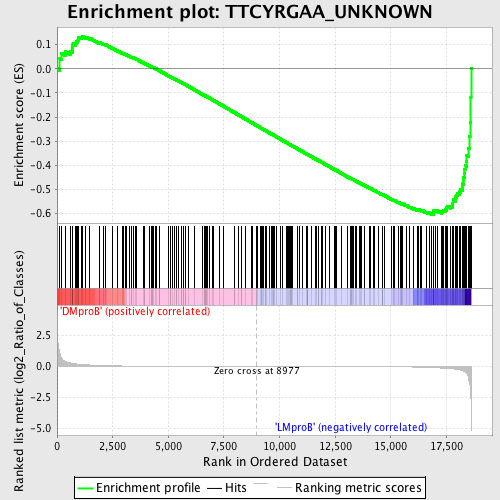
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_DMproB_versus_LMproB.phenotype_DMproB_versus_LMproB.cls #DMproB_versus_LMproB |
| Phenotype | phenotype_DMproB_versus_LMproB.cls#DMproB_versus_LMproB |
| Upregulated in class | LMproB |
| GeneSet | TTCYRGAA_UNKNOWN |
| Enrichment Score (ES) | -0.6035927 |
| Normalized Enrichment Score (NES) | -1.4772714 |
| Nominal p-value | 0.0069284067 |
| FDR q-value | 1.0 |
| FWER p-Value | 0.996 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | DNMT3B | 2840 14785 | 123 | 1.151 | 0.0414 | No | ||
| 2 | PROM1 | 5293 | 200 | 0.652 | 0.0645 | No | ||
| 3 | BCL6 | 22624 | 380 | 0.381 | 0.0707 | No | ||
| 4 | GTF3C2 | 7749 | 585 | 0.276 | 0.0712 | No | ||
| 5 | FRAG1 | 6129 18165 1724 | 690 | 0.242 | 0.0756 | No | ||
| 6 | RAB24 | 12584 5344 | 700 | 0.238 | 0.0851 | No | ||
| 7 | STAT4 | 14251 9907 | 707 | 0.235 | 0.0946 | No | ||
| 8 | WASL | 17203 | 713 | 0.234 | 0.1041 | No | ||
| 9 | UPF2 | 11591 | 821 | 0.207 | 0.1070 | No | ||
| 10 | ADRBK1 | 23960 | 877 | 0.196 | 0.1122 | No | ||
| 11 | PRDM2 | 2522 4289 | 935 | 0.186 | 0.1169 | No | ||
| 12 | GNA13 | 20617 | 946 | 0.184 | 0.1240 | No | ||
| 13 | AP3D1 | 19690 | 978 | 0.178 | 0.1298 | No | ||
| 14 | GPX1 | 19310 | 1094 | 0.159 | 0.1302 | No | ||
| 15 | ATP2C1 | 10660 6178 | 1118 | 0.156 | 0.1354 | No | ||
| 16 | ELOVL1 | 16114 2484 12026 | 1294 | 0.133 | 0.1315 | No | ||
| 17 | CDK5 | 16591 | 1476 | 0.113 | 0.1264 | No | ||
| 18 | BRD2 | 4735 8984 7628 | 1885 | 0.080 | 0.1076 | No | ||
| 19 | CDIPT | 18071 | 1899 | 0.080 | 0.1102 | No | ||
| 20 | CRSP7 | 18838 | 2104 | 0.065 | 0.1018 | No | ||
| 21 | MAPT | 9420 1347 | 2188 | 0.061 | 0.0999 | No | ||
| 22 | LTC4S | 20478 | 2489 | 0.048 | 0.0855 | No | ||
| 23 | PDE4C | 8481 | 2729 | 0.039 | 0.0742 | No | ||
| 24 | FBXO33 | 528 | 2940 | 0.033 | 0.0641 | No | ||
| 25 | AOAH | 11296 | 2993 | 0.032 | 0.0626 | No | ||
| 26 | VPS52 | 23287 1622 | 3076 | 0.030 | 0.0594 | No | ||
| 27 | LGI2 | 6371 | 3125 | 0.029 | 0.0580 | No | ||
| 28 | PPP2R5B | 23994 | 3236 | 0.027 | 0.0532 | No | ||
| 29 | PASK | 11242 | 3344 | 0.025 | 0.0484 | No | ||
| 30 | PRDX6 | 4389 8596 | 3356 | 0.024 | 0.0488 | No | ||
| 31 | TMC5 | 18111 | 3437 | 0.023 | 0.0454 | No | ||
| 32 | CLU | 21979 | 3520 | 0.022 | 0.0419 | No | ||
| 33 | STAG2 | 5521 | 3577 | 0.021 | 0.0397 | No | ||
| 34 | BCOR | 12934 2561 | 3862 | 0.018 | 0.0250 | No | ||
| 35 | PIK3R2 | 18850 | 3914 | 0.017 | 0.0230 | No | ||
| 36 | XPO1 | 4172 | 3943 | 0.017 | 0.0221 | No | ||
| 37 | NETO1 | 6372 1970 | 4143 | 0.015 | 0.0120 | No | ||
| 38 | COG7 | 10605 | 4224 | 0.014 | 0.0082 | No | ||
| 39 | SATB2 | 5604 | 4247 | 0.014 | 0.0076 | No | ||
| 40 | SELP | 14073 | 4282 | 0.014 | 0.0063 | No | ||
| 41 | IL22RA2 | 20082 | 4300 | 0.014 | 0.0060 | No | ||
| 42 | BTBD11 | 13147 | 4324 | 0.013 | 0.0053 | No | ||
| 43 | TAC4 | 20696 | 4439 | 0.013 | -0.0004 | No | ||
| 44 | TLR8 | 9308 | 4450 | 0.013 | -0.0004 | No | ||
| 45 | HSPE1 | 9133 | 4607 | 0.012 | -0.0084 | No | ||
| 46 | EVC2 | 16859 | 4623 | 0.012 | -0.0087 | No | ||
| 47 | ESR1 | 20097 4685 | 5004 | 0.010 | -0.0289 | No | ||
| 48 | FBXO5 | 19830 | 5098 | 0.009 | -0.0336 | No | ||
| 49 | MEPE | 16774 | 5113 | 0.009 | -0.0340 | No | ||
| 50 | SCN3A | 14573 | 5187 | 0.009 | -0.0376 | No | ||
| 51 | PER2 | 3940 13881 | 5277 | 0.009 | -0.0420 | No | ||
| 52 | CBX3 | 8705 4484 | 5346 | 0.008 | -0.0454 | No | ||
| 53 | DNAJA1 | 4878 16242 2345 | 5352 | 0.008 | -0.0453 | No | ||
| 54 | DAZL | 22944 | 5474 | 0.008 | -0.0516 | No | ||
| 55 | PHF6 | 2648 24340 | 5581 | 0.008 | -0.0570 | No | ||
| 56 | GOLGA3 | 6535 | 5582 | 0.008 | -0.0567 | No | ||
| 57 | ABCA3 | 6581 | 5697 | 0.007 | -0.0626 | No | ||
| 58 | NFATC3 | 5169 | 5702 | 0.007 | -0.0625 | No | ||
| 59 | UBE2B | 10245 | 5759 | 0.007 | -0.0653 | No | ||
| 60 | CDH22 | 14340 | 5784 | 0.007 | -0.0663 | No | ||
| 61 | GLULD1 | 14285 | 5909 | 0.006 | -0.0727 | No | ||
| 62 | TNFSF15 | 15863 | 6172 | 0.006 | -0.0867 | No | ||
| 63 | NPPB | 15994 | 6533 | 0.005 | -0.1061 | No | ||
| 64 | SLC25A27 | 7880 13149 | 6606 | 0.005 | -0.1098 | No | ||
| 65 | LRRC1 | 19059 3129 | 6611 | 0.005 | -0.1099 | No | ||
| 66 | WNT3 | 20635 | 6653 | 0.004 | -0.1119 | No | ||
| 67 | MYOHD1 | 20724 | 6667 | 0.004 | -0.1124 | No | ||
| 68 | NUP153 | 21474 | 6707 | 0.004 | -0.1144 | No | ||
| 69 | ELP4 | 8094 | 6709 | 0.004 | -0.1142 | No | ||
| 70 | PCDH8 | 21739 3017 | 6765 | 0.004 | -0.1170 | No | ||
| 71 | SYNCRIP | 3078 3035 3107 | 6835 | 0.004 | -0.1206 | No | ||
| 72 | FLRT1 | 11852 | 6987 | 0.004 | -0.1287 | No | ||
| 73 | GABRA1 | 20492 | 7028 | 0.004 | -0.1307 | No | ||
| 74 | PIK3R1 | 3170 | 7286 | 0.003 | -0.1445 | No | ||
| 75 | MYOD1 | 18237 | 7459 | 0.003 | -0.1538 | No | ||
| 76 | JAG1 | 14415 | 7963 | 0.002 | -0.1810 | No | ||
| 77 | LGI1 | 23867 | 7973 | 0.002 | -0.1814 | No | ||
| 78 | CEPT1 | 13624 | 8169 | 0.001 | -0.1920 | No | ||
| 79 | RAB2B | 21842 | 8307 | 0.001 | -0.1994 | No | ||
| 80 | MTA2 | 23934 18250 | 8308 | 0.001 | -0.1993 | No | ||
| 81 | SERPINA10 | 10151 | 8482 | 0.001 | -0.2087 | No | ||
| 82 | CNIH | 4536 23971 | 8715 | 0.000 | -0.2213 | No | ||
| 83 | NODAL | 333 20010 | 8777 | 0.000 | -0.2246 | No | ||
| 84 | SEMA3A | 16919 | 8963 | 0.000 | -0.2346 | No | ||
| 85 | RPS6KA3 | 8490 | 8979 | -0.000 | -0.2354 | No | ||
| 86 | PPP1R7 | 14176 | 9013 | -0.000 | -0.2372 | No | ||
| 87 | MAP3K6 | 16054 | 9123 | -0.000 | -0.2431 | No | ||
| 88 | PGM1 | 16841 | 9175 | -0.000 | -0.2459 | No | ||
| 89 | PRPH | 5294 | 9184 | -0.000 | -0.2463 | No | ||
| 90 | KCNH2 | 16592 | 9241 | -0.000 | -0.2493 | No | ||
| 91 | P518 | 14624 | 9289 | -0.001 | -0.2519 | No | ||
| 92 | PTHLH | 16931 | 9355 | -0.001 | -0.2554 | No | ||
| 93 | IL2 | 15354 | 9362 | -0.001 | -0.2557 | No | ||
| 94 | PIGW | 20312 | 9403 | -0.001 | -0.2578 | No | ||
| 95 | SPA17 | 19175 | 9542 | -0.001 | -0.2653 | No | ||
| 96 | JARID2 | 9200 | 9651 | -0.001 | -0.2711 | No | ||
| 97 | GJB6 | 21805 | 9674 | -0.001 | -0.2722 | No | ||
| 98 | ATP6V0D2 | 15934 | 9711 | -0.001 | -0.2742 | No | ||
| 99 | HSPB1 | 4879 | 9739 | -0.001 | -0.2756 | No | ||
| 100 | ARHGEF12 | 19156 | 9754 | -0.001 | -0.2763 | No | ||
| 101 | ACRV1 | 19521 | 9866 | -0.001 | -0.2823 | No | ||
| 102 | ARHGAP6 | 24207 | 10038 | -0.002 | -0.2915 | No | ||
| 103 | KLHL10 | 20666 | 10121 | -0.002 | -0.2958 | No | ||
| 104 | TWIST2 | 14185 | 10313 | -0.002 | -0.3061 | No | ||
| 105 | EIF5B | 10391 5963 | 10350 | -0.002 | -0.3080 | No | ||
| 106 | SEMA6A | 23439 9802 | 10413 | -0.002 | -0.3113 | No | ||
| 107 | CYP39A1 | 1538 23232 | 10461 | -0.002 | -0.3137 | No | ||
| 108 | TNRC4 | 15514 | 10496 | -0.003 | -0.3155 | No | ||
| 109 | RIMS2 | 4370 | 10503 | -0.003 | -0.3157 | No | ||
| 110 | PPM2C | 491 | 10534 | -0.003 | -0.3172 | No | ||
| 111 | EGR2 | 8886 | 10551 | -0.003 | -0.3180 | No | ||
| 112 | DNAJB4 | 12451 1916 1832 | 10586 | -0.003 | -0.3197 | No | ||
| 113 | PLAT | 18652 | 10787 | -0.003 | -0.3304 | No | ||
| 114 | GABRB3 | 4747 | 10791 | -0.003 | -0.3305 | No | ||
| 115 | GRB10 | 4799 | 10908 | -0.003 | -0.3366 | No | ||
| 116 | IER5 | 13802 | 11026 | -0.004 | -0.3428 | No | ||
| 117 | HOXA3 | 9108 1075 | 11197 | -0.004 | -0.3519 | No | ||
| 118 | RPS18 | 1529 5397 1603 | 11243 | -0.004 | -0.3542 | No | ||
| 119 | ADAMTS1 | 1755 4347 | 11257 | -0.004 | -0.3547 | No | ||
| 120 | PAFAH1B2 | 5221 9525 | 11266 | -0.004 | -0.3550 | No | ||
| 121 | SIX3 | 5444 | 11414 | -0.004 | -0.3628 | No | ||
| 122 | NNAT | 2764 5180 | 11602 | -0.005 | -0.3728 | No | ||
| 123 | MGAT3 | 22418 | 11647 | -0.005 | -0.3750 | No | ||
| 124 | ADRA2C | 16869 | 11751 | -0.005 | -0.3803 | No | ||
| 125 | LMOD1 | 14119 | 11770 | -0.005 | -0.3811 | No | ||
| 126 | PDGFB | 22199 2311 | 11888 | -0.005 | -0.3872 | No | ||
| 127 | GLRA2 | 24008 | 11891 | -0.005 | -0.3871 | No | ||
| 128 | CDK5R2 | 14220 | 11932 | -0.006 | -0.3891 | No | ||
| 129 | MYT1 | 36 | 12071 | -0.006 | -0.3963 | No | ||
| 130 | LGALS8 | 21544 | 12224 | -0.006 | -0.4043 | No | ||
| 131 | SCGB1A1 | 23750 | 12458 | -0.007 | -0.4167 | No | ||
| 132 | KCNAB1 | 1791 15577 | 12491 | -0.007 | -0.4181 | No | ||
| 133 | SLC2A12 | 20073 | 12537 | -0.007 | -0.4203 | No | ||
| 134 | CYP46A1 | 21158 | 12773 | -0.008 | -0.4327 | No | ||
| 135 | GPC4 | 24153 | 13037 | -0.009 | -0.4467 | No | ||
| 136 | KRT20 | 12410 | 13181 | -0.009 | -0.4540 | No | ||
| 137 | DMRTB1 | 15813 | 13199 | -0.009 | -0.4546 | No | ||
| 138 | KCNQ1 | 18001 4948 | 13210 | -0.009 | -0.4547 | No | ||
| 139 | RTN1 | 4177 2099 | 13237 | -0.009 | -0.4558 | No | ||
| 140 | SPACA3 | 13680 | 13274 | -0.010 | -0.4573 | No | ||
| 141 | CNTNAP2 | 17463 12406 | 13330 | -0.010 | -0.4599 | No | ||
| 142 | STAMBP | 17097 | 13392 | -0.010 | -0.4628 | No | ||
| 143 | FLNC | 4302 8510 | 13455 | -0.010 | -0.4657 | No | ||
| 144 | EMX2 | 23806 | 13605 | -0.011 | -0.4733 | No | ||
| 145 | SLC35C1 | 14517 | 13644 | -0.011 | -0.4749 | No | ||
| 146 | FGF16 | 24270 | 13678 | -0.012 | -0.4762 | No | ||
| 147 | ABL1 | 2693 4301 2794 | 13802 | -0.012 | -0.4824 | No | ||
| 148 | ACTA1 | 18715 | 13818 | -0.013 | -0.4827 | No | ||
| 149 | GPR22 | 21087 | 14036 | -0.014 | -0.4939 | No | ||
| 150 | GCNT2 | 9168 3166 3243 21661 3270 | 14068 | -0.014 | -0.4950 | No | ||
| 151 | GABRB1 | 16832 | 14217 | -0.015 | -0.5024 | No | ||
| 152 | HSPB2 | 19121 | 14278 | -0.016 | -0.5050 | No | ||
| 153 | NUP98 | 17726 | 14434 | -0.017 | -0.5127 | No | ||
| 154 | AGTR2 | 24365 | 14611 | -0.018 | -0.5215 | No | ||
| 155 | GRIN2D | 9042 | 14626 | -0.019 | -0.5215 | No | ||
| 156 | HNRPA2B1 | 1187 7001 7000 1040 | 14647 | -0.019 | -0.5218 | No | ||
| 157 | GAB2 | 1821 18184 2025 | 14705 | -0.019 | -0.5241 | No | ||
| 158 | NTF5 | 18248 | 15031 | -0.023 | -0.5408 | No | ||
| 159 | E2F1 | 14384 | 15124 | -0.025 | -0.5448 | No | ||
| 160 | IDH1 | 4894 | 15142 | -0.025 | -0.5447 | No | ||
| 161 | NPY2R | 343 | 15341 | -0.028 | -0.5542 | No | ||
| 162 | RECQL | 16942 935 9713 | 15442 | -0.031 | -0.5584 | No | ||
| 163 | SAG | 5404 | 15484 | -0.032 | -0.5593 | No | ||
| 164 | TNMD | 24260 | 15488 | -0.032 | -0.5581 | No | ||
| 165 | PHF15 | 20467 8050 | 15517 | -0.032 | -0.5583 | No | ||
| 166 | NPR1 | 9480 | 15690 | -0.037 | -0.5661 | No | ||
| 167 | CYR61 | 9159 15145 9311 5024 | 15858 | -0.042 | -0.5735 | No | ||
| 168 | YWHAG | 16339 | 16001 | -0.047 | -0.5792 | No | ||
| 169 | TXNDC9 | 13972 | 16002 | -0.047 | -0.5773 | No | ||
| 170 | TLK1 | 14563 | 16181 | -0.054 | -0.5847 | No | ||
| 171 | SOD2 | 23379 | 16206 | -0.056 | -0.5836 | No | ||
| 172 | MAT2A | 10553 10554 6099 | 16236 | -0.057 | -0.5828 | No | ||
| 173 | ETV1 | 4688 8925 8924 | 16353 | -0.062 | -0.5865 | No | ||
| 174 | DNAJB5 | 16236 | 16388 | -0.064 | -0.5857 | No | ||
| 175 | MTPN | 17183 | 16595 | -0.075 | -0.5938 | No | ||
| 176 | WBP2 | 20146 | 16723 | -0.084 | -0.5972 | No | ||
| 177 | STIP1 | 23798 | 16824 | -0.091 | -0.5988 | No | ||
| 178 | CAMK2B | 20536 | 16913 | -0.097 | -0.5995 | Yes | ||
| 179 | PLSCR4 | 19353 | 16921 | -0.097 | -0.5958 | Yes | ||
| 180 | IL21R | 1398 18084 | 16931 | -0.098 | -0.5922 | Yes | ||
| 181 | SERPINH1 | 8703 | 16934 | -0.098 | -0.5883 | Yes | ||
| 182 | SLC4A2 | 9828 | 16987 | -0.103 | -0.5868 | Yes | ||
| 183 | TPP2 | 14257 | 17108 | -0.113 | -0.5886 | Yes | ||
| 184 | ZDHHC14 | 5886 | 17281 | -0.131 | -0.5925 | Yes | ||
| 185 | ACVR1 | 4334 | 17342 | -0.138 | -0.5899 | Yes | ||
| 186 | SRPK2 | 5513 | 17366 | -0.141 | -0.5853 | Yes | ||
| 187 | CCT8 | 22551 | 17441 | -0.147 | -0.5832 | Yes | ||
| 188 | MAP1LC3B | 7443 | 17493 | -0.152 | -0.5796 | Yes | ||
| 189 | MORF4L2 | 12118 | 17514 | -0.156 | -0.5742 | Yes | ||
| 190 | EFHD1 | 8267 | 17562 | -0.162 | -0.5699 | Yes | ||
| 191 | PON2 | 11637 | 17694 | -0.179 | -0.5695 | Yes | ||
| 192 | CCT4 | 8710 | 17774 | -0.191 | -0.5659 | Yes | ||
| 193 | PPID | 12576 | 17788 | -0.193 | -0.5585 | Yes | ||
| 194 | HSPD1 | 4078 9129 | 17794 | -0.194 | -0.5507 | Yes | ||
| 195 | TOP1 | 5790 5789 10210 | 17800 | -0.195 | -0.5428 | Yes | ||
| 196 | CHORDC1 | 12430 | 17908 | -0.215 | -0.5396 | Yes | ||
| 197 | HAT1 | 14990 | 17910 | -0.215 | -0.5307 | Yes | ||
| 198 | VCP | 11254 | 17956 | -0.227 | -0.5236 | Yes | ||
| 199 | ATP5I | 8637 | 17997 | -0.239 | -0.5158 | Yes | ||
| 200 | CAP1 | 8684 | 18104 | -0.276 | -0.5101 | Yes | ||
| 201 | CRYAB | 19457 | 18139 | -0.286 | -0.5000 | Yes | ||
| 202 | YWHAQ | 10369 | 18214 | -0.327 | -0.4903 | Yes | ||
| 203 | APLP2 | 19190 | 18220 | -0.329 | -0.4769 | Yes | ||
| 204 | NUDC | 9495 | 18275 | -0.372 | -0.4643 | Yes | ||
| 205 | PAPOLA | 5261 9585 | 18279 | -0.376 | -0.4487 | Yes | ||
| 206 | ST13 | 12865 | 18297 | -0.389 | -0.4334 | Yes | ||
| 207 | UCHL1 | 16834 | 18321 | -0.409 | -0.4176 | Yes | ||
| 208 | CDKN1A | 4511 8729 | 18339 | -0.430 | -0.4005 | Yes | ||
| 209 | UTX | 10266 2574 | 18399 | -0.516 | -0.3822 | Yes | ||
| 210 | HSPH1 | 16282 | 18415 | -0.556 | -0.3597 | Yes | ||
| 211 | RAB18 | 1972 5342 | 18484 | -0.835 | -0.3286 | Yes | ||
| 212 | HSPA8 | 9124 3072 4871 | 18522 | -1.208 | -0.2801 | Yes | ||
| 213 | INVS | 2397 16211 | 18560 | -1.443 | -0.2218 | Yes | ||
| 214 | SFRS10 | 9818 1722 5441 | 18600 | -2.513 | -0.1189 | Yes | ||
| 215 | GAS7 | 20836 1299 | 18604 | -2.865 | 0.0007 | Yes |

