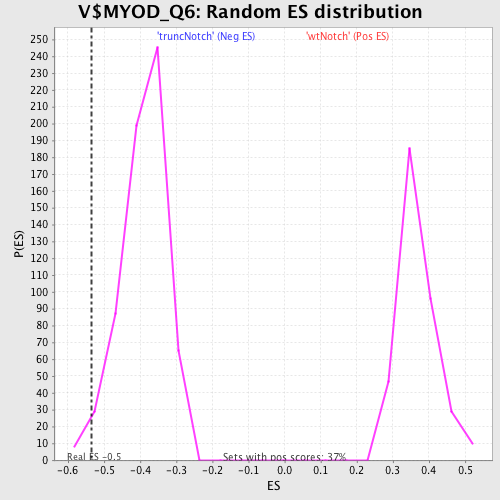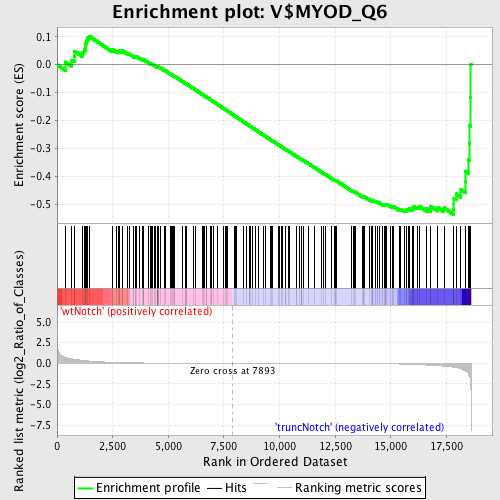
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_03_truncNotch_versus_wtNotch.phenotype_truncNotch_versus_wtNotch.cls #wtNotch_versus_truncNotch.phenotype_truncNotch_versus_wtNotch.cls #wtNotch_versus_truncNotch_repos |
| Phenotype | phenotype_truncNotch_versus_wtNotch.cls#wtNotch_versus_truncNotch_repos |
| Upregulated in class | truncNotch |
| GeneSet | V$MYOD_Q6 |
| Enrichment Score (ES) | -0.53597873 |
| Normalized Enrichment Score (NES) | -1.3711029 |
| Nominal p-value | 0.018957347 |
| FDR q-value | 0.7166681 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SPOP | 450035 | 364 | 0.722 | 0.0097 | No | ||
| 2 | NEDD4L | 6380368 | 661 | 0.535 | 0.0155 | No | ||
| 3 | ATP1B1 | 3130594 | 766 | 0.483 | 0.0296 | No | ||
| 4 | MAP3K7IP2 | 2340242 | 775 | 0.481 | 0.0488 | No | ||
| 5 | GALNT2 | 2260279 4060239 | 1140 | 0.351 | 0.0434 | No | ||
| 6 | H2AFZ | 1470168 | 1209 | 0.332 | 0.0532 | No | ||
| 7 | OSR1 | 1500025 | 1264 | 0.319 | 0.0633 | No | ||
| 8 | TLK2 | 1170161 | 1288 | 0.312 | 0.0748 | No | ||
| 9 | PLAGL2 | 1770148 | 1306 | 0.305 | 0.0863 | No | ||
| 10 | TMEM24 | 5860039 | 1346 | 0.295 | 0.0962 | No | ||
| 11 | KCNH2 | 1170451 | 1437 | 0.268 | 0.1023 | No | ||
| 12 | MMP11 | 1980619 | 2474 | 0.108 | 0.0505 | No | ||
| 13 | WNT6 | 4570364 | 2497 | 0.106 | 0.0537 | No | ||
| 14 | CHRNB1 | 1190671 | 2664 | 0.092 | 0.0484 | No | ||
| 15 | ACADSB | 520170 | 2765 | 0.084 | 0.0464 | No | ||
| 16 | STRN3 | 1450093 6200685 | 2806 | 0.081 | 0.0476 | No | ||
| 17 | PKP4 | 3710400 5890097 | 2809 | 0.081 | 0.0508 | No | ||
| 18 | STK3 | 7100427 | 2932 | 0.072 | 0.0471 | No | ||
| 19 | SMARCA5 | 6620050 | 2938 | 0.072 | 0.0498 | No | ||
| 20 | GALR3 | 4850113 | 3182 | 0.056 | 0.0389 | No | ||
| 21 | ADAM11 | 1050008 3130494 | 3259 | 0.052 | 0.0369 | No | ||
| 22 | ZIC4 | 1500082 | 3443 | 0.044 | 0.0288 | No | ||
| 23 | BLR1 | 1500300 5420017 | 3445 | 0.044 | 0.0305 | No | ||
| 24 | NRF1 | 2650195 | 3507 | 0.042 | 0.0289 | No | ||
| 25 | RIN1 | 510593 | 3517 | 0.041 | 0.0301 | No | ||
| 26 | NLK | 2030010 2450041 | 3545 | 0.040 | 0.0303 | No | ||
| 27 | HS6ST3 | 5910138 | 3701 | 0.034 | 0.0233 | No | ||
| 28 | LCN9 | 5050450 | 3845 | 0.030 | 0.0168 | No | ||
| 29 | FBXO32 | 110037 610750 | 3896 | 0.029 | 0.0153 | No | ||
| 30 | HTR7 | 2680176 | 3897 | 0.029 | 0.0164 | No | ||
| 31 | EIF2C1 | 6900551 | 3904 | 0.029 | 0.0173 | No | ||
| 32 | SSH2 | 360056 | 4096 | 0.024 | 0.0079 | No | ||
| 33 | TRIM3 | 5570156 | 4213 | 0.022 | 0.0026 | No | ||
| 34 | ASB14 | 5860427 | 4240 | 0.022 | 0.0020 | No | ||
| 35 | MID1 | 4070114 4760458 | 4285 | 0.021 | 0.0005 | No | ||
| 36 | CER1 | 840102 | 4379 | 0.020 | -0.0037 | No | ||
| 37 | PLCB1 | 2570050 7100102 | 4422 | 0.019 | -0.0052 | No | ||
| 38 | FGF17 | 3130022 | 4490 | 0.018 | -0.0081 | No | ||
| 39 | SREBF2 | 3390692 | 4495 | 0.018 | -0.0076 | No | ||
| 40 | USP15 | 610592 3520504 | 4508 | 0.018 | -0.0075 | No | ||
| 41 | COL4A4 | 1050541 | 4520 | 0.018 | -0.0074 | No | ||
| 42 | RHOBTB1 | 1410010 | 4528 | 0.018 | -0.0070 | No | ||
| 43 | TJP1 | 6350184 | 4547 | 0.017 | -0.0073 | No | ||
| 44 | ITGA6 | 3830129 | 4560 | 0.017 | -0.0073 | No | ||
| 45 | PPARGC1A | 4670040 | 4653 | 0.016 | -0.0116 | No | ||
| 46 | VKORC1L1 | 110220 1400050 | 4817 | 0.014 | -0.0199 | No | ||
| 47 | KCNE1L | 4670092 | 4877 | 0.014 | -0.0225 | No | ||
| 48 | KCNN3 | 6520600 6420138 | 5093 | 0.012 | -0.0337 | No | ||
| 49 | HR | 2690095 6200300 | 5150 | 0.011 | -0.0362 | No | ||
| 50 | FUT8 | 1340068 2340056 | 5196 | 0.011 | -0.0382 | No | ||
| 51 | SEMA4C | 1980315 | 5227 | 0.011 | -0.0394 | No | ||
| 52 | DSCAM | 1780050 2450731 2810438 | 5269 | 0.011 | -0.0412 | No | ||
| 53 | EMX1 | 6130537 | 5285 | 0.010 | -0.0416 | No | ||
| 54 | DSCAML1 | 1850403 5550079 | 5653 | 0.008 | -0.0611 | No | ||
| 55 | DCT | 1090347 3840494 | 5754 | 0.008 | -0.0663 | No | ||
| 56 | ANKRD2 | 6590427 | 5790 | 0.007 | -0.0679 | No | ||
| 57 | STARD13 | 540722 | 5793 | 0.007 | -0.0677 | No | ||
| 58 | LIN28 | 6590672 | 5827 | 0.007 | -0.0692 | No | ||
| 59 | GRID2 | 2680242 | 6135 | 0.006 | -0.0856 | No | ||
| 60 | KCNQ4 | 3800438 7050400 | 6238 | 0.005 | -0.0909 | No | ||
| 61 | TREX2 | 4920707 | 6535 | 0.004 | -0.1067 | No | ||
| 62 | NPEPPS | 2630731 | 6563 | 0.004 | -0.1080 | No | ||
| 63 | COL4A3 | 5910075 | 6607 | 0.004 | -0.1102 | No | ||
| 64 | NEB | 580735 | 6693 | 0.004 | -0.1147 | No | ||
| 65 | NBL1 | 2480021 | 6734 | 0.003 | -0.1167 | No | ||
| 66 | B3GALT2 | 430136 | 6876 | 0.003 | -0.1242 | No | ||
| 67 | GUCY1A3 | 110253 2640735 4070037 | 6919 | 0.003 | -0.1264 | No | ||
| 68 | SLC12A5 | 1980692 | 6961 | 0.003 | -0.1285 | No | ||
| 69 | CHRNG | 5860273 | 7035 | 0.002 | -0.1323 | No | ||
| 70 | NHLH1 | 6100452 | 7193 | 0.002 | -0.1408 | No | ||
| 71 | FBXW11 | 6450632 | 7459 | 0.001 | -0.1551 | No | ||
| 72 | RASGRF2 | 1990047 | 7574 | 0.001 | -0.1612 | No | ||
| 73 | PTPRJ | 4010707 | 7635 | 0.001 | -0.1645 | No | ||
| 74 | CHCHD3 | 4200044 | 7675 | 0.001 | -0.1665 | No | ||
| 75 | CCNL2 | 6520167 | 7961 | -0.000 | -0.1820 | No | ||
| 76 | EFNA1 | 3840672 | 8028 | -0.000 | -0.1855 | No | ||
| 77 | DPF3 | 2650128 2970168 | 8043 | -0.000 | -0.1863 | No | ||
| 78 | FOXI1 | 3710095 6620195 | 8372 | -0.001 | -0.2040 | No | ||
| 79 | CHRND | 840403 2260670 | 8515 | -0.002 | -0.2116 | No | ||
| 80 | ASB15 | 450563 | 8636 | -0.002 | -0.2180 | No | ||
| 81 | MTUS1 | 780348 4920609 | 8638 | -0.002 | -0.2180 | No | ||
| 82 | ABL2 | 580021 | 8680 | -0.002 | -0.2201 | No | ||
| 83 | SOST | 1170195 | 8793 | -0.003 | -0.2261 | No | ||
| 84 | ARL4A | 6290091 | 8931 | -0.003 | -0.2334 | No | ||
| 85 | TAC4 | 4730070 | 9070 | -0.003 | -0.2408 | No | ||
| 86 | TRPV3 | 540341 | 9289 | -0.004 | -0.2524 | No | ||
| 87 | USF1 | 4280156 4610114 6370113 | 9354 | -0.004 | -0.2557 | No | ||
| 88 | MAP3K13 | 3190017 | 9570 | -0.005 | -0.2672 | No | ||
| 89 | PAK3 | 4210136 | 9648 | -0.005 | -0.2711 | No | ||
| 90 | GATA2 | 6590280 | 9658 | -0.005 | -0.2714 | No | ||
| 91 | ELAVL3 | 2850014 | 9961 | -0.006 | -0.2875 | No | ||
| 92 | HDAC9 | 1990010 2260133 | 10013 | -0.006 | -0.2901 | No | ||
| 93 | ARHGEF2 | 3360577 | 10082 | -0.006 | -0.2935 | No | ||
| 94 | TTN | 2320161 4670056 6550026 | 10134 | -0.007 | -0.2960 | No | ||
| 95 | TGM1 | 1090500 | 10251 | -0.007 | -0.3020 | No | ||
| 96 | INHA | 6100102 | 10271 | -0.007 | -0.3027 | No | ||
| 97 | CMYA1 | 6900632 | 10404 | -0.008 | -0.3096 | No | ||
| 98 | ATOH7 | 5550121 | 10460 | -0.008 | -0.3122 | No | ||
| 99 | LRAT | 5130603 | 10743 | -0.009 | -0.3271 | No | ||
| 100 | ALG6 | 4060131 | 10752 | -0.009 | -0.3272 | No | ||
| 101 | MYOZ3 | 1990138 | 10895 | -0.010 | -0.3345 | No | ||
| 102 | ANK2 | 6510546 | 10963 | -0.010 | -0.3377 | No | ||
| 103 | HSD11B2 | 5900053 | 10970 | -0.010 | -0.3377 | No | ||
| 104 | CDON | 540204 | 10987 | -0.010 | -0.3381 | No | ||
| 105 | KAZALD1 | 380603 | 11091 | -0.011 | -0.3433 | No | ||
| 106 | ERBB4 | 4610400 4810017 | 11308 | -0.012 | -0.3545 | No | ||
| 107 | BEX2 | 5670433 | 11569 | -0.013 | -0.3681 | No | ||
| 108 | ERBB3 | 5890372 | 11878 | -0.015 | -0.3842 | No | ||
| 109 | CASQ1 | 6290035 7000707 | 11953 | -0.015 | -0.3875 | No | ||
| 110 | CYP26A1 | 380022 2690600 | 12057 | -0.016 | -0.3925 | No | ||
| 111 | POU3F3 | 6400047 | 12323 | -0.018 | -0.4061 | No | ||
| 112 | AP4S1 | 4730292 | 12481 | -0.020 | -0.4138 | No | ||
| 113 | FSCN2 | 1230039 | 12490 | -0.020 | -0.4134 | No | ||
| 114 | NPPC | 2320647 | 12504 | -0.020 | -0.4133 | No | ||
| 115 | RIMS2 | 670725 | 12541 | -0.020 | -0.4145 | No | ||
| 116 | CUGBP1 | 450292 510022 7050176 7050215 | 13251 | -0.028 | -0.4517 | No | ||
| 117 | NOL4 | 5050446 5360519 | 13315 | -0.029 | -0.4540 | No | ||
| 118 | PROX1 | 4210193 | 13362 | -0.030 | -0.4553 | No | ||
| 119 | MLLT6 | 3870168 | 13426 | -0.031 | -0.4574 | No | ||
| 120 | CASKIN2 | 2030113 | 13732 | -0.036 | -0.4725 | No | ||
| 121 | HN1 | 3360156 | 13757 | -0.036 | -0.4723 | No | ||
| 122 | SYNGR1 | 5130452 6650056 | 13763 | -0.036 | -0.4711 | No | ||
| 123 | PRKAG2 | 2340100 | 13820 | -0.037 | -0.4726 | No | ||
| 124 | GGN | 1410528 4210082 5290170 | 14054 | -0.042 | -0.4835 | No | ||
| 125 | RND1 | 5080300 | 14147 | -0.045 | -0.4867 | No | ||
| 126 | NKX6-2 | 2450332 3120333 | 14168 | -0.045 | -0.4859 | No | ||
| 127 | MDGA1 | 1170139 | 14197 | -0.046 | -0.4856 | No | ||
| 128 | PHACTR3 | 3850435 5900445 | 14303 | -0.048 | -0.4893 | No | ||
| 129 | TBR1 | 5130053 | 14393 | -0.050 | -0.4921 | No | ||
| 130 | CRAT | 540020 2060364 3520148 | 14478 | -0.053 | -0.4945 | No | ||
| 131 | SCG2 | 5670438 130671 | 14639 | -0.058 | -0.5007 | No | ||
| 132 | POFUT1 | 1570458 | 14707 | -0.060 | -0.5019 | No | ||
| 133 | NR2E1 | 4730288 | 14740 | -0.061 | -0.5012 | No | ||
| 134 | USP2 | 1190292 1240253 4850035 | 14758 | -0.062 | -0.4996 | No | ||
| 135 | ELAVL4 | 50735 3360086 5220167 | 14810 | -0.064 | -0.4997 | No | ||
| 136 | PDGFB | 3060440 6370008 | 14969 | -0.070 | -0.5054 | No | ||
| 137 | SYT4 | 5080193 | 15095 | -0.076 | -0.5091 | No | ||
| 138 | ACTN3 | 3140541 6480598 | 15137 | -0.078 | -0.5081 | No | ||
| 139 | ITGB4 | 1740021 3840482 | 15402 | -0.093 | -0.5187 | No | ||
| 140 | TNFRSF21 | 6380100 | 15441 | -0.096 | -0.5168 | No | ||
| 141 | ITPR3 | 4010632 | 15594 | -0.105 | -0.5208 | No | ||
| 142 | INVS | 4200632 6620528 | 15703 | -0.113 | -0.5220 | Yes | ||
| 143 | SIPA1 | 5220687 | 15726 | -0.115 | -0.5185 | Yes | ||
| 144 | NCKIPSD | 2940156 | 15787 | -0.120 | -0.5168 | Yes | ||
| 145 | SPI1 | 1410397 | 15838 | -0.125 | -0.5144 | Yes | ||
| 146 | CDH1 | 1940736 | 15952 | -0.136 | -0.5150 | Yes | ||
| 147 | DEF6 | 840593 | 16024 | -0.144 | -0.5130 | Yes | ||
| 148 | TEC | 1400576 | 16025 | -0.144 | -0.5071 | Yes | ||
| 149 | BCL9 | 7100112 | 16185 | -0.161 | -0.5092 | Yes | ||
| 150 | CRELD1 | 2060253 | 16307 | -0.176 | -0.5086 | Yes | ||
| 151 | PPP2R4 | 670280 2900097 | 16595 | -0.213 | -0.5154 | Yes | ||
| 152 | ARID5A | 2450133 | 16789 | -0.242 | -0.5161 | Yes | ||
| 153 | TAGAP | 3290433 | 16791 | -0.242 | -0.5062 | Yes | ||
| 154 | PSTPIP1 | 3190156 | 17111 | -0.293 | -0.5116 | Yes | ||
| 155 | HMGA1 | 6580408 | 17390 | -0.349 | -0.5124 | Yes | ||
| 156 | PRKACA | 2640731 4050048 | 17826 | -0.471 | -0.5168 | Yes | ||
| 157 | GFRA1 | 3610152 | 17836 | -0.478 | -0.4977 | Yes | ||
| 158 | ELOVL1 | 2450019 4760138 5340685 | 17838 | -0.478 | -0.4783 | Yes | ||
| 159 | SELL | 1190500 | 17938 | -0.519 | -0.4625 | Yes | ||
| 160 | POLD4 | 2320082 | 18150 | -0.660 | -0.4470 | Yes | ||
| 161 | GMPR | 3990022 | 18346 | -0.933 | -0.4196 | Yes | ||
| 162 | HEYL | 5860528 | 18352 | -0.937 | -0.3816 | Yes | ||
| 163 | TUFT1 | 540047 2370010 | 18473 | -1.181 | -0.3400 | Yes | ||
| 164 | HES6 | 540411 6550504 | 18520 | -1.504 | -0.2812 | Yes | ||
| 165 | CKM | 1450524 | 18536 | -1.575 | -0.2177 | Yes | ||
| 166 | IGF2 | 6510020 | 18590 | -2.564 | -0.1161 | Yes | ||
| 167 | GNAS | 630441 1850373 4050152 | 18597 | -2.879 | 0.0010 | Yes |