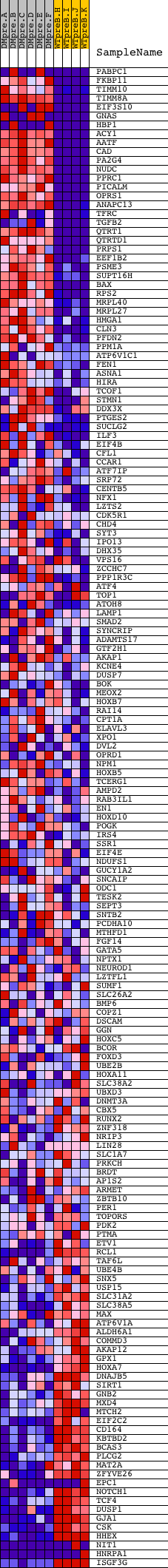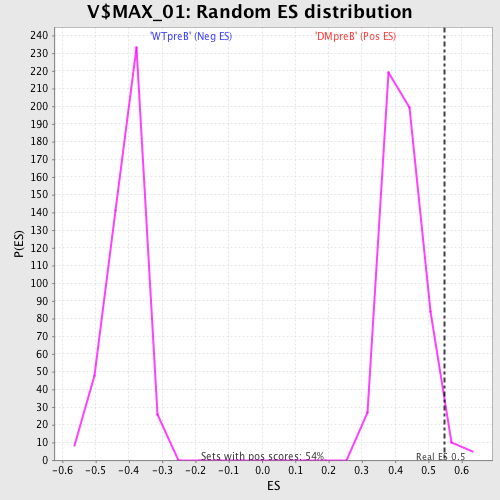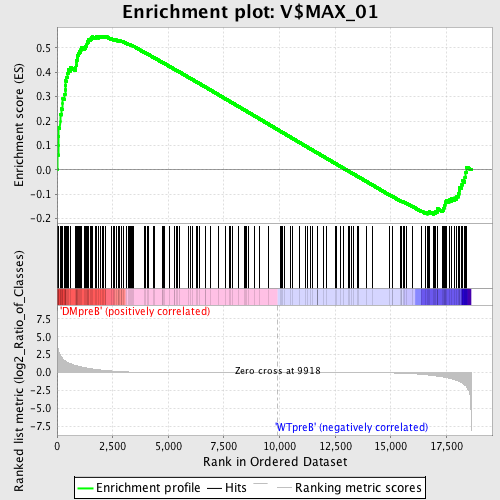
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_DMpreB_versus_WTpreB.phenotype_DMpreB_versus_WTpreB.cls #DMpreB_versus_WTpreB |
| Phenotype | phenotype_DMpreB_versus_WTpreB.cls#DMpreB_versus_WTpreB |
| Upregulated in class | DMpreB |
| GeneSet | V$MAX_01 |
| Enrichment Score (ES) | 0.5481195 |
| Normalized Enrichment Score (NES) | 1.2944953 |
| Nominal p-value | 0.02389706 |
| FDR q-value | 0.69204617 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PABPC1 | 5219 9522 9523 23572 | 7 | 5.273 | 0.0625 | Yes | ||
| 2 | FKBP11 | 22136 | 46 | 3.327 | 0.1001 | Yes | ||
| 3 | TIMM10 | 11390 2158 | 50 | 3.231 | 0.1385 | Yes | ||
| 4 | TIMM8A | 24062 | 72 | 2.883 | 0.1718 | Yes | ||
| 5 | EIF3S10 | 4659 8887 | 129 | 2.520 | 0.1988 | Yes | ||
| 6 | GNAS | 9025 2963 2752 | 131 | 2.517 | 0.2288 | Yes | ||
| 7 | HBP1 | 2048 2162 21086 | 207 | 2.101 | 0.2498 | Yes | ||
| 8 | ACY1 | 19012 2996 | 255 | 1.924 | 0.2702 | Yes | ||
| 9 | AATF | 20315 | 258 | 1.910 | 0.2928 | Yes | ||
| 10 | CAD | 16886 | 321 | 1.716 | 0.3100 | Yes | ||
| 11 | PA2G4 | 5265 | 360 | 1.642 | 0.3275 | Yes | ||
| 12 | NUDC | 9495 | 368 | 1.624 | 0.3465 | Yes | ||
| 13 | PPRC1 | 23808 | 379 | 1.599 | 0.3650 | Yes | ||
| 14 | PICALM | 18191 | 434 | 1.477 | 0.3797 | Yes | ||
| 15 | OPRS1 | 9515 2402 | 458 | 1.437 | 0.3956 | Yes | ||
| 16 | ANAPC13 | 12761 | 497 | 1.383 | 0.4100 | Yes | ||
| 17 | TFRC | 22785 | 595 | 1.245 | 0.4196 | Yes | ||
| 18 | TGFB2 | 5743 | 839 | 0.982 | 0.4182 | Yes | ||
| 19 | QTRT1 | 19540 | 849 | 0.970 | 0.4293 | Yes | ||
| 20 | QTRTD1 | 22590 | 873 | 0.942 | 0.4392 | Yes | ||
| 21 | PRPS1 | 24233 | 878 | 0.937 | 0.4502 | Yes | ||
| 22 | EEF1B2 | 4131 12063 | 901 | 0.924 | 0.4600 | Yes | ||
| 23 | PSME3 | 20657 | 912 | 0.917 | 0.4704 | Yes | ||
| 24 | SUPT16H | 8541 | 974 | 0.861 | 0.4774 | Yes | ||
| 25 | BAX | 17832 | 1000 | 0.840 | 0.4861 | Yes | ||
| 26 | RPS2 | 9279 | 1069 | 0.794 | 0.4919 | Yes | ||
| 27 | MRPL40 | 22641 | 1084 | 0.785 | 0.5005 | Yes | ||
| 28 | MRPL27 | 13573 | 1223 | 0.700 | 0.5013 | Yes | ||
| 29 | HMGA1 | 23323 | 1282 | 0.655 | 0.5060 | Yes | ||
| 30 | CLN3 | 17635 | 1313 | 0.637 | 0.5120 | Yes | ||
| 31 | PFDN2 | 9551 | 1343 | 0.621 | 0.5178 | Yes | ||
| 32 | PPM1A | 5279 9607 | 1352 | 0.616 | 0.5247 | Yes | ||
| 33 | ATP6V1C1 | 22487 12329 | 1403 | 0.586 | 0.5290 | Yes | ||
| 34 | FEN1 | 8961 | 1416 | 0.580 | 0.5353 | Yes | ||
| 35 | ASNA1 | 18810 | 1514 | 0.538 | 0.5364 | Yes | ||
| 36 | HIRA | 4852 9090 | 1541 | 0.528 | 0.5413 | Yes | ||
| 37 | TCOF1 | 5660 | 1573 | 0.513 | 0.5457 | Yes | ||
| 38 | STMN1 | 9261 | 1739 | 0.442 | 0.5421 | Yes | ||
| 39 | DDX3X | 24379 | 1756 | 0.436 | 0.5464 | Yes | ||
| 40 | PTGES2 | 15038 | 1874 | 0.392 | 0.5447 | Yes | ||
| 41 | SUCLG2 | 17062 5545 | 1933 | 0.374 | 0.5461 | Yes | ||
| 42 | ILF3 | 3110 3030 9176 | 2021 | 0.347 | 0.5455 | Yes | ||
| 43 | EIF4B | 13279 7979 | 2068 | 0.332 | 0.5469 | Yes | ||
| 44 | CFL1 | 4516 | 2158 | 0.303 | 0.5457 | Yes | ||
| 45 | CCAR1 | 7454 | 2180 | 0.296 | 0.5481 | Yes | ||
| 46 | ATF7IP | 12028 13038 | 2434 | 0.228 | 0.5371 | No | ||
| 47 | SRP72 | 16819 | 2533 | 0.205 | 0.5343 | No | ||
| 48 | CENTB5 | 15961 | 2560 | 0.198 | 0.5352 | No | ||
| 49 | NFX1 | 2475 | 2650 | 0.179 | 0.5325 | No | ||
| 50 | LZTS2 | 10362 | 2743 | 0.156 | 0.5294 | No | ||
| 51 | CDK5R1 | 20745 | 2758 | 0.153 | 0.5305 | No | ||
| 52 | CHD4 | 8418 17281 4225 | 2821 | 0.139 | 0.5288 | No | ||
| 53 | SYT3 | 18264 | 2886 | 0.130 | 0.5269 | No | ||
| 54 | IPO13 | 15784 | 2900 | 0.126 | 0.5277 | No | ||
| 55 | DHX35 | 14754 | 2967 | 0.115 | 0.5255 | No | ||
| 56 | VPS16 | 2927 14845 2825 | 3129 | 0.093 | 0.5178 | No | ||
| 57 | ZCCHC7 | 16225 | 3192 | 0.086 | 0.5155 | No | ||
| 58 | PPP1R3C | 23688 | 3231 | 0.083 | 0.5144 | No | ||
| 59 | ATF4 | 22417 2239 | 3287 | 0.078 | 0.5124 | No | ||
| 60 | TOP1 | 5790 5789 10210 | 3339 | 0.073 | 0.5105 | No | ||
| 61 | ATOH8 | 17120 | 3380 | 0.070 | 0.5092 | No | ||
| 62 | LAMP1 | 18679 | 3425 | 0.067 | 0.5076 | No | ||
| 63 | SMAD2 | 23511 | 3937 | 0.043 | 0.4804 | No | ||
| 64 | SYNCRIP | 3078 3035 3107 | 3969 | 0.042 | 0.4792 | No | ||
| 65 | ADAMTS17 | 598 | 4079 | 0.038 | 0.4737 | No | ||
| 66 | GTF2H1 | 4069 18236 | 4104 | 0.038 | 0.4729 | No | ||
| 67 | AKAP1 | 4364 1326 1286 20299 | 4322 | 0.032 | 0.4615 | No | ||
| 68 | KCNE4 | 12196 | 4366 | 0.031 | 0.4595 | No | ||
| 69 | DUSP7 | 10661 | 4741 | 0.024 | 0.4396 | No | ||
| 70 | BOK | 11953 14174 | 4745 | 0.024 | 0.4397 | No | ||
| 71 | MEOX2 | 21285 | 4769 | 0.024 | 0.4387 | No | ||
| 72 | HOXB7 | 666 9110 | 4847 | 0.023 | 0.4348 | No | ||
| 73 | RAI14 | 22327 2191 | 5060 | 0.020 | 0.4236 | No | ||
| 74 | CPT1A | 23756 | 5276 | 0.018 | 0.4121 | No | ||
| 75 | ELAVL3 | 9136 | 5292 | 0.018 | 0.4115 | No | ||
| 76 | XPO1 | 4172 | 5368 | 0.017 | 0.4077 | No | ||
| 77 | DVL2 | 20813 | 5423 | 0.017 | 0.4049 | No | ||
| 78 | OPRD1 | 15737 | 5479 | 0.016 | 0.4022 | No | ||
| 79 | NPM1 | 1196 | 5915 | 0.013 | 0.3787 | No | ||
| 80 | HOXB5 | 20687 | 5996 | 0.012 | 0.3745 | No | ||
| 81 | TCERG1 | 12075 2027 7068 | 6078 | 0.012 | 0.3703 | No | ||
| 82 | AMPD2 | 1931 15197 | 6243 | 0.011 | 0.3615 | No | ||
| 83 | RAB3IL1 | 3723 13229 | 6292 | 0.011 | 0.3591 | No | ||
| 84 | EN1 | 387 14155 | 6409 | 0.010 | 0.3529 | No | ||
| 85 | HOXD10 | 14983 | 6660 | 0.009 | 0.3395 | No | ||
| 86 | POGK | 7739 | 6677 | 0.009 | 0.3387 | No | ||
| 87 | IRS4 | 9183 4926 | 6909 | 0.008 | 0.3263 | No | ||
| 88 | SSR1 | 4211 | 7248 | 0.007 | 0.3080 | No | ||
| 89 | EIF4E | 15403 1827 8890 | 7585 | 0.006 | 0.2899 | No | ||
| 90 | NDUFS1 | 13940 | 7764 | 0.005 | 0.2803 | No | ||
| 91 | GUCY1A2 | 19580 | 7790 | 0.005 | 0.2790 | No | ||
| 92 | SNCAIP | 12589 | 7871 | 0.005 | 0.2747 | No | ||
| 93 | ODC1 | 9500 | 8133 | 0.004 | 0.2606 | No | ||
| 94 | TESK2 | 10504 | 8420 | 0.004 | 0.2452 | No | ||
| 95 | SEPT3 | 6275 10775 | 8485 | 0.003 | 0.2418 | No | ||
| 96 | SNTB2 | 3887 18476 | 8521 | 0.003 | 0.2399 | No | ||
| 97 | PCDHA10 | 8792 | 8612 | 0.003 | 0.2351 | No | ||
| 98 | MTHFD1 | 2132 21238 | 8890 | 0.002 | 0.2201 | No | ||
| 99 | FGF14 | 21716 3005 3463 | 9074 | 0.002 | 0.2102 | No | ||
| 100 | GATA5 | 14315 | 9491 | 0.001 | 0.1876 | No | ||
| 101 | NPTX1 | 20126 | 10028 | -0.000 | 0.1586 | No | ||
| 102 | NEUROD1 | 14550 | 10083 | -0.000 | 0.1557 | No | ||
| 103 | LZTFL1 | 8210 | 10116 | -0.000 | 0.1539 | No | ||
| 104 | SUMF1 | 17049 | 10228 | -0.001 | 0.1479 | No | ||
| 105 | SLC26A2 | 23427 | 10489 | -0.001 | 0.1338 | No | ||
| 106 | BMP6 | 8659 | 10582 | -0.002 | 0.1289 | No | ||
| 107 | COPZ1 | 22338 | 10878 | -0.002 | 0.1129 | No | ||
| 108 | DSCAM | 1667 4641 22530 | 11144 | -0.003 | 0.0986 | No | ||
| 109 | GGN | 2472 1171 18312 | 11258 | -0.003 | 0.0925 | No | ||
| 110 | HOXC5 | 22340 | 11403 | -0.004 | 0.0847 | No | ||
| 111 | BCOR | 12934 2561 | 11461 | -0.004 | 0.0817 | No | ||
| 112 | FOXD3 | 9088 | 11682 | -0.005 | 0.0698 | No | ||
| 113 | UBE2B | 10245 | 11683 | -0.005 | 0.0699 | No | ||
| 114 | HOXA11 | 17146 | 11691 | -0.005 | 0.0696 | No | ||
| 115 | SLC38A2 | 22149 2219 | 11720 | -0.005 | 0.0681 | No | ||
| 116 | UBXD3 | 15701 | 11966 | -0.005 | 0.0549 | No | ||
| 117 | DNMT3A | 2167 21330 | 12095 | -0.006 | 0.0480 | No | ||
| 118 | CBX5 | 22108 4485 | 12504 | -0.008 | 0.0260 | No | ||
| 119 | RUNX2 | 4480 8700 | 12563 | -0.008 | 0.0229 | No | ||
| 120 | ZNF318 | 12198 7179 | 12747 | -0.009 | 0.0131 | No | ||
| 121 | NRIP3 | 17679 | 12891 | -0.010 | 0.0055 | No | ||
| 122 | LIN28 | 15723 | 13077 | -0.011 | -0.0044 | No | ||
| 123 | SLC1A7 | 16147 | 13126 | -0.011 | -0.0069 | No | ||
| 124 | PRKCH | 21246 | 13221 | -0.012 | -0.0119 | No | ||
| 125 | BRDT | 4327 | 13337 | -0.012 | -0.0179 | No | ||
| 126 | AP1S2 | 8423 4228 | 13483 | -0.013 | -0.0256 | No | ||
| 127 | ARMET | 19005 | 13552 | -0.014 | -0.0292 | No | ||
| 128 | ZBTB10 | 10468 | 13886 | -0.018 | -0.0470 | No | ||
| 129 | PER1 | 20827 | 14179 | -0.022 | -0.0626 | No | ||
| 130 | TOPORS | 15920 | 14955 | -0.044 | -0.1040 | No | ||
| 131 | PDK2 | 20285 | 15054 | -0.048 | -0.1088 | No | ||
| 132 | PTMA | 9657 | 15085 | -0.050 | -0.1098 | No | ||
| 133 | ETV1 | 4688 8925 8924 | 15435 | -0.071 | -0.1279 | No | ||
| 134 | RCL1 | 3728 3743 23894 | 15487 | -0.076 | -0.1297 | No | ||
| 135 | TAF6L | 5923 | 15498 | -0.077 | -0.1294 | No | ||
| 136 | UBE4B | 2433 15672 | 15556 | -0.083 | -0.1315 | No | ||
| 137 | SNX5 | 12783 14410 | 15624 | -0.089 | -0.1340 | No | ||
| 138 | USP15 | 4758 3377 | 15712 | -0.099 | -0.1376 | No | ||
| 139 | SLC31A2 | 9827 | 15970 | -0.141 | -0.1498 | No | ||
| 140 | SLC38A5 | 24388 | 16386 | -0.239 | -0.1695 | No | ||
| 141 | MAX | 21034 | 16577 | -0.301 | -0.1762 | No | ||
| 142 | ATP6V1A | 8638 | 16670 | -0.331 | -0.1772 | No | ||
| 143 | ALDH6A1 | 21025 | 16695 | -0.338 | -0.1745 | No | ||
| 144 | COMMD3 | 73 | 16724 | -0.346 | -0.1719 | No | ||
| 145 | AKAP12 | 17600 | 16932 | -0.417 | -0.1781 | No | ||
| 146 | GPX1 | 19310 | 16950 | -0.425 | -0.1740 | No | ||
| 147 | HOXA7 | 4862 | 17020 | -0.453 | -0.1723 | No | ||
| 148 | DNAJB5 | 16236 | 17082 | -0.478 | -0.1699 | No | ||
| 149 | SIRT1 | 19745 | 17091 | -0.483 | -0.1646 | No | ||
| 150 | GNB2 | 9026 | 17106 | -0.490 | -0.1595 | No | ||
| 151 | MXD4 | 9346 | 17337 | -0.594 | -0.1649 | No | ||
| 152 | MTCH2 | 14954 7120 | 17366 | -0.612 | -0.1591 | No | ||
| 153 | EIF2C2 | 6247 | 17393 | -0.631 | -0.1530 | No | ||
| 154 | CD164 | 20046 | 17425 | -0.654 | -0.1468 | No | ||
| 155 | KBTBD2 | 17137 | 17437 | -0.664 | -0.1395 | No | ||
| 156 | BCAS3 | 5314 | 17440 | -0.668 | -0.1317 | No | ||
| 157 | PLCG2 | 18453 | 17511 | -0.711 | -0.1270 | No | ||
| 158 | MAT2A | 10553 10554 6099 | 17615 | -0.768 | -0.1234 | No | ||
| 159 | ZFYVE26 | 9999 | 17714 | -0.848 | -0.1186 | No | ||
| 160 | EPC1 | 2039 8907 2033 | 17873 | -0.981 | -0.1155 | No | ||
| 161 | NOTCH1 | 14649 | 17966 | -1.076 | -0.1076 | No | ||
| 162 | TCF4 | 5642 | 18039 | -1.160 | -0.0977 | No | ||
| 163 | DUSP1 | 23061 | 18067 | -1.196 | -0.0849 | No | ||
| 164 | GJA1 | 20023 | 18097 | -1.227 | -0.0718 | No | ||
| 165 | CSK | 8805 | 18193 | -1.378 | -0.0605 | No | ||
| 166 | HHEX | 23872 | 18228 | -1.455 | -0.0450 | No | ||
| 167 | NIT1 | 11293 933 | 18330 | -1.725 | -0.0299 | No | ||
| 168 | HNRPA1 | 9103 4860 | 18370 | -1.836 | -0.0101 | No | ||
| 169 | ISGF3G | 2686 9184 | 18410 | -1.961 | 0.0112 | No |