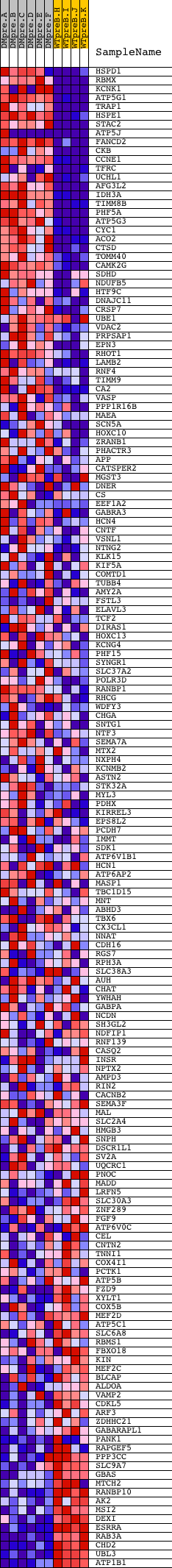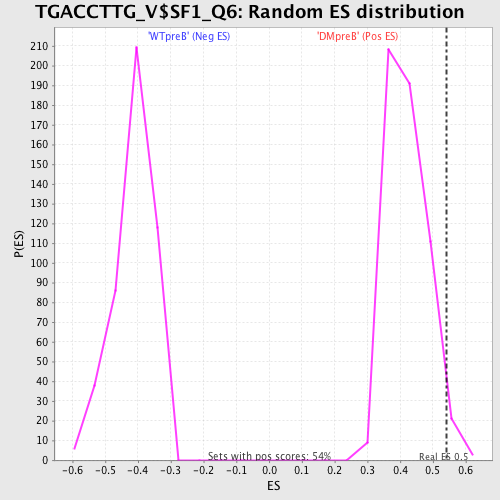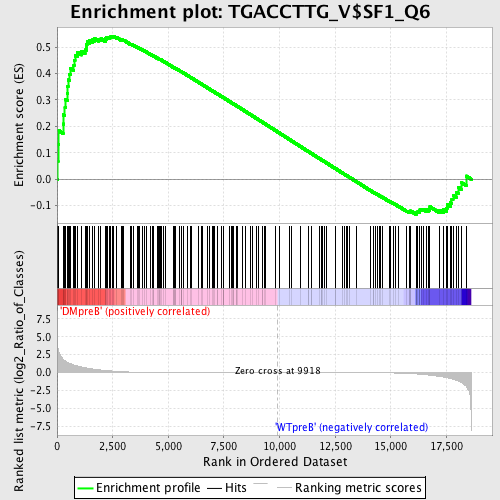
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_DMpreB_versus_WTpreB.phenotype_DMpreB_versus_WTpreB.cls #DMpreB_versus_WTpreB |
| Phenotype | phenotype_DMpreB_versus_WTpreB.cls#DMpreB_versus_WTpreB |
| Upregulated in class | DMpreB |
| GeneSet | TGACCTTG_V$SF1_Q6 |
| Enrichment Score (ES) | 0.54174685 |
| Normalized Enrichment Score (NES) | 1.2893597 |
| Nominal p-value | 0.023941068 |
| FDR q-value | 0.6363295 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | HSPD1 | 4078 9129 | 35 | 3.635 | 0.0680 | Yes | ||
| 2 | RBMX | 5367 9708 | 42 | 3.376 | 0.1325 | Yes | ||
| 3 | KCNK1 | 18415 | 82 | 2.753 | 0.1833 | Yes | ||
| 4 | ATP5G1 | 8636 | 270 | 1.873 | 0.2092 | Yes | ||
| 5 | TRAP1 | 22683 | 282 | 1.845 | 0.2441 | Yes | ||
| 6 | HSPE1 | 9133 | 353 | 1.656 | 0.2721 | Yes | ||
| 7 | STAC2 | 20265 | 386 | 1.572 | 0.3006 | Yes | ||
| 8 | ATP5J | 880 22555 | 462 | 1.432 | 0.3240 | Yes | ||
| 9 | FANCD2 | 17326 464 | 478 | 1.404 | 0.3502 | Yes | ||
| 10 | CKB | 4523 | 495 | 1.384 | 0.3759 | Yes | ||
| 11 | CCNE1 | 17857 | 561 | 1.308 | 0.3975 | Yes | ||
| 12 | TFRC | 22785 | 595 | 1.245 | 0.4197 | Yes | ||
| 13 | UCHL1 | 16834 | 751 | 1.057 | 0.4316 | Yes | ||
| 14 | AFG3L2 | 23416 | 766 | 1.044 | 0.4509 | Yes | ||
| 15 | IDH3A | 19447 | 806 | 1.016 | 0.4683 | Yes | ||
| 16 | TIMM8B | 11391 | 907 | 0.918 | 0.4806 | Yes | ||
| 17 | PHF5A | 22194 | 1098 | 0.775 | 0.4852 | Yes | ||
| 18 | ATP5G3 | 14558 | 1257 | 0.672 | 0.4895 | Yes | ||
| 19 | CYC1 | 12348 | 1309 | 0.639 | 0.4990 | Yes | ||
| 20 | ACO2 | 8527 | 1318 | 0.634 | 0.5108 | Yes | ||
| 21 | CTSD | 22 | 1349 | 0.617 | 0.5210 | Yes | ||
| 22 | TOMM40 | 6997 | 1461 | 0.559 | 0.5258 | Yes | ||
| 23 | CAMK2G | 21905 | 1605 | 0.498 | 0.5276 | Yes | ||
| 24 | SDHD | 19125 | 1692 | 0.461 | 0.5318 | Yes | ||
| 25 | NDUFB5 | 12277 | 1859 | 0.398 | 0.5304 | Yes | ||
| 26 | HTF9C | 4886 1678 | 1971 | 0.362 | 0.5314 | Yes | ||
| 27 | DNAJC11 | 2486 6068 | 2166 | 0.301 | 0.5267 | Yes | ||
| 28 | CRSP7 | 18838 | 2168 | 0.300 | 0.5324 | Yes | ||
| 29 | UBE1 | 24370 2551 | 2211 | 0.288 | 0.5356 | Yes | ||
| 30 | VDAC2 | 22081 | 2265 | 0.270 | 0.5380 | Yes | ||
| 31 | PRPSAP1 | 20141 | 2370 | 0.245 | 0.5370 | Yes | ||
| 32 | EPN3 | 20291 | 2416 | 0.233 | 0.5391 | Yes | ||
| 33 | RHOT1 | 12227 | 2475 | 0.217 | 0.5401 | Yes | ||
| 34 | LAMB2 | 9265 | 2519 | 0.207 | 0.5417 | Yes | ||
| 35 | RNF4 | 5385 9729 | 2671 | 0.175 | 0.5369 | No | ||
| 36 | TIMM9 | 19686 | 2891 | 0.129 | 0.5275 | No | ||
| 37 | CA2 | 15631 1825 | 2932 | 0.122 | 0.5277 | No | ||
| 38 | VASP | 5847 | 2991 | 0.112 | 0.5267 | No | ||
| 39 | PPP1R16B | 6014 | 3308 | 0.077 | 0.5111 | No | ||
| 40 | MAEA | 16877 | 3332 | 0.074 | 0.5112 | No | ||
| 41 | SCN5A | 5412 | 3453 | 0.065 | 0.5060 | No | ||
| 42 | HOXC10 | 22342 | 3592 | 0.058 | 0.4996 | No | ||
| 43 | ZRANB1 | 6884 11702 | 3635 | 0.056 | 0.4984 | No | ||
| 44 | PHACTR3 | 7900 2770 | 3703 | 0.052 | 0.4958 | No | ||
| 45 | APP | 4402 | 3821 | 0.047 | 0.4903 | No | ||
| 46 | CATSPER2 | 10014 | 3911 | 0.043 | 0.4863 | No | ||
| 47 | MGST3 | 13771 12349 | 4003 | 0.040 | 0.4822 | No | ||
| 48 | DNER | 13901 | 4176 | 0.036 | 0.4735 | No | ||
| 49 | CS | 19839 | 4292 | 0.033 | 0.4679 | No | ||
| 50 | EEF1A2 | 8880 14309 | 4348 | 0.031 | 0.4656 | No | ||
| 51 | GABRA3 | 24139 | 4354 | 0.031 | 0.4659 | No | ||
| 52 | HCN4 | 9080 | 4491 | 0.028 | 0.4591 | No | ||
| 53 | CNTF | 3741 8761 3680 3736 | 4547 | 0.027 | 0.4566 | No | ||
| 54 | VSNL1 | 6527 | 4615 | 0.026 | 0.4535 | No | ||
| 55 | NTNG2 | 2731 9330 | 4620 | 0.026 | 0.4538 | No | ||
| 56 | KLK15 | 11404 | 4637 | 0.026 | 0.4534 | No | ||
| 57 | KIF5A | 9222 | 4677 | 0.025 | 0.4518 | No | ||
| 58 | COMTD1 | 21903 | 4797 | 0.023 | 0.4458 | No | ||
| 59 | TUBB4 | 22917 | 4860 | 0.023 | 0.4428 | No | ||
| 60 | AMY2A | 8587 | 5249 | 0.018 | 0.4222 | No | ||
| 61 | FSTL3 | 19961 | 5251 | 0.018 | 0.4225 | No | ||
| 62 | ELAVL3 | 9136 | 5292 | 0.018 | 0.4206 | No | ||
| 63 | TCF2 | 1395 20727 | 5297 | 0.018 | 0.4208 | No | ||
| 64 | DIRAS1 | 9915 | 5315 | 0.018 | 0.4202 | No | ||
| 65 | HOXC13 | 22344 | 5316 | 0.018 | 0.4205 | No | ||
| 66 | KCNG4 | 18728 3845 | 5507 | 0.016 | 0.4105 | No | ||
| 67 | PHF15 | 20467 8050 | 5576 | 0.015 | 0.4071 | No | ||
| 68 | SYNGR1 | 22420 2290 | 5581 | 0.015 | 0.4072 | No | ||
| 69 | SLC37A2 | 19178 | 5601 | 0.015 | 0.4065 | No | ||
| 70 | POLR3D | 21760 12456 | 5668 | 0.015 | 0.4032 | No | ||
| 71 | RANBP1 | 9692 5357 | 5860 | 0.013 | 0.3931 | No | ||
| 72 | RHCG | 17788 | 5995 | 0.012 | 0.3861 | No | ||
| 73 | WDFY3 | 7789 3607 16462 | 6049 | 0.012 | 0.3834 | No | ||
| 74 | CHGA | 21178 2151 | 6344 | 0.011 | 0.3677 | No | ||
| 75 | SNTG1 | 7717 | 6490 | 0.010 | 0.3600 | No | ||
| 76 | NTF3 | 16991 | 6533 | 0.010 | 0.3579 | No | ||
| 77 | SEMA7A | 19426 9803 | 6738 | 0.009 | 0.3470 | No | ||
| 78 | MTX2 | 14980 | 6865 | 0.008 | 0.3404 | No | ||
| 79 | NXPH4 | 19602 | 6972 | 0.008 | 0.3348 | No | ||
| 80 | KCNMB2 | 1939 15615 | 7028 | 0.008 | 0.3319 | No | ||
| 81 | ASTN2 | 15860 2524 | 7071 | 0.008 | 0.3298 | No | ||
| 82 | STK32A | 6509 | 7197 | 0.007 | 0.3232 | No | ||
| 83 | MYL3 | 19291 | 7219 | 0.007 | 0.3222 | No | ||
| 84 | PDHX | 14510 | 7375 | 0.007 | 0.3139 | No | ||
| 85 | KIRREL3 | 12571 | 7459 | 0.006 | 0.3095 | No | ||
| 86 | EPS8L2 | 18012 | 7460 | 0.006 | 0.3096 | No | ||
| 87 | PCDH7 | 7029 3460 | 7486 | 0.006 | 0.3084 | No | ||
| 88 | IMMT | 367 8029 | 7763 | 0.005 | 0.2935 | No | ||
| 89 | SDK1 | 16640 | 7824 | 0.005 | 0.2904 | No | ||
| 90 | ATP6V1B1 | 8499 | 7902 | 0.005 | 0.2863 | No | ||
| 91 | HCN1 | 21557 | 7934 | 0.005 | 0.2847 | No | ||
| 92 | ATP6AP2 | 2567 24380 2579 2656 | 7947 | 0.005 | 0.2842 | No | ||
| 93 | MASP1 | 1632 22626 | 8046 | 0.004 | 0.2789 | No | ||
| 94 | TBC1D15 | 19625 | 8053 | 0.004 | 0.2787 | No | ||
| 95 | MNT | 20784 | 8099 | 0.004 | 0.2763 | No | ||
| 96 | ABHD3 | 23491 | 8323 | 0.004 | 0.2643 | No | ||
| 97 | TBX6 | 1993 18075 | 8479 | 0.003 | 0.2560 | No | ||
| 98 | CX3CL1 | 9791 | 8685 | 0.003 | 0.2449 | No | ||
| 99 | NNAT | 2764 5180 | 8773 | 0.003 | 0.2403 | No | ||
| 100 | CDH16 | 4507 | 8955 | 0.002 | 0.2305 | No | ||
| 101 | RGS7 | 13743 | 9047 | 0.002 | 0.2256 | No | ||
| 102 | RPH3A | 13628 | 9228 | 0.002 | 0.2159 | No | ||
| 103 | SLC38A3 | 13319 | 9233 | 0.002 | 0.2157 | No | ||
| 104 | AUH | 21464 | 9305 | 0.001 | 0.2119 | No | ||
| 105 | CHAT | 21882 | 9349 | 0.001 | 0.2096 | No | ||
| 106 | YWHAH | 5937 10368 | 9813 | 0.000 | 0.1845 | No | ||
| 107 | GABPA | 8995 883 4744 | 9977 | -0.000 | 0.1756 | No | ||
| 108 | NCDN | 15756 2487 6471 | 10452 | -0.001 | 0.1500 | No | ||
| 109 | SH3GL2 | 9811 5429 | 10521 | -0.001 | 0.1463 | No | ||
| 110 | NDFIP1 | 23573 | 10927 | -0.002 | 0.1244 | No | ||
| 111 | RNF139 | 7992 13291 | 11281 | -0.003 | 0.1053 | No | ||
| 112 | CASQ2 | 15477 | 11315 | -0.003 | 0.1036 | No | ||
| 113 | INSR | 18950 | 11422 | -0.004 | 0.0979 | No | ||
| 114 | NPTX2 | 16630 | 11424 | -0.004 | 0.0980 | No | ||
| 115 | AMPD3 | 4383 2110 | 11808 | -0.005 | 0.0773 | No | ||
| 116 | RIN2 | 14817 | 11866 | -0.005 | 0.0743 | No | ||
| 117 | CACNB2 | 4466 8678 | 11878 | -0.005 | 0.0738 | No | ||
| 118 | SEMA3F | 18999 3097 | 11939 | -0.005 | 0.0706 | No | ||
| 119 | MAL | 2760 14437 | 11997 | -0.006 | 0.0677 | No | ||
| 120 | SLC2A4 | 20380 | 12102 | -0.006 | 0.0621 | No | ||
| 121 | HMGB3 | 9096 | 12507 | -0.008 | 0.0404 | No | ||
| 122 | SNPH | 14395 | 12521 | -0.008 | 0.0398 | No | ||
| 123 | DSCR1L1 | 23231 | 12845 | -0.009 | 0.0225 | No | ||
| 124 | SV2A | 15498 12250 | 12913 | -0.010 | 0.0191 | No | ||
| 125 | UQCRC1 | 10259 | 12986 | -0.010 | 0.0153 | No | ||
| 126 | PNOC | 21783 | 13074 | -0.011 | 0.0108 | No | ||
| 127 | MADD | 2926 14526 | 13136 | -0.011 | 0.0077 | No | ||
| 128 | LRFN5 | 6213 | 13441 | -0.013 | -0.0085 | No | ||
| 129 | SLC30A3 | 5995 | 14101 | -0.021 | -0.0438 | No | ||
| 130 | ZNF289 | 8060 13377 2750 8059 | 14204 | -0.023 | -0.0489 | No | ||
| 131 | FGF9 | 8966 | 14207 | -0.023 | -0.0486 | No | ||
| 132 | ATP6V0C | 8643 | 14304 | -0.025 | -0.0533 | No | ||
| 133 | CEL | 8734 | 14389 | -0.026 | -0.0574 | No | ||
| 134 | CNTN2 | 5627 | 14506 | -0.029 | -0.0631 | No | ||
| 135 | TNNI1 | 14116 | 14536 | -0.029 | -0.0641 | No | ||
| 136 | COX4I1 | 18444 | 14606 | -0.032 | -0.0672 | No | ||
| 137 | PCTK1 | 24369 | 14935 | -0.043 | -0.0842 | No | ||
| 138 | ATP5B | 19846 | 14963 | -0.044 | -0.0848 | No | ||
| 139 | FZD9 | 16345 | 15107 | -0.051 | -0.0916 | No | ||
| 140 | XYLT1 | 18113 | 15210 | -0.056 | -0.0960 | No | ||
| 141 | COX5B | 4552 | 15362 | -0.066 | -0.1029 | No | ||
| 142 | MEF2D | 9379 | 15725 | -0.101 | -0.1206 | No | ||
| 143 | ATP5C1 | 8635 | 15830 | -0.118 | -0.1240 | No | ||
| 144 | SLC6A8 | 24306 | 15837 | -0.119 | -0.1220 | No | ||
| 145 | RBMS1 | 14580 | 15872 | -0.124 | -0.1215 | No | ||
| 146 | FBXO18 | 14686 2796 | 15892 | -0.127 | -0.1201 | No | ||
| 147 | KIN | 15120 | 16140 | -0.177 | -0.1301 | No | ||
| 148 | MEF2C | 3204 9378 | 16157 | -0.182 | -0.1274 | No | ||
| 149 | BLCAP | 14371 | 16170 | -0.184 | -0.1245 | No | ||
| 150 | ALDOA | 8572 | 16216 | -0.195 | -0.1232 | No | ||
| 151 | VAMP2 | 20826 | 16297 | -0.215 | -0.1234 | No | ||
| 152 | CDKL5 | 24019 | 16305 | -0.216 | -0.1197 | No | ||
| 153 | ARF3 | 22137 | 16310 | -0.218 | -0.1157 | No | ||
| 154 | ZDHHC21 | 15854 | 16376 | -0.237 | -0.1147 | No | ||
| 155 | GABARAPL1 | 17268 | 16451 | -0.261 | -0.1137 | No | ||
| 156 | PANK1 | 1687 23692 | 16596 | -0.305 | -0.1156 | No | ||
| 157 | RAPGEF5 | 5739 10155 21123 | 16692 | -0.337 | -0.1143 | No | ||
| 158 | PPP3CC | 21763 | 16753 | -0.357 | -0.1107 | No | ||
| 159 | SLC9A7 | 24177 | 16757 | -0.359 | -0.1039 | No | ||
| 160 | GBAS | 16690 | 17185 | -0.521 | -0.1170 | No | ||
| 161 | MTCH2 | 14954 7120 | 17366 | -0.612 | -0.1150 | No | ||
| 162 | RANBP10 | 18764 | 17512 | -0.712 | -0.1092 | No | ||
| 163 | AK2 | 8563 2479 16073 | 17531 | -0.720 | -0.0963 | No | ||
| 164 | MSI2 | 8030 1268 | 17679 | -0.815 | -0.0886 | No | ||
| 165 | DEXI | 7193 12212 | 17741 | -0.863 | -0.0753 | No | ||
| 166 | ESRRA | 23802 | 17796 | -0.905 | -0.0609 | No | ||
| 167 | RAB3A | 9681 | 17940 | -1.048 | -0.0485 | No | ||
| 168 | CHD2 | 10847 | 18049 | -1.174 | -0.0318 | No | ||
| 169 | UBL3 | 16284 | 18166 | -1.334 | -0.0124 | No | ||
| 170 | ATP1B1 | 4420 | 18390 | -1.913 | 0.0123 | No |