98 | FZD4 | 18193 | 9098 | -0.000 | 0.3198 | No | ||
| 99 | SFRP5 | 23674 | 9303 | -0.001 | 0.3088 | No | ||
| 100 | CNN1 | 8759 | 9326 | -0.001 | 0.3076 | No | ||
| 101 | CALD1 | 4273 8463 | 9344 | -0.001 | 0.3067 | No | ||
| 102 | NEUROG3 | 20005 | 9459 | -0.001 | 0.3006 | No | ||
| 103 | FEV | 11153 6434 | 9520 | -0.001 | 0.2974 | No | ||
| 104 | FMO1 | 8976 | 9725 | -0.001 | 0.2864 | No | ||
| 105 | ELA3B | 12592 7494 | 9772 | -0.001 | 0.2840 | No | ||
| 106 | GPR44 | 23926 | 9833 | -0.001 | 0.2809 | No | ||
| 107 | SPATS1 | 22973 | 9847 | -0.001 | 0.2802 | No | ||
| 108 | PI15 | 13586 | 9948 | -0.002 | 0.2749 | No | ||
| 109 | COL17A1 | 23647 3671 | 10079 | -0.002 | 0.2680 | No | ||
| 110 | HNF4G | 15640 | 10349 | -0.002 | 0.2535 | No | ||
| 111 | PRSS1 | 5800 | 10509 | -0.003 | 0.2451 | No | ||
| 112 | EGR2 | 8886 | 10551 | -0.003 | 0.2430 | No | ||
| 113 | DNAJB4 | 12451 1916 1832 | 10586 | -0.003 | 0.2413 | No | ||
| 114 | CLCN5 | 24204 | 10611 | -0.003 | 0.2402 | No | ||
| 115 | BMP6 | 8659 | 10887 | -0.003 | 0.2255 | No | ||
| 116 | HOXC12 | 22343 | 10911 | -0.003 | 0.2245 | No | ||
| 117 | GRID2 | 4803 | 10920 | -0.003 | 0.2242 | No | ||
| 118 | CSF2 | 20454 | 10990 | -0.003 | 0.2207 | No | ||
| 119 | AMBN | 4380 | 11059 | -0.004 | 0.2172 | No | ||
| 120 | PITX2 | 15424 1878 | 11271 | -0.004 | 0.2061 | No | ||
| 121 | ALDH1A1 | 8569 | 11399 | -0.004 | 0.1994 | No | ||
| 122 | IL5 | 20884 10220 | 11453 | -0.004 | 0.1968 | No | ||
| 123 | SIX1 | 9821 | 11513 | -0.005 | 0.1939 | No | ||
| 124 | PTPRR | 3436 19876 | 11540 | -0.005 | 0.1928 | No | ||
| 125 | ADAMTSL1 | 16181 | 11596 | -0.005 | 0.1901 | No | ||
| 126 | HOXA13 | 9107 | 11614 | -0.005 | 0.1895 | No | ||
| 127 | RARB | 3011 21915 | 11666 | -0.005 | 0.1870 | No | ||
| 128 | HOXD1 | 14981 | 11940 | -0.006 | 0.1726 | No | ||
| 129 | TFEC | 17213 | 12044 | -0.006 | 0.1674 | No | ||
| 130 | PON3 | 1141 17223 | 12129 | -0.006 | 0.1632 | No | ||
| 131 | MSR1 | 5417 | 12179 | -0.006 | 0.1609 | No | ||
| 132 | CACNB3 | 22373 8679 | 12327 | -0.007 | 0.1534 | No | ||
| 133 | IL3 | 20453 | 12431 | -0.007 | 0.1482 | No | ||
| 134 | CLDN14 | 12081 | 12585 | -0.007 | 0.1403 | No | ||
| 135 | LIN28 | 15723 | 13032 | -0.009 | 0.1167 | No | ||
| 136 | ABCG8 | 23146 | 13114 | -0.009 | 0.1129 | No | ||
| 137 | DSPP | 8867 | 13172 | -0.009 | 0.1104 | No | ||
| 138 | EVI1 | 4689 8926 | 13186 | -0.009 | 0.1102 | No | ||
| 139 | TSHB | 15218 | 13287 | -0.010 | 0.1054 | No | ||
| 140 | SLC8A3 | 2065 4293 | 13292 | -0.010 | 0.1058 | No | ||
| 141 | HTR7 | 23690 | 13350 | -0.010 | 0.1033 | No | ||
| 142 | PLAG1 | 7148 15949 12161 | 13525 | -0.011 | 0.0946 | No | ||
| 143 | CREB5 | 10551 | 13526 | -0.011 | 0.0952 | No | ||
| 144 | RHOBTB2 | 3069 6373 10902 | 13567 | -0.011 | 0.0938 | No | ||
| 145 | KIRREL2 | 17889 3922 | 13623 | -0.011 | 0.0915 | No | ||
| 146 | PAPPA | 16190 | 13762 | -0.012 | 0.0848 | No | ||
| 147 | BSN | 18997 | 13882 | -0.013 | 0.0791 | No | ||
| 148 | RORA | 3019 5388 | 13925 | -0.013 | 0.0777 | No | ||
| 149 | PNOC | 21783 | 13929 | -0.013 | 0.0783 | No | ||
| 150 | WNT6 | 5881 | 14077 | -0.014 | 0.0712 | No | ||
| 151 | FOXP2 | 4312 17523 8523 | 14306 | -0.016 | 0.0599 | No | ||
| 152 | TNXB | 1557 8174 878 8175 | 14366 | -0.016 | 0.0577 | No | ||
| 153 | TBX5 | 3531 5634 | 14576 | -0.018 | 0.0475 | No | ||
| 154 | NDRG2 | 21843 | 14583 | -0.018 | 0.0483 | No | ||
| 155 | EN1 | 387 14155 | 14727 | -0.019 | 0.0418 | No | ||
| 156 | GRM3 | 16929 | 14761 | -0.020 | 0.0412 | No | ||
| 157 | ZDHHC1 | 18770 | 14832 | -0.021 | 0.0387 | No | ||
| 158 | UBE3A | 1209 1084 1830 | 14993 | -0.023 | 0.0314 | No | ||
| 159 | SLC14A1 | 4231 23403 8425 | 15204 | -0.026 | 0.0217 | No | ||
| 160 | MAP4K5 | 11853 21048 2101 2142 | 15377 | -0.029 | 0.0142 | No | ||
| 161 | TWIST1 | 21291 | 15387 | -0.029 | 0.0155 | No | ||
| 162 | CXCL13 | 16789 | 15469 | -0.031 | 0.0131 | No | ||
| 163 | HOXC10 | 22342 | 15612 | -0.035 | 0.0076 | No | ||
| 164 | PIK3R3 | 5248 | 15620 | -0.035 | 0.0094 | No | ||
| 165 | MTMR4 | 20712 | 15646 | -0.036 | 0.0102 | No | ||
| 166 | RBPMS | 5369 | 16056 | -0.049 | -0.0089 | No | ||
| 167 | MAP3K7IP2 | 19827 | 17385 | -0.143 | -0.0720 | No | ||
| 168 | ESRRB | 21196 | 17859 | -0.205 | -0.0848 | No | ||
| 169 | EDG5 | 19216 | 17960 | -0.229 | -0.0759 | No | ||
| 170 | CPLX2 | 8779 | 18122 | -0.280 | -0.0671 | No | ||
| 171 | MAGED2 | 24034 | 18156 | -0.293 | -0.0506 | No | ||
| 172 | MSI2 | 8030 1268 | 18210 | -0.326 | -0.0331 | No | ||
| 173 | KIF3C | 9220 4954 | 18242 | -0.348 | -0.0130 | No | ||
| 174 | JUN | 15832 | 18408 | -0.531 | 0.0113 | No |

