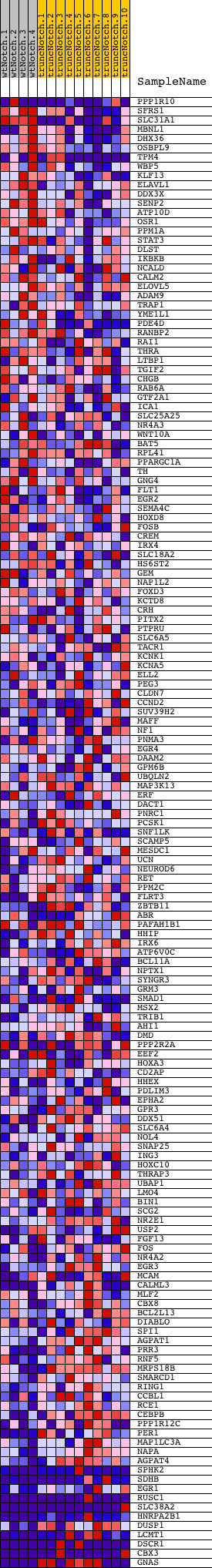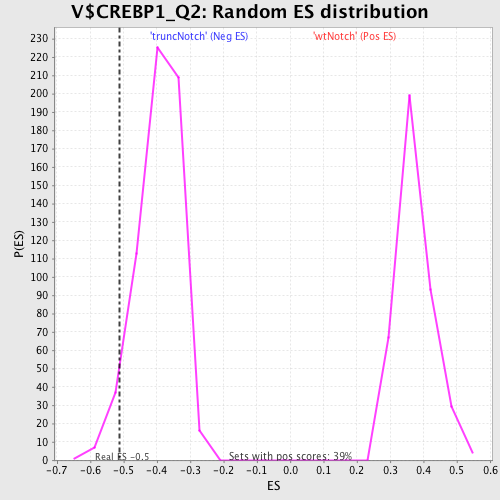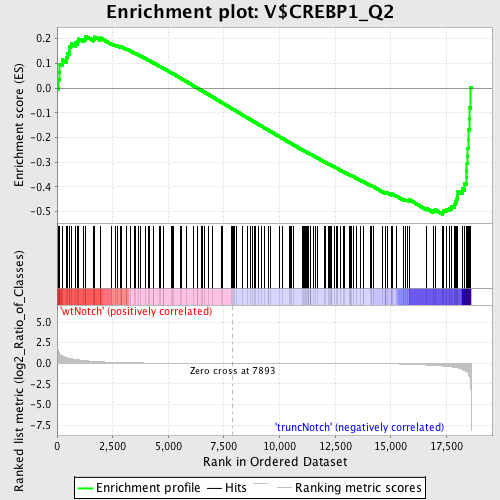
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_03_truncNotch_versus_wtNotch.phenotype_truncNotch_versus_wtNotch.cls #wtNotch_versus_truncNotch.phenotype_truncNotch_versus_wtNotch.cls #wtNotch_versus_truncNotch_repos |
| Phenotype | phenotype_truncNotch_versus_wtNotch.cls#wtNotch_versus_truncNotch_repos |
| Upregulated in class | truncNotch |
| GeneSet | V$CREBP1_Q2 |
| Enrichment Score (ES) | -0.51126 |
| Normalized Enrichment Score (NES) | -1.3003324 |
| Nominal p-value | 0.049342107 |
| FDR q-value | 0.9342972 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PPP1R10 | 110136 3170725 3610484 | 77 | 1.334 | 0.0340 | No | ||
| 2 | SFRS1 | 2360440 | 108 | 1.134 | 0.0648 | No | ||
| 3 | SLC31A1 | 4590161 6940154 | 124 | 1.090 | 0.0951 | No | ||
| 4 | MBNL1 | 2640762 7100048 | 235 | 0.865 | 0.1138 | No | ||
| 5 | DHX36 | 2470465 | 436 | 0.659 | 0.1218 | No | ||
| 6 | OSBPL9 | 2570373 | 460 | 0.647 | 0.1391 | No | ||
| 7 | TPM4 | 2100059 3850072 4920162 | 537 | 0.604 | 0.1522 | No | ||
| 8 | WBP5 | 7050563 | 566 | 0.585 | 0.1674 | No | ||
| 9 | KLF13 | 4670563 | 625 | 0.551 | 0.1800 | No | ||
| 10 | ELAVL1 | 4730497 | 820 | 0.464 | 0.1828 | No | ||
| 11 | DDX3X | 2190020 | 934 | 0.421 | 0.1887 | No | ||
| 12 | SENP2 | 4540452 | 974 | 0.407 | 0.1982 | No | ||
| 13 | ATP10D | 4850139 | 1192 | 0.336 | 0.1961 | No | ||
| 14 | OSR1 | 1500025 | 1264 | 0.319 | 0.2013 | No | ||
| 15 | PPM1A | 1170301 5900273 | 1276 | 0.317 | 0.2098 | No | ||
| 16 | STAT3 | 460040 3710341 | 1643 | 0.227 | 0.1964 | No | ||
| 17 | DLST | 2320722 | 1680 | 0.221 | 0.2008 | No | ||
| 18 | IKBKB | 6840072 | 1694 | 0.218 | 0.2063 | No | ||
| 19 | NCALD | 730176 | 1939 | 0.177 | 0.1982 | No | ||
| 20 | CALM2 | 6620463 | 1943 | 0.176 | 0.2030 | No | ||
| 21 | ELOVL5 | 3800170 | 2460 | 0.110 | 0.1782 | No | ||
| 22 | ADAM9 | 3360411 | 2609 | 0.096 | 0.1730 | No | ||
| 23 | TRAP1 | 6040168 | 2705 | 0.089 | 0.1703 | No | ||
| 24 | YME1L1 | 2360102 2510520 2650451 | 2844 | 0.079 | 0.1651 | No | ||
| 25 | PDE4D | 2470528 6660014 | 2880 | 0.076 | 0.1654 | No | ||
| 26 | RANBP2 | 4280338 | 2886 | 0.075 | 0.1673 | No | ||
| 27 | RAI1 | 110377 5340670 | 3123 | 0.060 | 0.1562 | No | ||
| 28 | THRA | 6770341 110164 1990600 | 3137 | 0.059 | 0.1572 | No | ||
| 29 | LTBP1 | 780035 2100746 2760402 3780551 6550184 6860193 | 3289 | 0.050 | 0.1505 | No | ||
| 30 | TGIF2 | 3840364 4050520 | 3482 | 0.043 | 0.1413 | No | ||
| 31 | CHGB | 1980053 | 3528 | 0.041 | 0.1400 | No | ||
| 32 | RAB6A | 3840463 4780731 | 3663 | 0.035 | 0.1338 | No | ||
| 33 | GTF2A1 | 4120138 4920524 5390110 1230397 | 3729 | 0.034 | 0.1312 | No | ||
| 34 | ICA1 | 3830059 | 3973 | 0.027 | 0.1188 | No | ||
| 35 | SLC25A25 | 1230605 | 4103 | 0.024 | 0.1125 | No | ||
| 36 | NR4A3 | 2900021 5860095 5910039 | 4134 | 0.024 | 0.1116 | No | ||
| 37 | WNT10A | 2100706 | 4314 | 0.021 | 0.1025 | No | ||
| 38 | BAT5 | 5720687 | 4347 | 0.020 | 0.1013 | No | ||
| 39 | RPL41 | 6940112 | 4590 | 0.017 | 0.0887 | No | ||
| 40 | PPARGC1A | 4670040 | 4653 | 0.016 | 0.0858 | No | ||
| 41 | TH | 2100056 | 4778 | 0.015 | 0.0795 | No | ||
| 42 | GNG4 | 2630600 | 4796 | 0.015 | 0.0790 | No | ||
| 43 | FLT1 | 3830167 4920438 | 5123 | 0.012 | 0.0617 | No | ||
| 44 | EGR2 | 3800403 | 5194 | 0.011 | 0.0582 | No | ||
| 45 | SEMA4C | 1980315 | 5227 | 0.011 | 0.0568 | No | ||
| 46 | HOXD8 | 3710156 | 5567 | 0.009 | 0.0386 | No | ||
| 47 | FOSB | 1940142 | 5612 | 0.008 | 0.0365 | No | ||
| 48 | CREM | 840156 6380438 6660041 6660168 | 5828 | 0.007 | 0.0250 | No | ||
| 49 | IRX4 | 1980102 | 6110 | 0.006 | 0.0100 | No | ||
| 50 | SLC18A2 | 6420541 | 6319 | 0.005 | -0.0011 | No | ||
| 51 | HS6ST2 | 520133 5550603 | 6323 | 0.005 | -0.0012 | No | ||
| 52 | GEM | 5290082 | 6475 | 0.004 | -0.0092 | No | ||
| 53 | NAP1L2 | 130047 | 6522 | 0.004 | -0.0116 | No | ||
| 54 | FOXD3 | 6550156 | 6611 | 0.004 | -0.0162 | No | ||
| 55 | KCTD8 | 6660593 | 6822 | 0.003 | -0.0275 | No | ||
| 56 | CRH | 3710301 | 6975 | 0.003 | -0.0357 | No | ||
| 57 | PITX2 | 870537 2690139 | 7394 | 0.001 | -0.0583 | No | ||
| 58 | PTPRU | 3170180 | 7424 | 0.001 | -0.0598 | No | ||
| 59 | SLC6A5 | 870121 | 7826 | 0.000 | -0.0815 | No | ||
| 60 | TACR1 | 70358 3840411 | 7875 | 0.000 | -0.0841 | No | ||
| 61 | KCNK1 | 1050068 | 7920 | -0.000 | -0.0865 | No | ||
| 62 | KCNA5 | 5270750 | 7979 | -0.000 | -0.0896 | No | ||
| 63 | ELL2 | 6420091 | 8041 | -0.000 | -0.0929 | No | ||
| 64 | PEG3 | 3710020 | 8338 | -0.001 | -0.1089 | No | ||
| 65 | CLDN7 | 1190301 | 8575 | -0.002 | -0.1217 | No | ||
| 66 | CCND2 | 5340167 | 8714 | -0.002 | -0.1291 | No | ||
| 67 | SUV39H2 | 1450672 6370242 | 8802 | -0.003 | -0.1337 | No | ||
| 68 | MAFF | 4850577 5720286 | 8885 | -0.003 | -0.1381 | No | ||
| 69 | NF1 | 6980433 | 8907 | -0.003 | -0.1391 | No | ||
| 70 | PNMA3 | 3170309 | 9062 | -0.003 | -0.1474 | No | ||
| 71 | EGR4 | 3120750 | 9179 | -0.004 | -0.1536 | No | ||
| 72 | DAAM2 | 1400358 7650669 | 9340 | -0.004 | -0.1621 | No | ||
| 73 | GPM6B | 6220458 | 9482 | -0.005 | -0.1696 | No | ||
| 74 | UBQLN2 | 430397 1170594 4070121 | 9490 | -0.005 | -0.1699 | No | ||
| 75 | MAP3K13 | 3190017 | 9570 | -0.005 | -0.1740 | No | ||
| 76 | ERF | 110551 1770278 | 9994 | -0.006 | -0.1968 | No | ||
| 77 | DACT1 | 1500050 | 10116 | -0.007 | -0.2031 | No | ||
| 78 | PNRC1 | 360025 | 10438 | -0.008 | -0.2203 | No | ||
| 79 | PCSK1 | 2350148 | 10489 | -0.008 | -0.2228 | No | ||
| 80 | SNF1LK | 6110403 | 10547 | -0.008 | -0.2256 | No | ||
| 81 | SCAMP5 | 5550368 6290021 | 10609 | -0.008 | -0.2287 | No | ||
| 82 | MESDC1 | 2030019 | 11048 | -0.010 | -0.2521 | No | ||
| 83 | UCN | 6620338 | 11062 | -0.010 | -0.2525 | No | ||
| 84 | NEUROD6 | 4670731 | 11115 | -0.011 | -0.2551 | No | ||
| 85 | RET | 3850746 | 11163 | -0.011 | -0.2573 | No | ||
| 86 | PPM2C | 3360209 | 11219 | -0.011 | -0.2600 | No | ||
| 87 | FLRT3 | 70685 2760497 6040519 | 11275 | -0.011 | -0.2626 | No | ||
| 88 | ZBTB11 | 6520092 | 11280 | -0.011 | -0.2625 | No | ||
| 89 | ABR | 610079 1170609 3610195 5670050 | 11310 | -0.012 | -0.2637 | No | ||
| 90 | PAFAH1B1 | 4230333 6420121 6450066 | 11376 | -0.012 | -0.2669 | No | ||
| 91 | HHIP | 6940438 | 11385 | -0.012 | -0.2670 | No | ||
| 92 | IRX6 | 4590025 | 11410 | -0.012 | -0.2680 | No | ||
| 93 | ATP6V0C | 1780609 | 11525 | -0.013 | -0.2738 | No | ||
| 94 | BCL11A | 6860369 | 11620 | -0.013 | -0.2785 | No | ||
| 95 | NPTX1 | 3710736 | 11686 | -0.014 | -0.2816 | No | ||
| 96 | SYNGR3 | 4590576 | 11997 | -0.016 | -0.2980 | No | ||
| 97 | GRM3 | 4540242 | 12077 | -0.016 | -0.3018 | No | ||
| 98 | SMAD1 | 630537 1850333 | 12217 | -0.017 | -0.3088 | No | ||
| 99 | MSX2 | 1570022 | 12236 | -0.017 | -0.3093 | No | ||
| 100 | TRIB1 | 2320435 | 12289 | -0.018 | -0.3116 | No | ||
| 101 | AHI1 | 460520 | 12312 | -0.018 | -0.3123 | No | ||
| 102 | DMD | 1740041 3990332 | 12328 | -0.018 | -0.3126 | No | ||
| 103 | PPP2R2A | 2900014 5700575 | 12489 | -0.020 | -0.3207 | No | ||
| 104 | EEF2 | 1050369 4670035 5890598 | 12561 | -0.020 | -0.3240 | No | ||
| 105 | HOXA3 | 3610632 6020358 | 12594 | -0.021 | -0.3251 | No | ||
| 106 | CD2AP | 1940369 | 12742 | -0.022 | -0.3324 | No | ||
| 107 | HHEX | 2340575 | 12859 | -0.023 | -0.3380 | No | ||
| 108 | PDLIM3 | 1410095 | 12906 | -0.024 | -0.3399 | No | ||
| 109 | EPHA2 | 5890056 | 13132 | -0.027 | -0.3513 | No | ||
| 110 | GPR3 | 2970452 | 13186 | -0.027 | -0.3534 | No | ||
| 111 | DDX51 | 5550044 6350270 6590750 | 13196 | -0.027 | -0.3531 | No | ||
| 112 | SLC6A4 | 2230372 | 13246 | -0.028 | -0.3549 | No | ||
| 113 | NOL4 | 5050446 5360519 | 13315 | -0.029 | -0.3578 | No | ||
| 114 | SNAP25 | 360520 | 13442 | -0.031 | -0.3637 | No | ||
| 115 | ING3 | 110438 2060390 | 13616 | -0.034 | -0.3721 | No | ||
| 116 | HOXC10 | 5360170 | 13767 | -0.036 | -0.3792 | No | ||
| 117 | THRAP3 | 6180309 | 13781 | -0.037 | -0.3789 | No | ||
| 118 | UBAP1 | 4280059 | 14067 | -0.043 | -0.3931 | No | ||
| 119 | LMO4 | 3800746 | 14148 | -0.045 | -0.3962 | No | ||
| 120 | BIN1 | 5420348 5670500 | 14218 | -0.046 | -0.3986 | No | ||
| 121 | SCG2 | 5670438 130671 | 14639 | -0.058 | -0.4197 | No | ||
| 122 | NR2E1 | 4730288 | 14740 | -0.061 | -0.4233 | No | ||
| 123 | USP2 | 1190292 1240253 4850035 | 14758 | -0.062 | -0.4225 | No | ||
| 124 | FGF13 | 630575 1570440 5360121 | 14771 | -0.063 | -0.4213 | No | ||
| 125 | FOS | 1850315 | 14838 | -0.065 | -0.4230 | No | ||
| 126 | NR4A2 | 60273 | 15039 | -0.073 | -0.4318 | No | ||
| 127 | EGR3 | 6940128 | 15044 | -0.073 | -0.4299 | No | ||
| 128 | MCAM | 5130270 6620577 | 15055 | -0.074 | -0.4283 | No | ||
| 129 | CALML3 | 1980593 | 15267 | -0.085 | -0.4373 | No | ||
| 130 | MLF2 | 1940176 | 15564 | -0.103 | -0.4504 | No | ||
| 131 | CBX8 | 50671 | 15674 | -0.111 | -0.4532 | No | ||
| 132 | BCL2L13 | 3170593 | 15732 | -0.116 | -0.4529 | No | ||
| 133 | DIABLO | 2360170 | 15828 | -0.124 | -0.4545 | No | ||
| 134 | SPI1 | 1410397 | 15838 | -0.125 | -0.4514 | No | ||
| 135 | AGPAT1 | 610056 | 16609 | -0.215 | -0.4870 | No | ||
| 136 | PRR3 | 3440112 | 16907 | -0.260 | -0.4957 | No | ||
| 137 | RNF5 | 2760088 | 16985 | -0.271 | -0.4921 | No | ||
| 138 | MRPS18B | 5570086 | 17340 | -0.340 | -0.5015 | Yes | ||
| 139 | SMARCD1 | 3060193 3850184 6400369 | 17376 | -0.347 | -0.4935 | Yes | ||
| 140 | RING1 | 4560180 | 17518 | -0.374 | -0.4905 | Yes | ||
| 141 | CCBL1 | 2350082 | 17636 | -0.409 | -0.4851 | Yes | ||
| 142 | RCE1 | 1980372 | 17732 | -0.440 | -0.4777 | Yes | ||
| 143 | CEBPB | 2970019 | 17856 | -0.485 | -0.4705 | Yes | ||
| 144 | PPP1R12C | 580309 4760131 | 17897 | -0.500 | -0.4584 | Yes | ||
| 145 | PER1 | 3290315 | 17967 | -0.532 | -0.4469 | Yes | ||
| 146 | MAP1LC3A | 840519 5570605 | 18008 | -0.555 | -0.4332 | Yes | ||
| 147 | NAPA | 1780537 5720102 | 18014 | -0.561 | -0.4174 | Yes | ||
| 148 | AGPAT4 | 6040079 | 18213 | -0.728 | -0.4074 | Yes | ||
| 149 | SPHK2 | 2030487 4120670 4850114 | 18315 | -0.894 | -0.3873 | Yes | ||
| 150 | SDHB | 3120398 4780113 3710435 | 18391 | -0.996 | -0.3629 | Yes | ||
| 151 | EGR1 | 4610347 | 18402 | -1.010 | -0.3346 | Yes | ||
| 152 | RUSC1 | 6450093 6980575 | 18422 | -1.058 | -0.3053 | Yes | ||
| 153 | SLC38A2 | 1470242 3800026 | 18436 | -1.086 | -0.2750 | Yes | ||
| 154 | HNRPA2B1 | 1240619 6200161 6200494 6980408 | 18464 | -1.154 | -0.2435 | Yes | ||
| 155 | DUSP1 | 6860121 | 18501 | -1.337 | -0.2072 | Yes | ||
| 156 | LCMT1 | 2230551 3830497 | 18504 | -1.364 | -0.1683 | Yes | ||
| 157 | DSCR1 | 4540048 6220039 | 18535 | -1.573 | -0.1250 | Yes | ||
| 158 | CBX3 | 1780592 3940008 | 18544 | -1.645 | -0.0784 | Yes | ||
| 159 | GNAS | 630441 1850373 4050152 | 18597 | -2.879 | 0.0010 | Yes |